| n.genes | n.arrays | score | residual | profile | network | motif1 | motif2 | motif.posns | orf.names | short.names | long.names |
| 205 | 5 | 433 | -7.903 | 0.186 | 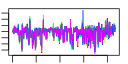 |  | 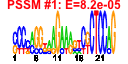 |  |  | YHL033C YLL045C YML073C YKL180W YML063W | RPL8A RPL8B RPL6A RPL17A RPS1B | Ribosomal protein L4 of the large (60S) ribosomal subunit, nearly identical to Rpl8Bp and has similarity to rat L7a ribosomal protein; mutation results in decreased amounts of free 60S subunits | Ribosomal protein L4 of the large (60S) ribosomal subunit, nearly identical to Rpl8Ap and has similarity to rat L7a ribosomal protein; mutation results in decreased amounts of free 60S subunits | N-terminally acetylated protein component of the large (60S) ribosomal subunit, has similarity to Rpl6Bp and to rat L6 ribosomal protein; binds to 5.8S rRNA | Protein component of the large (60S) ribosomal subunit, nearly identical to Rpl17Bp and has similarity to E. coli L22 and rat L17 ribosomal proteins; copurifies with the Dam1 complex (aka DASH complex) | Ribosomal protein 10 (rp10) of the small (40S) subunit; nearly identical to Rps1Ap and has similarity to rat S3a ribosomal protein |
| 235 | 6 | 435 | -6.141 | 0.225 | 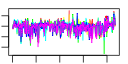 | 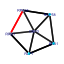 | 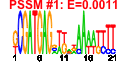 | 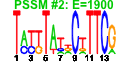 |  | YGL078C YJL148W YMR131C YNL002C YOL077C YOR310C | DBP3 RPA34 RRB1 RLP7 BRX1 NOP58 | Putative ATP-dependent RNA helicase of the DEAD-box family involved in ribosomal biogenesis | RNA polymerase I subunit A34.5 | Essential nuclear protein involved in early steps of ribosome biogenesis; physically interacts with the ribosomal protein Rpl3p | Nucleolar protein with similarity to large ribosomal subunit L7 proteins; constituent of 66S pre-ribosomal particles; plays an essential role in processing of precursors to the large ribosomal subunit RNAs | Nucleolar protein, constituent of 66S pre-ribosomal particles; depletion leads to defects in rRNA processing and a block in the assembly of large ribosomal subunits; possesses a sigma(70)-like RNA-binding motif | Protein involved in pre-rRNA processing, 18S rRNA synthesis, and snoRNA synthesis; component of the small subunit processome complex, which is required for processing of pre-18S rRNA |
| 115 | 7 | 442 | -5.694 | 0.247 |  |  | 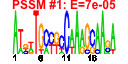 |  |  | YLR333C YML024W YMR230W YOR293W YGR027C YLR167W YLR344W | RPS25B RPS17A RPS10B RPS10A RPS25A RPS31 RPL26A | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps25Ap and has similarity to rat S25 ribosomal protein | Ribosomal protein 51 (rp51) of the small (40s) subunit; nearly identical to Rps17Bp and has similarity to rat S17 ribosomal protein | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps10Ap and has similarity to rat ribosomal protein S10 | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps10Bp and has similarity to rat ribosomal protein S10 | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps25Bp and has similarity to rat S25 ribosomal protein | Fusion protein that is cleaved to yield a ribosomal protein of the small (40S) subunit and ubiquitin; ubiquitin may facilitate assembly of the ribosomal protein into ribosomes; interacts genetically with translation factor eIF2B | Protein component of the large (60S) ribosomal subunit, nearly identical to Rpl26Bp and has similarity to E. coli L24 and rat L26 ribosomal proteins; binds to 5.8S rRNA |
| 210 | 7 | 443 | -5.212 | 0.228 |  |  |  |  |  | YMR290C YNL132W YOR310C YDR165W YGL120C YLR197W YNL248C | HAS1 KRE33 NOP58 TRM82 PRP43 SIK1 RPA49 | ATP-dependent RNA helicase; localizes to both the nuclear periphery and nucleolus; highly enriched in nuclear pore complex fractions; constituent of 66S pre-ribosomal particles | Essential protein of unknown function; heterozygous mutant shows haploinsufficiency in K1 killer toxin resistance | Protein involved in pre-rRNA processing, 18S rRNA synthesis, and snoRNA synthesis; component of the small subunit processome complex, which is required for processing of pre-18S rRNA | Subunit of a tRNA methyltransferase complex composed of Trm8p and Trm82p that catalyzes 7-methylguanosine modification of tRNA | RNA helicase in the DEAH-box family, functions in both RNA polymerase I and polymerase II transcript metabolism, involved in release of the lariat-intron from the spliceosome | Essential evolutionarily-conserved nucleolar protein component of the box C/D snoRNP complexes that direct 2'-O-methylation of pre-rRNA during its maturation; overexpression causes spindle orientation defects | RNA polymerase I subunit A49 |
| 209 | 9 | 435 | -4.838 | 0.280 | 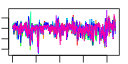 | 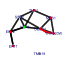 | 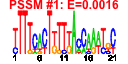 | 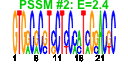 |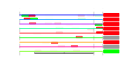  | YDR385W YGR264C YLR249W YNL209W YOR133W YHR019C YHR020W YIL078W YKL056C | EFT2 MES1 YEF3 SSB2 EFT1 DED81 THS1 TMA19 | Elongation factor 2 (EF-2), also encoded by EFT1; catalyzes ribosomal translocation during protein synthesis; contains diphthamide, the unique posttranslationally modified histidine residue specifically ADP-ribosylated by diphtheria toxin | Methionyl-tRNA synthetase, forms a complex with glutamyl-tRNA synthetase (Gus1p) and Arc1p, which increases the catalytic efficiency of both tRNA synthetases; also has a role in nuclear export of tRNAs | Translational elongation factor 3, stimulates the binding of aminoacyl-tRNA (AA-tRNA) to ribosomes by releasing EF-1 alpha from the ribosomal complex; contains two ABC cassettes; binds and hydrolyses ATP | Cytoplasmic ATPase that is a ribosome-associated molecular chaperone, functions with J-protein partner Zuo1p; may be involved in the folding of newly-synthesized polypeptide chains; member of the HSP70 family; homolog of SSB1 | Elongation factor 2 (EF-2), also encoded by EFT2; catalyzes ribosomal translocation during protein synthesis; contains diphthamide, the unique posttranslationally modified histidine residue specifically ADP-ribosylated by diphtheria toxin | Cytosolic asparaginyl-tRNA synthetase, required for protein synthesis, catalyzes the specific attachment of asparagine to its cognate tRNA | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; has similarity to proline-tRNA ligase; YHR020W is an essential gene | Threonyl-tRNA synthetase, essential cytoplasmic protein | Protein that associates with ribosomes; homolog of translationally controlled tumor protein; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and relocates to the mitochondrial outer surface upon oxidative stress |
| 58 | 9 | 447 | -4.786 | 0.261 | 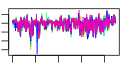 | 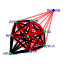 | 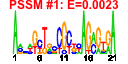 |  |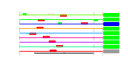  | YKR057W YNL162W YOR234C YPR132W YBL087C YGL030W YKR094C YMR143W YOR167C | RPS21A YNL162W-A RPL33B RPS23B RPL23A RPL30 RPL40B RPS16A RPS28A | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps21Bp and has similarity to rat S21 ribosomal protein | Putative protein of unknown function, identified by homology | Ribosomal protein L37 of the large (60S) ribosomal subunit, nearly identical to Rpl33Ap and has similarity to rat L35a; rpl33b null mutant exhibits normal growth while rpl33a rpl33b double null mutant is inviable | Ribosomal protein 28 (rp28) of the small (40S) ribosomal subunit, required for translational accuracy; nearly identical to Rps23Ap and similar to E. coli S12 and rat S23 ribosomal proteins; deletion of both RPS23A and RPS23B is lethal | Protein component of the large (60S) ribosomal subunit, identical to Rpl23Bp and has similarity to E. coli L14 and rat L23 ribosomal proteins | Protein component of the large (60S) ribosomal subunit, has similarity to rat L30 ribosomal protein; involved in pre-rRNA processing in the nucleolus; autoregulates splicing of its transcript | Fusion protein, identical to Rpl40Ap, that is cleaved to yield ubiquitin and a ribosomal protein of the large (60S) ribosomal subunit with similarity to rat L40; ubiquitin may facilitate assembly of the ribosomal protein into ribosomes | Protein component of the small (40S) ribosomal subunit; identical to Rps16Bp and has similarity to E. coli S9 and rat S16 ribosomal proteins | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps28Bp and has similarity to rat S28 ribosomal protein |
| 40 | 9 | 436 | -4.729 | 0.252 | 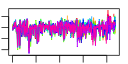 | 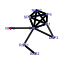 | 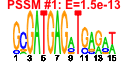 |  |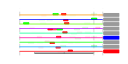  | YDR083W YDR299W YDR496C YGL078C YGL171W YHR052W YHR066W YLR276C YOR206W | RRP8 BFR2 PUF6 DBP3 ROK1 CIC1 SPT16 DBP9 NOC2 | Nucleolar protein involved in rRNA processing, pre-rRNA cleavage at site A2; also involved in telomere maintenance; mutation is synthetically lethal with a gar1 mutation | Essential protein possibly involved in secretion; multicopy suppressor of sensitivity to Brefeldin A | Pumilio-homology domain protein that binds ASH1 mRNA at PUF consensus sequences in the 3' UTR and represses its translation, resulting in proper asymmetric localization of ASH1 mRNA | Putative ATP-dependent RNA helicase of the DEAD-box family involved in ribosomal biogenesis | ATP-dependent RNA helicase of the DEAD box family; required for 18S rRNA synthesis | Essential protein that interacts with proteasome components and has a potential role in proteasome substrate specificity; also copurifies with 66S pre-ribosomal particles | Subunit of the heterodimeric FACT complex (Spt16p-Pob3p), facilitates RNA Polymerase II transcription elongation through nucleosomes by destabilizing and then reassembling nucleosome structure | ATP-dependent RNA helicase of the DEAD-box family involved in biogenesis of the 60S ribosomal subunit | Protein that forms a nucleolar complex with Mak21p that binds to 90S and 66S pre-ribosomes, as well as a nuclear complex with Noc3p that binds to 66S pre-ribosomes; both complexes mediate intranuclear transport of ribosomal precursors |
| 139 | 8 | 441 | -4.205 | 0.257 |  | 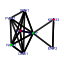 |  |  |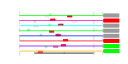  | YER006W YGR103W YGR145W YGR245C YLL008W YLR196W YNL061W YNL132W | NUG1 NOP7 ENP2 SDA1 DRS1 PWP1 NOP2 KRE33 | GTPase that associates with nuclear 60S pre-ribosomes, required for export of 60S ribosomal subunits from the nucleus | Nucleolar protein involved in rRNA processing and 60S ribosomal subunit biogenesis; constituent of several different pre-ribosomal particles; required for exit from G0 and the initiation of cell proliferation | Essential nucleolar protein of unknown function; contains WD repeats, interacts with Mpp10p and Bfr2p, and has homology to Spb1p | Highly conserved nuclear protein required for actin cytoskeleton organization and passage through Start, plays a critical role in G1 events, binds Nap1p, also involved in 60S ribosome biogenesis | Nucleolar DEAD-box protein required for ribosome assembly and function, including synthesis of 60S ribosomal subunits; constituent of 66S pre-ribosomal particles | Protein with WD-40 repeats involved in rRNA processing; associates with trans-acting ribosome biogenesis factors; similar to beta-transducin superfamily | Probable RNA m(5)C methyltransferase, essential for processing and maturation of 27S pre-rRNA and large ribosomal subunit biogenesis; localized to the nucleolus; constituent of 66S pre-ribosomal particles | Essential protein of unknown function; heterozygous mutant shows haploinsufficiency in K1 killer toxin resistance |
| 44 | 8 | 441 | -3.944 | 0.272 | 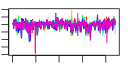 | 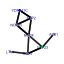 |  | 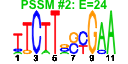 |  | YJL122W YKL143W YNL110C YOR004W YGL171W YKL172W YOR294W YPL043W | ALB1 LTV1 NOP15 UTP23 ROK1 EBP2 YDR341C NOP4 | Shuttling pre-60S factor; involved in the biogenesis of ribosomal large subunit; interacts directly with Arx1p; responsible for Tif6p recycling defects in absence of Rei1p | Component of the GSE complex, which is required for proper sorting of amino acid permease Gap1p; required for ribosomal small subunit export from nucleus; required for growth at low temperature | Constituent of 66S pre-ribosomal particles, involved in 60S ribosomal subunit biogenesis; localizes to both nucleolus and cytoplasm | Essential nucleolar protein that is a component of the SSU (small subunit) processome involved in 40S ribosomal subunit biogenesis; has homology to PINc domain protein Fcf1p, although the PINc domain of Utp23p is not required for function | ATP-dependent RNA helicase of the DEAD box family; required for 18S rRNA synthesis | Essential protein required for the maturation of 25S rRNA and 60S ribosomal subunit assembly, localizes to the nucleolus; constituent of 66S pre-ribosomal particles | Arginyl-tRNA synthetase, proposed to be cytoplasmic but the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Nucleolar protein, essential for processing and maturation of 27S pre-rRNA and large ribosomal subunit biogenesis; constituent of 66S pre-ribosomal particles; contains four RNA recognition motifs (RRMs) |
| 273 | 11 | 441 | -3.852 | 0.289 |  |  |  | 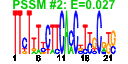 |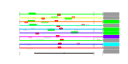  | YBL028C YER126C YER127W YHR052W YIL127C YJL122W YKL009W YKR081C YLR009W YNL113W YNL114C | NSA2 LCP5 CIC1 ALB1 MRT4 RPF2 RLP24 RPC19 | Protein of unknown function that may interact with ribosomes; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus; predicted to be involved in ribosome biogenesis | Protein constituent of 66S pre-ribosomal particles, contributes to processing of the 27S pre-rRNA | Essential protein involved in maturation of 18S rRNA; depletion leads to inhibited pre-rRNA processing and reduced polysome levels; localizes primarily to the nucleolus | Essential protein that interacts with proteasome components and has a potential role in proteasome substrate specificity; also copurifies with 66S pre-ribosomal particles | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus | Shuttling pre-60S factor; involved in the biogenesis of ribosomal large subunit; interacts directly with Arx1p; responsible for Tif6p recycling defects in absence of Rei1p | Protein involved in mRNA turnover and ribosome assembly, localizes to the nucleolus | Essential protein involved in the processing of pre-rRNA and the assembly of the 60S ribosomal subunit; interacts with ribosomal protein L11; localizes predominantly to the nucleolus; constituent of 66S pre-ribosomal particles | Essential protein with similarity to Rpl24Ap and Rpl24Bp, associated with pre-60S ribosomal subunits and required for ribosomal large subunit biogenesis | RNA polymerase subunit, common to RNA polymerases I and III |
| 224 | 12 | 457 | -3.809 | 0.308 |  |  |  | 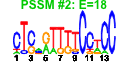 |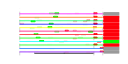  | YDL136W YER074W YOR312C YDL191W YDR447C YHR021C YIL148W YJL136C YKL156W YKR057W YOR182C YPR043W | RPL35B RPS24A RPL20B RPL35A RPS17B RPS27B RPL40A RPS21B RPS27A RPS21A RPS30B RPL43A | Protein component of the large (60S) ribosomal subunit, identical to Rpl35Ap and has similarity to rat L35 ribosomal protein | Protein component of the small (40S) ribosomal subunit; identical to Rps24Bp and has similarity to rat S24 ribosomal protein | Protein component of the large (60S) ribosomal subunit, nearly identical to Rpl20Ap and has similarity to rat L18a ribosomal protein | Protein component of the large (60S) ribosomal subunit, identical to Rpl35Bp and has similarity to rat L35 ribosomal protein | Ribosomal protein 51 (rp51) of the small (40s) subunit; nearly identical to Rps17Ap and has similarity to rat S17 ribosomal protein | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps27Ap and has similarity to rat S27 ribosomal protein | Fusion protein, identical to Rpl40Bp, that is cleaved to yield ubiquitin and a ribosomal protein of the large (60S) ribosomal subunit with similarity to rat L40; ubiquitin may facilitate assembly of the ribosomal protein into ribosomes | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps21Ap and has similarity to rat S21 ribosomal protein | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps27Bp and has similarity to rat S27 ribosomal protein | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps21Bp and has similarity to rat S21 ribosomal protein | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps30Ap and has similarity to rat S30 ribosomal protein | Protein component of the large (60S) ribosomal subunit, identical to Rpl43Bp and has similarity to rat L37a ribosomal protein; null mutation confers a dominant lethal phenotype |
| 289 | 9 | 437 | -3.546 | 0.270 |  |  |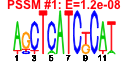  |  |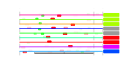  | YBL039C YDR101C YHR170W YMR290C YNL141W YOR095C YOR272W YPL086C YPL211W | URA7 ARX1 NMD3 HAS1 AAH1 RKI1 YTM1 ELP3 NIP7 | Major CTP synthase isozyme (see also URA8), catalyzes the ATP-dependent transfer of the amide nitrogen from glutamine to UTP, forming CTP, the final step in de novo biosynthesis of pyrimidines; involved in phospholipid biosynthesis | Shuttling pre-60S factor; involved in the biogenesis of ribosomal large subunit biogenesis; interacts directly with Alb1; responsible for Tif6 recycling defects in absence of Rei1; associated with the ribosomal export complex | Protein involved in nuclear export of the large ribosomal subunit; acts as a Crm1p-dependent adapter protein for export of nascent ribosomal subunits through the nuclear pore complex | ATP-dependent RNA helicase; localizes to both the nuclear periphery and nucleolus; highly enriched in nuclear pore complex fractions; constituent of 66S pre-ribosomal particles | Adenine deaminase (adenine aminohydrolase), involved in purine salvage and nitrogen catabolism | Ribose-5-phosphate ketol-isomerase, catalyzes the interconversion of ribose 5-phosphate and ribulose 5-phosphate in the pentose phosphate pathway; participates in pyridoxine biosynthesis | Constituent of 66S pre-ribosomal particles, required for maturation of the large ribosomal subunit | Subunit of Elongator complex, which is required for modification of wobble nucleosides in tRNA; exhibits histone acetyltransferase activity that is directed to histones H3 and H4; disruption confers resistance to K. lactis zymotoxin | Nucleolar protein required for 60S ribosome subunit biogenesis, constituent of 66S pre-ribosomal particles; physically interacts with Nop8p and the exosome subunit Rrp43p |
| 45 | 11 | 429 | -3.395 | 0.280 |  |  | 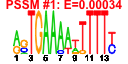 |  |  | YEL026W YHR089C YJR070C YLR175W YLR197W YLR198C YMR217W YNL248C YOR309C YPR110C YPR163C | SNU13 GAR1 LIA1 CBF5 SIK1 GUA1 RPA49 RPC40 STM1 | RNA binding protein, part of U3 snoRNP involved in rRNA processing, part of U4/U6-U5 tri-snRNP involved in mRNA splicing, similar to human 15.5K protein | Protein component of the H/ACA snoRNP pseudouridylase complex, involved in the modification and cleavage of the 18S pre-rRNA | Deoxyhypusine hydroxylase, a HEAT-repeat containing metalloenzyme that catalyses hypusine formation; binds to and is required for the modification of Hyp2p (eIF5A); complements S. pombe mmd1 mutants defective in mitochondrial positioning | Pseudouridine synthase catalytic subunit of box H/ACA small nucleolar ribonucleoprotein particles (snoRNPs), acts on both large and small rRNAs and on snRNA U2 | Essential evolutionarily-conserved nucleolar protein component of the box C/D snoRNP complexes that direct 2'-O-methylation of pre-rRNA during its maturation; overexpression causes spindle orientation defects | GMP synthase, an enzyme that catalyzes the second step in the biosynthesis of GMP from inosine 5'-phosphate (IMP); transcription is not subject to regulation by guanine but is negatively regulated by nutrient starvation | RNA polymerase I subunit A49 | RNA polymerase subunit, common to RNA polymerase I and III | Protein that binds G4 quadruplex and purine motif triplex nucleic acid; acts with Cdc13p to maintain telomere structure; interacts with ribosomes and subtelomeric Y' DNA; multicopy suppressor of tom1 and pop2 mutations |
| 267 | 9 | 443 | -3.208 | 0.293 |  |  |  |  |  | YDR120C YDL148C YDL153C YDR083W YDR496C YJR002W YNL174W YNL175C YOR206W | TRM1 NOP14 SAS10 RRP8 PUF6 MPP10 NOP13 NOC2 | tRNA methyltransferase, localizes to both the nucleus and mitochondrion to produce the modified base N2,N2-dimethylguanosine in tRNAs in both compartments | Nucleolar protein, forms a complex with Noc4p that mediates maturation and nuclear export of 40S ribosomal subunits; also present in the small subunit processome complex, which is required for processing of pre-18S rRNA | Component of the small (ribosomal) subunit (SSU) processosome required for pre-18S rRNa processing; essential nucleolar protein that, when overproduced, disrupts silencing | Nucleolar protein involved in rRNA processing, pre-rRNA cleavage at site A2; also involved in telomere maintenance; mutation is synthetically lethal with a gar1 mutation | Pumilio-homology domain protein that binds ASH1 mRNA at PUF consensus sequences in the 3' UTR and represses its translation, resulting in proper asymmetric localization of ASH1 mRNA | Component of the SSU processome, which is required for pre-18S rRNA processing, interacts with and controls the stability of Imp3p and Imp4p, essential for viability; similar to human Mpp10p | Protein of unknown function, localizes to the nucleolus and nucleoplasm; contains an RNA recognition motif (RRM) and has similarity to Nop12p, which is required for processing of pre-18S rRNA | Protein that forms a nucleolar complex with Mak21p that binds to 90S and 66S pre-ribosomes, as well as a nuclear complex with Noc3p that binds to 66S pre-ribosomes; both complexes mediate intranuclear transport of ribosomal precursors |
| 105 | 14 | 443 | -3.117 | 0.389 |  |  |  | 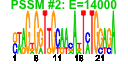 |  | YDR465C YGR245C YJL010C YJL033W YKL078W YLR009W YLR017W YML043C YNL313C YPL012W YJL050W YLR276C YLR401C YPL126W | RMT2 SDA1 NOP9 HCA4 DHR2 RLP24 MEU1 RRN11 RRP12 MTR4 DBP9 DUS3 NAN1 | Arginine methyltransferase; ribosomal protein L12 is a substrate | Highly conserved nuclear protein required for actin cytoskeleton organization and passage through Start, plays a critical role in G1 events, binds Nap1p, also involved in 60S ribosome biogenesis | Essential component of pre-40S ribosomes that is required for early cleavages of 35S pre-rRNA and hence formation of 18S rRNA; binds RNA in vitro and contains multiple pumilio-like repeats | Putative nucleolar DEAD box RNA helicase; high-copy number suppression of a U14 snoRNA processing mutant suggests an involvement in 18S rRNA synthesis | Predominantly nucleolar DEAH-box ATP-dependent RNA helicase, required for 18S rRNA synthesis | Essential protein with similarity to Rpl24Ap and Rpl24Bp, associated with pre-60S ribosomal subunits and required for ribosomal large subunit biogenesis | Methylthioadenosine phosphorylase (MTAP), catalyzes the initial step in the methionine salvage pathway; affects polyamine biosynthesis through regulation of ornithine decarboxylase (Spe1p) activity; regulates ADH2 gene expression | Protein required for rDNA transcription by RNA polymerase I, component of the core factor (CF) of rDNA transcription factor, which also contains Rrn6p and Rrn7p | Putative protein of unknown function; identified as interacting with Kar2p, Grs1p, and Tub3p in high-throughput TAP-tagging experiments; YNL313C is an essential gene | Protein required for export of the ribosomal subunits; associates with the RNA components of the pre-ribosomes; contains HEAT-repeats | Dead-box family ATP dependent helicase required for mRNA export from the nucleus; co-factor of the exosome complex, required for 3' end formation of 5.8S rRNA | ATP-dependent RNA helicase of the DEAD-box family involved in biogenesis of the 60S ribosomal subunit | Dihydrouridine synthase, member of a widespread family of conserved proteins including Smm1p, Dus1p, and Dus4p; contains a consensus oleate response element (ORE) in its promoter region | U3 snoRNP protein, component of the small (ribosomal) subunit (SSU) processosome containing U3 snoRNA; required for the biogenesis of18S rRNA |
| 277 | 14 | 438 | -3.034 | 0.303 |  |  | 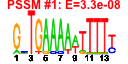 |  |  | YPR041W YBR079C YBR084W YDR091C YGR264C YKR059W YLR249W YML056C YMR217W YMR309C YNL209W YNL247W YOR243C YPL226W | TIF5 RPG1 MIS1 RLI1 MES1 TIF1 YEF3 IMD4 GUA1 NIP1 SSB2 PUS7 NEW1 | Translation initiation factor eIF-5; N-terminal domain functions as a GTPase-activating protein to mediate hydrolysis of ribosome-bound GTP; C-terminal domain is the core of ribosomal preinitiation complex formation | Subunit of the core complex of translation initiation factor 3(eIF3), essential for translation; part of a subcomplex (Prt1p-Rpg1p-Nip1p) that stimulates binding of mRNA and tRNA(i)Met to ribosomes | Mitochondrial C1-tetrahydrofolate synthase, involved in interconversion between different oxidation states of tetrahydrofolate (THF); provides activities of formyl-THF synthetase, methenyl-THF cyclohydrolase, and methylene-THF dehydrogenase | Essential iron-sulfur protein required for ribosome biogenesis and translation initiation; facilitates binding of a multifactor complex (MFC) of translation initiation factors to the small ribosomal subunit; predicted ABC family ATPase | Methionyl-tRNA synthetase, forms a complex with glutamyl-tRNA synthetase (Gus1p) and Arc1p, which increases the catalytic efficiency of both tRNA synthetases; also has a role in nuclear export of tRNAs | Translation initiation factor eIF4A, identical to Tif2p; DEA(D/H)-box RNA helicase that couples ATPase activity to RNA binding and unwinding; forms a dumbbell structure of two compact domains connected by a linker; interacts with eIF4G | Translational elongation factor 3, stimulates the binding of aminoacyl-tRNA (AA-tRNA) to ribosomes by releasing EF-1 alpha from the ribosomal complex; contains two ABC cassettes; binds and hydrolyses ATP | Inosine monophosphate dehydrogenase, catalyzes the first step of GMP biosynthesis, member of a four-gene family in S. cerevisiae, constitutively expressed | GMP synthase, an enzyme that catalyzes the second step in the biosynthesis of GMP from inosine 5'-phosphate (IMP); transcription is not subject to regulation by guanine but is negatively regulated by nutrient starvation | Subunit of the eukaryotic translation initiation factor 3 (eIF3), involved in the assembly of preinitiation complex and start codon selection | Cytoplasmic ATPase that is a ribosome-associated molecular chaperone, functions with J-protein partner Zuo1p; may be involved in the folding of newly-synthesized polypeptide chains; member of the HSP70 family; homolog of SSB1 | Essential protein of unknown function; may interact with ribosomes, based on co-purification experiments | Pseudouridine synthase, catalyzes pseudouridylation at position 35 in U2 snRNA, position 13 in cytoplasmic tRNAs, and position 35 in pre-tRNA(Tyr); Asp(256) mutation abolishes activity; conserved in archaea, some bacteria, and vertebrates | ATP binding cassette family member; Asn/Gln-rich rich region supports [NU+] prion formation, susceptibility to [PSI+] prion induction and aggregation of a fragment of the human Machado-Joseph Disease protein |
| 84 | 11 | 435 | -3.015 | 0.302 |  |  |  | 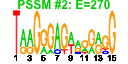 |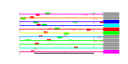  | YLR178C YOL053C YBR072W YDL204W YDR070C YER150W YFL014W YGR043C YGR088W YML128C YMR170C | TFS1 DDR2 HSP26 RTN2 FMP16 SPI1 HSP12 NQM1 CTT1 MSC1 ALD2 | Carboxypeptidase Y inhibitor, function requires acetylation by the NatB N-terminal acetyltransferase; phosphatidylethanolamine-binding protein involved in protein kinase A signaling pathway | Multistress response protein, expression is activated by a variety of xenobiotic agents and environmental or physiological stresses | Small heat shock protein (sHSP) with chaperone activity; forms hollow, sphere-shaped oligomers that suppress unfolded proteins aggregation; oligomer activation requires a heat-induced conformational change; not expressed in unstressed cells | Protein of unknown function; has similarity to mammalian reticulon proteins; member of the RTNLA (reticulon-like A) subfamily | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | GPI-anchored cell wall protein involved in weak acid resistance; basal expression requires Msn2p/Msn4p; expression is induced under conditions of stress and during the diauxic shift; similar to Sed1p | Plasma membrane localized protein that protects membranes from desiccation; induced by heat shock, oxidative stress, osmostress, stationary phase entry, glucose depletion, oleate and alcohol; regulated by the HOG and Ras-Pka pathways | Protein of unknown function; transcription is repressed by Mot1p and induced by alpha-factor and during diauxic shift; null mutant non-quiescent cells exhibit reduced reproductive capacity | Cytosolic catalase T, has a role in protection from oxidative damage by hydrogen peroxide | Protein of unknown function; mutant is defective in directing meiotic recombination events to homologous chromatids; the authentic, non-tagged protein is detected in highly purified mitochondria and is phosphorylated | Cytoplasmic aldehyde dehydrogenase, involved in ethanol oxidation and beta-alanine biosynthesis; uses NAD+ as the preferred coenzyme; expression is stress induced and glucose repressed; very similar to Ald3p |
| 36 | 11 | 441 | -2.996 | 0.328 |  |  |  |  |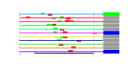  | YGR128C YJL069C YJL109C YJR041C YLL008W YLR409C YMR229C YNL182C YPL086C YPR010C YPR112C | UTP8 UTP18 UTP10 URB2 DRS1 UTP21 RRP5 IPI3 ELP3 RFA2 MRD1 | Nucleolar protein required for export of tRNAs from the nucleus; also copurifies with the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Possible U3 snoRNP protein involved in maturation of pre-18S rRNA, based on computational analysis of large-scale protein-protein interaction data | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Nucleolar protein required for normal metabolism of the rRNA primary transcript, proposed to be involved in ribosome biogenesis | Nucleolar DEAD-box protein required for ribosome assembly and function, including synthesis of 60S ribosomal subunits; constituent of 66S pre-ribosomal particles | Protein required for the synthesis of both 18S and 5.8S rRNA; C-terminal region is crucial for the formation of 18S rRNA and N-terminal region is required for the 5.8S rRNA; component of small ribosomal subunit (SSU) processosome | Essential component of the Rix1 complex (Rix1p, Ipi1p, Ipi3p) that is required for processing of ITS2 sequences from 35S pre-rRNA; highly conserved and contains WD40 motifs; Rix1 complex associates with Mdn1p in pre-60S ribosomal particles | Subunit of Elongator complex, which is required for modification of wobble nucleosides in tRNA; exhibits histone acetyltransferase activity that is directed to histones H3 and H4; disruption confers resistance to K. lactis zymotoxin | Subunit of heterotrimeric Replication Protein A (RPA), which is a highly conserved single-stranded DNA binding protein involved in DNA replication, repair, and recombination | Essential conserved protein that associates with 35S precursor rRNA and is required for its initial processing at the A(0)-A(2) cleavage sites, shows partial nucleolar localization, contains five consensus RNA-binding domains |
| 95 | 11 | 450 | -2.825 | 0.340 |  | 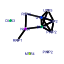 |  |  |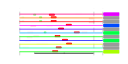  | YCR055C YCR057C YCR072C YDR398W YJL050W YLR129W YLR401C YPL217C YDR087C YML093W YPL093W | PWP2 RSA4 UTP5 MTR4 DIP2 DUS3 BMS1 RRP1 UTP14 NOG1 | Conserved 90S pre-ribosomal component essential for proper endonucleolytic cleavage of the 35 S rRNA precursor at A0, A1, and A2 sites; contains eight WD-repeats; PWP2 deletion leads to defects in cell cycle and bud morphogenesis | WD-repeat protein involved in ribosome biogenesis; may interact with ribosomes; required for maturation and efficient intra-nuclear transport or pre-60S ribosomal subunits, localizes to the nucleolus | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Dead-box family ATP dependent helicase required for mRNA export from the nucleus; co-factor of the exosome complex, required for 3' end formation of 5.8S rRNA | Nucleolar protein, specifically associated with the U3 snoRNA, part of the large ribonucleoprotein complex known as the small subunit (SSU) processome, required for 18S rRNA biogenesis, part of the active pre-rRNA processing complex | Dihydrouridine synthase, member of a widespread family of conserved proteins including Smm1p, Dus1p, and Dus4p; contains a consensus oleate response element (ORE) in its promoter region | Essential conserved nucleolar GTP-binding protein required for synthesis of 40S ribosomal subunits and for processing of the 35S pre-rRNA at sites A0, A1, and A2; interacts with Rcl1p, has similarity to Tsr1p | Essential evolutionarily conserved nucleolar protein necessary for biogenesis of 60S ribosomal subunits and processing of pre-rRNAs to mature rRNAs, associated with several distinct 66S pre-ribosomal particles | Putative GTPase that associates with free 60S ribosomal subunits in the nucleolus and is required for 60S ribosomal subunit biogenesis; constituent of 66S pre-ribosomal particles; member of the ODN family of nucleolar G-proteins |
| 10 | 11 | 432 | -2.791 | 0.320 |  |  |  |  |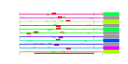  | YLL034C YLR186W YML093W YNL062C YNL075W YNL113W YOL080C YOR340C YPL043W YPL126W YPL266W | RIX7 EMG1 UTP14 GCD10 IMP4 RPC19 REX4 RPA43 NOP4 NAN1 CDH1 | Putative ATPase of the AAA family, required for export of pre-ribosomal large subunits from the nucleus; distributed between the nucleolus, nucleoplasm, and nuclear periphery depending on growth conditions | Protein required for the maturation of the 18S rRNA and for 40S ribosome production; associated with spindle/microtubules; nuclear localization depends on physical interaction with Nop14p; may bind snoRNAs | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Subunit of tRNA (1-methyladenosine) methyltransferase with Gcd14p, required for the modification of the adenine at position 58 in tRNAs, especially tRNAi-Met; first identified as a negative regulator of GCN4 expression | Component of the SSU processome, which is required for pre-18S rRNA processing; interacts with Mpp10p; member of a superfamily of proteins that contain a sigma(70)-like motif and associate with RNAs | RNA polymerase subunit, common to RNA polymerases I and III | Putative RNA exonuclease possibly involved in pre-rRNA processing and ribosome assembly | RNA polymerase I subunit A43 | Nucleolar protein, essential for processing and maturation of 27S pre-rRNA and large ribosomal subunit biogenesis; constituent of 66S pre-ribosomal particles; contains four RNA recognition motifs (RRMs) | U3 snoRNP protein, component of the small (ribosomal) subunit (SSU) processosome containing U3 snoRNA; required for the biogenesis of18S rRNA | Cell-cycle regulated activator of the anaphase-promoting complex/cyclosome (APC/C), which directs ubiquitination of cyclins resulting in mitotic exit; targets the APC/C to specific substrates including CDC20, ASE1, CIN8 and FIN1 |
| 64 | 11 | 452 | -2.746 | 0.396 |  |  | 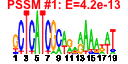 | 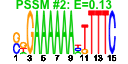 |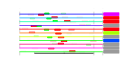  | YAL036C YBR142W YJL148W YMR310C YNL151C YOR145C YPL093W YPL094C YNL062C YOR340C YPR190C | RBG1 MAK5 RPA34 RPC31 PAN1 NOG1 SEC62 GCD10 RPA43 RPC82 | Member of the DRG family of GTP-binding proteins; interacts with translating ribosomes and with Tma46p | Essential nucleolar protein, putative DEAD-box RNA helicase required for maintenance of M1 dsRNA virus; involved in biogenesis of large (60S) ribosomal subunits | RNA polymerase I subunit A34.5 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; YMR310C is not an essential gene | RNA polymerase III subunit C31; contains HMG-like C-terminal domain | Part of actin cytoskeleton-regulatory complex Pan1p-Sla1p-End3p, associates with actin patches on the cell cortex; promotes protein-protein interactions essential for endocytosis; previously thought to be a subunit of poly(A) ribonuclease | Putative GTPase that associates with free 60S ribosomal subunits in the nucleolus and is required for 60S ribosomal subunit biogenesis; constituent of 66S pre-ribosomal particles; member of the ODN family of nucleolar G-proteins | Essential subunit of Sec63 complex (Sec63p, Sec62p, Sec66p and Sec72p); with Sec61 complex, Kar2p/BiP and Lhs1p forms a channel competent for SRP-dependent and post-translational SRP-independent protein targeting and import into the ER | Subunit of tRNA (1-methyladenosine) methyltransferase with Gcd14p, required for the modification of the adenine at position 58 in tRNAs, especially tRNAi-Met; first identified as a negative regulator of GCN4 expression | RNA polymerase I subunit A43 | RNA polymerase III subunit C82 |
| 85 | 11 | 435 | -2.717 | 0.345 |  |  |  | 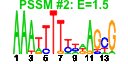 |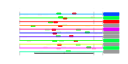  | YDL031W YDR087C YDR161W YER006W YGL099W YGL111W YIR026C YLR435W YOL022C YDL201W YPL266W | DBP10 RRP1 NUG1 LSG1 NSA1 YVH1 TSR2 TSR4 TRM8 CDH1 | Putative ATP-dependent RNA helicase of the DEAD-box protein family, constituent of 66S pre-ribosomal particles; essential protein involved in ribosome biogenesis | Essential evolutionarily conserved nucleolar protein necessary for biogenesis of 60S ribosomal subunits and processing of pre-rRNAs to mature rRNAs, associated with several distinct 66S pre-ribosomal particles | Putative protein of unknown function; non-essential gene; proposed function in rRNA and ribosome biosynthesis based on transcriptional co-regulation | GTPase that associates with nuclear 60S pre-ribosomes, required for export of 60S ribosomal subunits from the nucleus | Putative GTPase involved in 60S ribosomal subunit biogenesis; required for the release of Nmd3p from 60S subunits in the cytoplasm | Constituent of 66S pre-ribosomal particles, involved in 60S ribosomal subunit biogenesis | Protein phosphatase involved in vegetative growth at low temperatures, sporulation, and glycogen accumulation; transcription induced by low temperature and nitrogen starvation; member of the dual-specificity family of protein phosphatases | Protein with a potential role in pre-rRNA processing | Cytoplasmic protein of unknown function; essential gene in S288C, and non-essential with reduced growth rate in CEN.PK2; Null mutant accumulates 20S pre-rRNA | Subunit of a tRNA methyltransferase complex composed of Trm8p and Trm82p that catalyzes 7-methylguanosine modification of tRNA | Cell-cycle regulated activator of the anaphase-promoting complex/cyclosome (APC/C), which directs ubiquitination of cyclins resulting in mitotic exit; targets the APC/C to specific substrates including CDC20, ASE1, CIN8 and FIN1 |
| 202 | 11 | 449 | -2.643 | 0.351 |  |  | 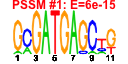 | 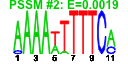 |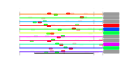  | YBL039C YDL148C YHR170W YAL059W YBR034C YDL060W YGR159C YHR148W YHR196W YHR197W YNL112W | URA7 NOP14 NMD3 ECM1 HMT1 TSR1 NSR1 IMP3 UTP9 RIX1 DBP2 | Major CTP synthase isozyme (see also URA8), catalyzes the ATP-dependent transfer of the amide nitrogen from glutamine to UTP, forming CTP, the final step in de novo biosynthesis of pyrimidines; involved in phospholipid biosynthesis | Nucleolar protein, forms a complex with Noc4p that mediates maturation and nuclear export of 40S ribosomal subunits; also present in the small subunit processome complex, which is required for processing of pre-18S rRNA | Protein involved in nuclear export of the large ribosomal subunit; acts as a Crm1p-dependent adapter protein for export of nascent ribosomal subunits through the nuclear pore complex | Protein of unknown function, localized in the nucleoplasm and the nucleolus, genetically interacts with MTR2 in 60S ribosomal protein subunit export | Nuclear SAM-dependent mono- and asymmetric arginine dimethylating methyltransferase that modifies hnRNPs, including Npl3p and Hrp1p, thus facilitating nuclear export of these proteins; required for viability of npl3 mutants | Protein required for processing of 20S pre-rRNA in the cytoplasm, associates with pre-40S ribosomal particles | Nucleolar protein that binds nuclear localization sequences, required for pre-rRNA processing and ribosome biogenesis | Component of the SSU processome, which is required for pre-18S rRNA processing, essential protein that interacts with Mpp10p and mediates interactions of Imp4p and Mpp10p with U3 snoRNA | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Essential component of the Rix1 complex (Rix1p, Ipi1p, Ipi3p) that is required for processing of ITS2 sequences from 35S pre-rRNA; Rix1 complex associates with Mdn1p in pre-60S ribosomal particles | Essential ATP-dependent RNA helicase of the DEAD-box protein family, involved in nonsense-mediated mRNA decay and rRNA processing |
| 137 | 13 | 449 | -2.639 | 0.364 | 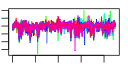 | 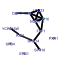 |  | 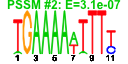 |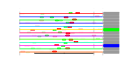  | YGR280C YKR024C YMR128W YNL308C YPR144C YCR016W YFL002C YGR272C YHR066W YIL019W YKL082C YOR004W YPR112C | PXR1 DBP7 ECM16 KRI1 NOC4 SPB4 EFG1 SPT16 FAF1 RRP14 UTP23 MRD1 | Essential protein involved in rRNA and snoRNA maturation; competes with TLC1 RNA for binding to Est2p, suggesting a role in negative regulation of telomerase; human homolog inhibits telomerase; contains a G-patch RNA interacting domain | Putative ATP-dependent RNA helicase of the DEAD-box family involved in ribosomal biogenesis | Essential DEAH-box ATP-dependent RNA helicase specific to the U3 snoRNP, predominantly nucleolar in distribution, required for 18S rRNA synthesis | Essential nucleolar protein required for 40S ribosome biogenesis; physically and functionally interacts with Krr1p | Nucleolar protein, forms a complex with Nop14p that mediates maturation and nuclear export of 40S ribosomal subunits | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus and nucleus; YCR016W is not an essential gene | Putative ATP-dependent RNA helicase, nucleolar protein required for synthesis of 60S ribosomal subunits at a late step in the pathway; sediments with 66S pre-ribosomes in sucrose gradients | Protein of unknown function; null mutant is sensitive to hydroxyurea and is delayed in recovering from alpha-factor arrest; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus | Subunit of the heterodimeric FACT complex (Spt16p-Pob3p), facilitates RNA Polymerase II transcription elongation through nucleosomes by destabilizing and then reassembling nucleosome structure | Protein required for pre-rRNA processing and 40S ribosomal subunit assembly | Essential protein, constituent of 66S pre-ribosomal particles; interacts with proteins involved in ribosomal biogenesis and cell polarity; member of the SURF-6 family | Essential nucleolar protein that is a component of the SSU (small subunit) processome involved in 40S ribosomal subunit biogenesis; has homology to PINc domain protein Fcf1p, although the PINc domain of Utp23p is not required for function | Essential conserved protein that associates with 35S precursor rRNA and is required for its initial processing at the A(0)-A(2) cleavage sites, shows partial nucleolar localization, contains five consensus RNA-binding domains |
| 136 | 12 | 445 | -2.637 | 0.324 |  |  | 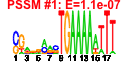 |  |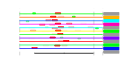  | YDR165W YGL120C YGR123C YMR093W YDR398W YGL099W YGR128C YJR070C YLR175W YLR186W YLR449W YOR309C | TRM82 PRP43 PPT1 UTP15 UTP5 LSG1 UTP8 LIA1 CBF5 EMG1 FPR4 | Subunit of a tRNA methyltransferase complex composed of Trm8p and Trm82p that catalyzes 7-methylguanosine modification of tRNA | RNA helicase in the DEAH-box family, functions in both RNA polymerase I and polymerase II transcript metabolism, involved in release of the lariat-intron from the spliceosome | Protein serine/threonine phosphatase with similarity to human phosphatase PP5; present in both the nucleus and cytoplasm; expressed during logarithmic growth; computational analyses suggest roles in phosphate metabolism and rRNA processing | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Putative GTPase involved in 60S ribosomal subunit biogenesis; required for the release of Nmd3p from 60S subunits in the cytoplasm | Nucleolar protein required for export of tRNAs from the nucleus; also copurifies with the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Deoxyhypusine hydroxylase, a HEAT-repeat containing metalloenzyme that catalyses hypusine formation; binds to and is required for the modification of Hyp2p (eIF5A); complements S. pombe mmd1 mutants defective in mitochondrial positioning | Pseudouridine synthase catalytic subunit of box H/ACA small nucleolar ribonucleoprotein particles (snoRNPs), acts on both large and small rRNAs and on snRNA U2 | Protein required for the maturation of the 18S rRNA and for 40S ribosome production; associated with spindle/microtubules; nuclear localization depends on physical interaction with Nop14p; may bind snoRNAs | Peptidyl-prolyl cis-trans isomerase (PPIase) (proline isomerase) localized to the nucleus; catalyzes isomerization of proline residues in histones H3 and H4, which affects lysine methylation of those histones |
| 53 | 31 | 446 | -2.636 | 0.535 | 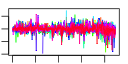 |  | 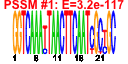 | 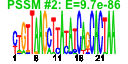 | 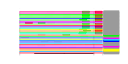 | YBL029W YCL038C YDR406W YEL049W YER037W YFL020C YGR144W YGR294W YHL046C YIL176C YIR041W YJL223C YKR046C YLL025W YLL064C YLR037C YLR053C YLR346C YLR350W YNL331C YOL161C YPL278C YCR104W YDR262W YGL260W YGL261C YJR155W YKL224C YMR325W YNR076W YPL282C | ATG22 PDR15 PAU2 PHM8 PAU5 THI4 PAU12 PAU13 PAU14 PAU15 PAU1 PET10 PAU17 PAU18 DAN2 ORM2 AAD14 PAU20 PAU3 PAU11 AAD10 PAU16 PAU19 PAU6 PAU22 | Non-essential protein of unknown function | Protein required for the breakdown of autophagic vesicles in the vacuole during autophagy, putative integral membrane protein that localizes to vacuolar membranes and punctate structures attached to the vacuole | Plasma membrane ATP binding cassette (ABC) transporter, multidrug transporter and general stress response factor implicated in cellular detoxification; regulated by Pdr1p, Pdr3p and Pdr8p; promoter contains a PDR responsive element | Part of 23-member seripauperin multigene family encoded mainly in subtelomeric regions, active during alcoholic fermentation, regulated by anaerobiosis, negatively regulated by oxygen, repressed by heme | Protein of unknown function, expression is induced by low phosphate levels and by inactivation of Pho85p | Thiazole synthase, catalyzes formation of the thiazole moiety of thiamin pyrophosphate; required for thiamine biosynthesis and for mitochondrial genome stability | hypothetical protein | Putative protein of unknown function; not an essential gene | Protein of unknown function that co-purifies with lipid particles; expression pattern suggests a role in respiratory growth; computational analysis of large-scale protein-protein interaction data suggests a role in ATP/ADP exchange | Putative protein of unknown function; YLL025W is not an essential gene | Cell wall mannoprotein with similarity to Tir1p, Tir2p, Tir3p, and Tir4p; expressed under anaerobic conditions, completely repressed during aerobic growth | Putative protein of unknown function | Putative protein of unknown function found in mitochondria; expression is regulated by transcription factors involved in pleiotropic drug resistance, Pdr1p and Yrr1p; YLR346C is not an essential gene | Evolutionarily conserved protein with similarity to Orm1p, required for resistance to agents that induce the unfolded protein response; human ortholog is located in the endoplasmic reticulum | Putative aryl-alcohol dehydrogenase with similarity to P. chrysosporium aryl-alcohol dehydrogenase; mutational analysis has not yet revealed a physiological role | Putative protein of unknown function; gene expression regulated by copper levels | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole and is induced in response to the DNA-damaging agent MMS; gene expression increases in response to Zymoliase treatment | Putative protein of unknown function; transcription is significantly increased in a NAP1 deletion background; deletion mutant has increased accumulation of nickel and selenium | Putative protein of unknown function; mRNA expression appears to be regulated by SUT1 and UPC2 |
| 97 | 13 | 437 | -2.584 | 0.319 | 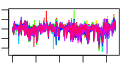 |  |  | 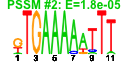 |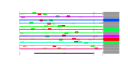  | YBR034C YGR159C YGR160W YHR169W YHR196W YIR012W YNL022C YNL112W YNR053C YOL010W YCR072C YML080W YNL182C | HMT1 NSR1 DBP8 UTP9 SQT1 DBP2 NOG2 RCL1 RSA4 DUS1 IPI3 | Nuclear SAM-dependent mono- and asymmetric arginine dimethylating methyltransferase that modifies hnRNPs, including Npl3p and Hrp1p, thus facilitating nuclear export of these proteins; required for viability of npl3 mutants | Nucleolar protein that binds nuclear localization sequences, required for pre-rRNA processing and ribosome biogenesis | Putative ATP-dependent RNA helicase of the DEAD-box family involved in biogenesis of the 40S ribosomal subunit | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Essential protein involved in a late step of 60S ribosomal subunit assembly or modification; contains multiple WD repeats; interacts with Qsr1p in a two-hybrid assay | Putative protein of unknown function with seven beta-strand methyltransferase motif similar to NOP2/YNL061W; green fluorescent protein (GFP)-fusion protein localizes to a single spot in the nucleus; YNL022C is not an essential gene | Essential ATP-dependent RNA helicase of the DEAD-box protein family, involved in nonsense-mediated mRNA decay and rRNA processing | Putative GTPase that associates with pre-60S ribosomal subunits in the nucleolus and is required for their nuclear export and maturation | RNA terminal phosphate cyclase-like protein involved in rRNA processing at sites A0, A1, and A2; does not possess detectable RNA cyclase activity | WD-repeat protein involved in ribosome biogenesis; may interact with ribosomes; required for maturation and efficient intra-nuclear transport or pre-60S ribosomal subunits, localizes to the nucleolus | Dihydrouridine synthase, member of a widespread family of conserved proteins including Smm1p, Dus3p, and Dus4p; modifies pre-tRNA(Phe) at U17 | Essential component of the Rix1 complex (Rix1p, Ipi1p, Ipi3p) that is required for processing of ITS2 sequences from 35S pre-rRNA; highly conserved and contains WD40 motifs; Rix1 complex associates with Mdn1p in pre-60S ribosomal particles |
| 153 | 12 | 445 | -2.547 | 0.358 | 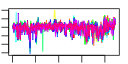 | 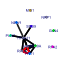 | 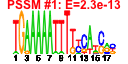 |  |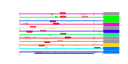  | YBR079C YBR084W YGR162W YMR309C YOR168W YOR243C YOR361C YPL081W YPL226W YCL037C YDL167C YPR010C | RPG1 MIS1 TIF4631 NIP1 GLN4 PUS7 PRT1 RPS9A NEW1 SRO9 NRP1 RFA2 | Subunit of the core complex of translation initiation factor 3(eIF3), essential for translation; part of a subcomplex (Prt1p-Rpg1p-Nip1p) that stimulates binding of mRNA and tRNA(i)Met to ribosomes | Mitochondrial C1-tetrahydrofolate synthase, involved in interconversion between different oxidation states of tetrahydrofolate (THF); provides activities of formyl-THF synthetase, methenyl-THF cyclohydrolase, and methylene-THF dehydrogenase | Translation initiation factor eIF4G, subunit of the mRNA cap-binding protein complex (eIF4F) that also contains eIF4E (Cdc33p); associates with the poly(A)-binding protein Pab1p, also interacts with eIF4A (Tif1p); homologous to Tif4632p | Subunit of the eukaryotic translation initiation factor 3 (eIF3), involved in the assembly of preinitiation complex and start codon selection | Glutamine tRNA synthetase, monomeric class I tRNA synthetase that catalyzes the specific glutaminylation of tRNA(Glu); N-terminal domain proposed to be involved in enzyme-tRNA interactions | Pseudouridine synthase, catalyzes pseudouridylation at position 35 in U2 snRNA, position 13 in cytoplasmic tRNAs, and position 35 in pre-tRNA(Tyr); Asp(256) mutation abolishes activity; conserved in archaea, some bacteria, and vertebrates | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps9Bp and has similarity to E. coli S4 and rat S9 ribosomal proteins | ATP binding cassette family member; Asn/Gln-rich rich region supports [NU+] prion formation, susceptibility to [PSI+] prion induction and aggregation of a fragment of the human Machado-Joseph Disease protein | Cytoplasmic RNA-binding protein that associates with translating ribosomes; involved in heme regulation of Hap1p as a component of the HMC complex, also involved in the organization of actin filaments; contains a La motif | Protein of unknown function, rich in asparagine residues | Subunit of heterotrimeric Replication Protein A (RPA), which is a highly conserved single-stranded DNA binding protein involved in DNA replication, repair, and recombination |
| 14 | 12 | 438 | -2.485 | 0.330 | 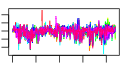 | 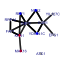 |  |  |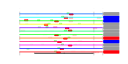  | YAL025C YCL059C YDR184C YER126C YGL029W YGR272C YHR088W YIL019W YIL127C YKL082C YKL172W YOR294W | MAK16 KRR1 ATC1 NSA2 CGR1 EFG1 RPF1 FAF1 RRP14 EBP2 YDR341C | Essential nuclear protein, constituent of 66S pre-ribosomal particles; required for maturation of 25S and 5.8S rRNAs; required for maintenance of M1 satellite double-stranded RNA of the L-A virus | Essential nucleolar protein required for the synthesis of 18S rRNA and for the assembly of 40S ribosomal subunit | Nuclear protein, possibly involved in regulation of cation stress responses and/or in the establishment of bipolar budding pattern | Protein constituent of 66S pre-ribosomal particles, contributes to processing of the 27S pre-rRNA | Protein involved in nucleolar integrity and processing of the pre-rRNA for the 60S ribosome subunit; transcript is induced in response to cytotoxic stress but not genotoxic stress | Protein of unknown function; null mutant is sensitive to hydroxyurea and is delayed in recovering from alpha-factor arrest; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus | Nucleolar protein involved in the assembly of the large ribosomal subunit; constituent of 66S pre-ribosomal particles; contains a sigma(70)-like motif, which is thought to bind RNA | Protein required for pre-rRNA processing and 40S ribosomal subunit assembly | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus | Essential protein, constituent of 66S pre-ribosomal particles; interacts with proteins involved in ribosomal biogenesis and cell polarity; member of the SURF-6 family | Essential protein required for the maturation of 25S rRNA and 60S ribosomal subunit assembly, localizes to the nucleolus; constituent of 66S pre-ribosomal particles | Arginyl-tRNA synthetase, proposed to be cytoplasmic but the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
| 174 | 10 | 454 | -2.458 | 0.383 | 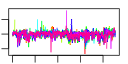 |  |  | 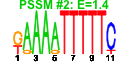 |  | YAL059W YBL054W YBR267W YDL060W YER127W YAL025C YDR184C YDR299W YDR465C YNL075W | ECM1 REI1 TSR1 LCP5 MAK16 ATC1 BFR2 RMT2 IMP4 | Protein of unknown function, localized in the nucleoplasm and the nucleolus, genetically interacts with MTR2 in 60S ribosomal protein subunit export | Protein of unknown function involved in rRNA and ribosome biosynthesis | Cytoplasmic pre-60S factor; required for the correct recycling of shuttling factors Alb1, Arx1 and Tif6 at the end of the ribosomal large subunit biogenesis; involved in bud growth in the mitotic signaling network | Protein required for processing of 20S pre-rRNA in the cytoplasm, associates with pre-40S ribosomal particles | Essential protein involved in maturation of 18S rRNA; depletion leads to inhibited pre-rRNA processing and reduced polysome levels; localizes primarily to the nucleolus | Essential nuclear protein, constituent of 66S pre-ribosomal particles; required for maturation of 25S and 5.8S rRNAs; required for maintenance of M1 satellite double-stranded RNA of the L-A virus | Nuclear protein, possibly involved in regulation of cation stress responses and/or in the establishment of bipolar budding pattern | Essential protein possibly involved in secretion; multicopy suppressor of sensitivity to Brefeldin A | Arginine methyltransferase; ribosomal protein L12 is a substrate | Component of the SSU processome, which is required for pre-18S rRNA processing; interacts with Mpp10p; member of a superfamily of proteins that contain a sigma(70)-like motif and associate with RNAs |
| 190 | 15 | 457 | -2.428 | 0.391 | 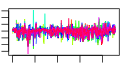 |  | 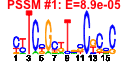 | 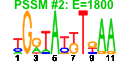 |  | YHR021C YIL148W YKL156W YLR406C YNL302C YOL127W YOR182C YDL130W YDL184C YIL069C YJL189W YJL191W YMR230W YOL039W YPR044C | RPS27B RPL40A RPS27A RPL31B RPS19B RPL25 RPS30B STF1 RPL41A RPS24B RPL39 RPS14B RPS10B RPP2A | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps27Ap and has similarity to rat S27 ribosomal protein | Fusion protein, identical to Rpl40Bp, that is cleaved to yield ubiquitin and a ribosomal protein of the large (60S) ribosomal subunit with similarity to rat L40; ubiquitin may facilitate assembly of the ribosomal protein into ribosomes | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps27Bp and has similarity to rat S27 ribosomal protein | Protein component of the large (60S) ribosomal subunit, nearly identical to Rpl31Ap and has similarity to rat L31 ribosomal protein; associates with the karyopherin Sxm1p | Protein component of the small (40S) ribosomal subunit, required for assembly and maturation of pre-40 S particles; mutations in human RPS19 are associated with Diamond Blackfan anemia; nearly identical to Rps19Ap | Primary rRNA-binding ribosomal protein component of the large (60S) ribosomal subunit, has similarity to E. coli L23 and rat L23a ribosomal proteins; binds to 26S rRNA via a conserved C-terminal motif | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps30Ap and has similarity to rat S30 ribosomal protein | Protein involved in regulation of the mitochondrial F1F0-ATP synthase; Stf1p and Stf2p act as stabilizing factors that enhance inhibitory action of the Inh1p protein | Ribosomal protein L47 of the large (60S) ribosomal subunit, identical to Rpl41Bp and has similarity to rat L41 ribosomal protein; comprised of only 25 amino acids; rpl41a rpl41b double null mutant is viable | Protein component of the small (40S) ribosomal subunit; identical to Rps24Ap and has similarity to rat S24 ribosomal protein | Protein component of the large (60S) ribosomal subunit, has similarity to rat L39 ribosomal protein; required for ribosome biogenesis; exhibits genetic interactions with SIS1 and PAB1 | Ribosomal protein 59 of the small subunit, required for ribosome assembly and 20S pre-rRNA processing; mutations confer cryptopleurine resistance; nearly identical to Rps14Ap and similar to E. coli S11 and rat S14 ribosomal proteins | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps10Ap and has similarity to rat ribosomal protein S10 | Ribosomal protein P2 alpha, a component of the ribosomal stalk, which is involved in the interaction between translational elongation factors and the ribosome; regulates the accumulation of P1 (Rpp1Ap and Rpp1Bp) in the cytoplasm |
| 192 | 14 | 449 | -2.422 | 0.344 |  |  |  | 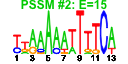 |  | YDL153C YDR312W YJR002W YMR239C YBR247C YCL059C YDR361C YHR088W YIL096C YKL099C YKL143W YLR003C YLR051C YNR003C | SAS10 SSF2 MPP10 RNT1 ENP1 KRR1 BCP1 RPF1 UTP11 LTV1 FCF2 RPC34 | Component of the small (ribosomal) subunit (SSU) processosome required for pre-18S rRNa processing; essential nucleolar protein that, when overproduced, disrupts silencing | Protein required for ribosomal large subunit maturation, functionally redundant with Ssf1p; member of the Brix family | Component of the SSU processome, which is required for pre-18S rRNA processing, interacts with and controls the stability of Imp3p and Imp4p, essential for viability; similar to human Mpp10p | RNAase III; involved in rDNA transcription and rRNA processing; also cleaves a stem-loop structure at the 3' end of U2 snRNA to ensure formation of the correct U2 3' end | Protein associated with U3 and U14 snoRNAs, required for pre-rRNA processing and 40S ribosomal subunit synthesis; localized in the nucleus and concentrated in the nucleolus | Essential nucleolar protein required for the synthesis of 18S rRNA and for the assembly of 40S ribosomal subunit | Essential protein involved in nuclear export of Mss4p, which is a lipid kinase that generates phosphatidylinositol 4,5-biphosphate and plays a role in actin cytoskeleton organization and vesicular transport | Nucleolar protein involved in the assembly of the large ribosomal subunit; constituent of 66S pre-ribosomal particles; contains a sigma(70)-like motif, which is thought to bind RNA | Putative protein of unknown function; associates with precursors of the 60S ribosomal subunit | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Component of the GSE complex, which is required for proper sorting of amino acid permease Gap1p; required for ribosomal small subunit export from nucleus; required for growth at low temperature | Putative protein of unknown function that may participate in DNA replication; green fluorescent protein (GFP)-fusion protein is localized to the nucleus; YLR003C is not an essential gene | Essential nucleolar protein involved in the early steps of 35S rRNA processing; interacts with Faf1p; member of a transcriptionally co-regulated set of genes called the RRB regulon | RNA polymerase III subunit C34; interacts with TFIIIB70 and is a key determinant in pol III recruitment by the preinitiation complex |
| 124 | 16 | 450 | -2.402 | 0.402 | 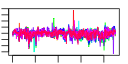 |  | 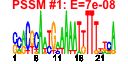 | 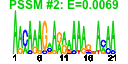 |  | YBL024W YCL037C YDR341C YDR395W YIL078W YJR143C YLR083C YLR355C YML022W YML056C YOL097C YPL263C YCL036W YDR023W YPR110C YPR163C | NCL1 SRO9 SXM1 THS1 PMT4 EMP70 ILV5 YLR118C IMD4 WRS1 KEL3 GFD2 SES1 RPC40 STM1 | S-adenosyl-L-methionine-dependent tRNA: m5C-methyltransferase, methylates cytosine to m5C at several positions in tRNAs and intron-containing pre-tRNAs; similar to Nop2p and human proliferation associated nucleolar protein p120 | Cytoplasmic RNA-binding protein that associates with translating ribosomes; involved in heme regulation of Hap1p as a component of the HMC complex, also involved in the organization of actin filaments; contains a La motif | Arginyl-tRNA synthetase, proposed to be cytoplasmic but the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Nuclear transport factor (karyopherin) involved in protein transport between the cytoplasm and nucleoplasm; similar to Nmd5p, Cse1p, Lph2p, and the human cellular apoptosis susceptibility protein, CAS1 | Threonyl-tRNA synthetase, essential cytoplasmic protein | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; appears to form homodimers in vivo and does not complex with other Pmt proteins; target for new antifungals | Protein whose 24kDa cleavage product is found in endosome-enriched membrane fractions, predicted to be a transmembrane protein | Acetohydroxyacid reductoisomerase, mitochondrial protein involved in branched-chain amino acid biosynthesis, also required for maintenance of wild-type mitochondrial DNA and found in mitochondrial nucleoids | Acyl-protein thioesterase responsible for depalmitoylation of Gpa1p; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and nucleus and is induced in response to the DNA-damaging agent MMS | Inosine monophosphate dehydrogenase, catalyzes the first step of GMP biosynthesis, member of a four-gene family in S. cerevisiae, constitutively expressed | Cytoplasmic tryptophanyl-tRNA synthetase, aminoacylates tryptophanyl-tRNA | Cytoplasmic protein of unknown function | Protein of unknown function, identified as a high-copy suppressor of a dbp5 mutation | Cytosolic seryl-tRNA synthetase, class II aminoacyl-tRNA synthetase that aminoacylates tRNA(Ser), displays tRNA-dependent amino acid recognition which enhances discrimination of the serine substrate, interacts with peroxin Pex21p | RNA polymerase subunit, common to RNA polymerase I and III | Protein that binds G4 quadruplex and purine motif triplex nucleic acid; acts with Cdc13p to maintain telomere structure; interacts with ribosomes and subtelomeric Y' DNA; multicopy suppressor of tom1 and pop2 mutations |
| 111 | 11 | 441 | -2.393 | 0.338 | 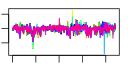 |  |  | 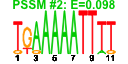 |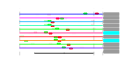  | YBR247C YCL054W YCLX02C YCR016W YMR049C YCR055C YCR057C YDR324C YLL011W YLR129W YMR229C | ENP1 SPB1 ERB1 PWP2 UTP4 SOF1 DIP2 RRP5 | Protein associated with U3 and U14 snoRNAs, required for pre-rRNA processing and 40S ribosomal subunit synthesis; localized in the nucleus and concentrated in the nucleolus | AdoMet-dependent methyltransferase involved in rRNA processing and 60S ribosomal subunit maturation; methylates G2922 in the tRNA docking site of the large subunit rRNA and in the absence of snR52, U2921; suppressor of PAB1 mutants | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus and nucleus; YCR016W is not an essential gene | Protein required for maturation of the 25S and 5.8S ribosomal RNAs; constituent of 66S pre-ribosomal particles; homologous to mammalian Bop1 | Conserved 90S pre-ribosomal component essential for proper endonucleolytic cleavage of the 35 S rRNA precursor at A0, A1, and A2 sites; contains eight WD-repeats; PWP2 deletion leads to defects in cell cycle and bud morphogenesis | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Essential protein required for biogenesis of 40S (small) ribosomal subunit; has similarity to the beta subunit of trimeric G-proteins and the splicing factor Prp4p | Nucleolar protein, specifically associated with the U3 snoRNA, part of the large ribonucleoprotein complex known as the small subunit (SSU) processome, required for 18S rRNA biogenesis, part of the active pre-rRNA processing complex | Protein required for the synthesis of both 18S and 5.8S rRNA; C-terminal region is crucial for the formation of 18S rRNA and N-terminal region is required for the 5.8S rRNA; component of small ribosomal subunit (SSU) processosome |
| 123 | 12 | 449 | -2.388 | 0.335 |  |  |  | 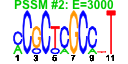 |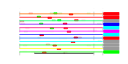  | YGL169W YHR148W YHR197W YJL208C YHR169W YIR012W YJL069C YNL022C YNR053C YOL010W YOL080C YOR145C | SUA5 IMP3 RIX1 NUC1 DBP8 SQT1 UTP18 NOG2 RCL1 REX4 PAN1 | Protein required for respiratory growth; null mutation suppresses the Cyc1p translation defect caused by the presence of an aberrant ATG codon upstream of the correct start | Component of the SSU processome, which is required for pre-18S rRNA processing, essential protein that interacts with Mpp10p and mediates interactions of Imp4p and Mpp10p with U3 snoRNA | Essential component of the Rix1 complex (Rix1p, Ipi1p, Ipi3p) that is required for processing of ITS2 sequences from 35S pre-rRNA; Rix1 complex associates with Mdn1p in pre-60S ribosomal particles | Major mitochondrial nuclease, has RNAse and DNA endo- and exonucleolytic activities; has roles in mitochondrial recombination, apoptosis and maintenance of polyploidy | Putative ATP-dependent RNA helicase of the DEAD-box family involved in biogenesis of the 40S ribosomal subunit | Essential protein involved in a late step of 60S ribosomal subunit assembly or modification; contains multiple WD repeats; interacts with Qsr1p in a two-hybrid assay | Possible U3 snoRNP protein involved in maturation of pre-18S rRNA, based on computational analysis of large-scale protein-protein interaction data | Putative protein of unknown function with seven beta-strand methyltransferase motif similar to NOP2/YNL061W; green fluorescent protein (GFP)-fusion protein localizes to a single spot in the nucleus; YNL022C is not an essential gene | Putative GTPase that associates with pre-60S ribosomal subunits in the nucleolus and is required for their nuclear export and maturation | RNA terminal phosphate cyclase-like protein involved in rRNA processing at sites A0, A1, and A2; does not possess detectable RNA cyclase activity | Putative RNA exonuclease possibly involved in pre-rRNA processing and ribosome assembly | Part of actin cytoskeleton-regulatory complex Pan1p-Sla1p-End3p, associates with actin patches on the cell cortex; promotes protein-protein interactions essential for endocytosis; previously thought to be a subunit of poly(A) ribonuclease |
| 234 | 14 | 443 | -2.259 | 0.384 | 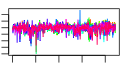 |  | 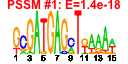 | 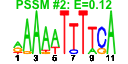 |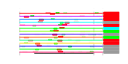  | YNL247W YBR142W YCL054W YDL031W YGR173W YGR187C YLR002C YLR222C YNL163C YNR054C YOL041C YPL212C YPL217C YPR137W | MAK5 SPB1 DBP10 RBG2 HGH1 NOC3 UTP13 RIA1 ESF2 NOP12 PUS1 BMS1 RRP9 | Essential protein of unknown function; may interact with ribosomes, based on co-purification experiments | Essential nucleolar protein, putative DEAD-box RNA helicase required for maintenance of M1 dsRNA virus; involved in biogenesis of large (60S) ribosomal subunits | AdoMet-dependent methyltransferase involved in rRNA processing and 60S ribosomal subunit maturation; methylates G2922 in the tRNA docking site of the large subunit rRNA and in the absence of snR52, U2921; suppressor of PAB1 mutants | Putative ATP-dependent RNA helicase of the DEAD-box protein family, constituent of 66S pre-ribosomal particles; essential protein involved in ribosome biogenesis | Protein with similarity to mammalian developmentally regulated GTP-binding protein | Protein of unknown function with similarity to human HMG1 and HMG2; localizes to the cytoplasm | Protein that forms a nuclear complex with Noc2p that binds to 66S ribosomal precursors to mediate their intranuclear transport; also binds to chromatin to promote the association of DNA replication factors and replication initiation | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Cytoplasmic GTPase involved in biogenesis of the 60S ribosome; has similarity to translation elongation factor 2 (Eft1p and Eft2p) | Essential nucleolar protein involved in pre-18S rRNA processing; component of the small subunit (SSU) processome; has sequence similarity to mABT1, a mouse transcription activator | Nucleolar protein, required for pre-25S rRNA processing; contains an RNA recognition motif (RRM) and has similarity to Nop13p, Nsr1p, and putative orthologs in Drosophila and S. pombe | tRNA:pseudouridine synthase, introduces pseudouridines at positions 26-28, 34-36, 65, and 67 of tRNA; nuclear protein that appears to be involved in tRNA export; also acts on U2 snRNA | Essential conserved nucleolar GTP-binding protein required for synthesis of 40S ribosomal subunits and for processing of the 35S pre-rRNA at sites A0, A1, and A2; interacts with Rcl1p, has similarity to Tsr1p | Protein involved in pre-rRNA processing, associated with U3 snRNP; component of small ribosomal subunit (SSU) processosome; ortholog of the human U3-55k protein |
| 51 | 11 | 443 | -2.154 | 0.366 |  |  | 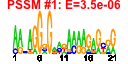 | 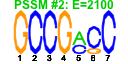 |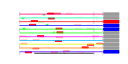  | YDR171W YER053C YER067W YFR014C YGR194C YKL091C YLR327C YMR105C YFR053C YGR008C YHR138C | HSP42 PIC2 CMK1 XKS1 TMA10 PGM2 HXK1 STF2 | Small heat shock protein (sHSP) with chaperone activity; forms barrel-shaped oligomers that suppress unfolded protein aggregation; involved in cytoskeleton reorganization after heat shock | Mitochondrial phosphate carrier, imports inorganic phosphate into mitochondria; functionally redundant with Mir1p but less abundant than Mir1p under normal conditions; expression is induced at high temperature | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YER067W is not an essential gene | Calmodulin-dependent protein kinase; may play a role in stress response, many CA++/calmodulan dependent phosphorylation substrates demonstrated in vitro, amino acid sequence similar to Cmk2p and mammalian Cam Kinase II | Xylulokinase, converts D-xylulose and ATP to xylulose 5-phosphate and ADP; rate limiting step in fermentation of xylulose; required for xylose fermentation by recombinant S. cerevisiae strains | Putative homolog of Sec14p, which is a phosphatidylinositol/phosphatidylcholine transfer protein involved in lipid metabolism; localizes to the nucleus | Protein of unknown function that associates with ribosomes | Phosphoglucomutase, catalyzes the conversion from glucose-1-phosphate to glucose-6-phosphate, which is a key step in hexose metabolism; functions as the acceptor for a Glc-phosphotransferase | Hexokinase isoenzyme 1, a cytosolic protein that catalyzes phosphorylation of glucose during glucose metabolism; expression is highest during growth on non-glucose carbon sources; glucose-induced repression involves the hexokinase Hxk2p | Protein involved in regulation of the mitochondrial F1F0-ATP synthase; Stf1p and Stf2p act as stabilizing factors that enhance inhibitory action of the Inh1p protein | Putative protein of unknown function; has similarity to Pbi2p; double null mutant lacking Pbi2p and Yhr138p exhibits highly fragmented vacuoles |
| 242 | 12 | 433 | -2.119 | 0.368 |  |  | 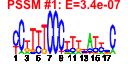 |  |  | YCR010C YGR201C YGR243W YIL057C YIL160C YKL217W YKR097W YML042W YMR107W YMR175W YPL223C YPR030W | ADY2 FMP43 POT1 JEN1 PCK1 CAT2 SPG4 SIP18 GRE1 CSR2 | Acetate transporter required for normal sporulation; phosphorylated in mitochondria | Putative protein of unknown function | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; expression induced under carbon limitation and repressed under high glucose | 3-ketoacyl-CoA thiolase with broad chain length specificity, cleaves 3-ketoacyl-CoA into acyl-CoA and acetyl-CoA during beta-oxidation of fatty acids | Lactate transporter, required for uptake of lactate and pyruvate; phosphorylated; expression is derepressed by transcriptional activator Cat8p during respiratory growth, and repressed in the presence of glucose, fructose, and mannose | Phosphoenolpyruvate carboxykinase, key enzyme in gluconeogenesis, catalyzes early reaction in carbohydrate biosynthesis, glucose represses transcription and accelerates mRNA degradation, regulated by Mcm1p and Cat8p, located in the cytosol | Carnitine acetyl-CoA transferase present in both mitochondria and peroxisomes, transfers activated acetyl groups to carnitine to form acetylcarnitine which can be shuttled across membranes | Protein required for survival at high temperature during stationary phase; not required for growth on nonfermentable carbon sources | Protein of unknown function whose expression is induced by osmotic stress | Hydrophilin of unknown function; stress induced (osmotic, ionic, oxidative, heat shock and heavy metals); regulated by the HOG pathway | Nuclear protein with a potential regulatory role in utilization of galactose and nonfermentable carbon sources; overproduction suppresses the lethality at high temperature of a chs5 spa2 double null mutation; potential Cdc28p substrate |
| 75 | 10 | 455 | -2.112 | 0.365 | 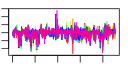 |  |  |  |  | YDL051W YDR321W YGR173W YGR200C YLR222C YNL174W YNL175C YOR095C YJL109C YLR409C | LHP1 ASP1 RBG2 ELP2 UTP13 NOP13 RKI1 UTP10 UTP21 | RNA binding protein required for maturation of tRNA and snRNA precursors; acts as a molecular chaperone for RNAs transcribed by polymerase III; homologous to human La (SS-B) autoantigen | Cytosolic L-asparaginase, involved in asparagine catabolism | Protein with similarity to mammalian developmentally regulated GTP-binding protein | Subunit of Elongator complex, which is required for modification of wobble nucleosides in tRNA; target of Kluyveromyces lactis zymocin | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Protein of unknown function, localizes to the nucleolus and nucleoplasm; contains an RNA recognition motif (RRM) and has similarity to Nop12p, which is required for processing of pre-18S rRNA | Ribose-5-phosphate ketol-isomerase, catalyzes the interconversion of ribose 5-phosphate and ribulose 5-phosphate in the pentose phosphate pathway; participates in pyridoxine biosynthesis | Possible U3 snoRNP protein involved in maturation of pre-18S rRNA, based on computational analysis of large-scale protein-protein interaction data |
| 86 | 18 | 441 | -2.083 | 0.407 |  |  | 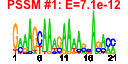 | 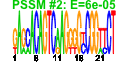 |  | YJR095W YKL093W YKL187C YMR114C YMR206W YMR322C YNL009W YOR391C YPL223C YPL280W YBL049W YGR289C YKL026C YLR312C YML054C YMR081C YNL195C YPR151C | SFC1 MBR1 SNO4 IDP3 HSP33 GRE1 HSP32 MOH1 MAL11 GPX1 QNQ1 CYB2 ISF1 SUE1 | Mitochondrial succinate-fumarate transporter, transports succinate into and fumarate out of the mitochondrion; required for ethanol and acetate utilization | Protein involved in mitochondrial functions and stress response; overexpression suppresses growth defects of hap2, hap3, and hap4 mutants | Putative protein of unknown function; the authentic, non-tagged protein is detected in a phosphorylated state in highly purified mitochondria in high-throughput studies | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and cytoplasm; YMR114C is not an essential gene | Putative protein of unknown function; YMR206W is not an essential gene | Possible chaperone and cysteine protease with similarity to E. coli Hsp31 and S. cerevisiae Hsp31p, Hsp32p, and Hsp33p; member of the DJ-1/ThiJ/PfpI superfamily; may have a role in pyridoxine metabolism | Peroxisomal NADP-dependent isocitrate dehydrogenase, catalyzes oxidation of isocitrate to alpha-ketoglutarate with the formation of NADP(H+), required for growth on unsaturated fatty acids | Possible chaperone and cysteine protease with similarity to E. coli Hsp31 and S. cerevisiae Hsp31p, Hsp32p, and Sno4p; member of the DJ-1/ThiJ/PfpI superfamily, which includes human DJ-1 involved in Parkinson's disease | Hydrophilin of unknown function; stress induced (osmotic, ionic, oxidative, heat shock and heavy metals); regulated by the HOG pathway | Possible chaperone and cysteine protease with similarity to E. coli Hsp31 and S. cerevisiae Hsp31p, Hsp33p, and Sno4p; member of the DJ-1/ThiJ/PfpI superfamily, which includes human DJ-1 involved in Parkinson's disease | Protein of unknown function, has homology to kinase Snf7p; not required for growth on nonfermentable carbon sources; essential for viability in stationary phase | Maltose permease, inducible high-affinity maltose transporter (alpha-glucoside transporter); encoded in the MAL1 complex locus; member of the 12 transmembrane domain superfamily of sugar transporters | Phospholipid hydroperoxide glutathione peroxidase induced by glucose starvation that protects cells from phospholipid hydroperoxides and nonphospholipid peroxides during oxidative stress | Protein of unknown function; null mutant quiescent and non-quiescent cells exhibit reduced reproductive capacity | Cytochrome b2 (L-lactate cytochrome-c oxidoreductase), component of the mitochondrial intermembrane space, required for lactate utilization; expression is repressed by glucose and anaerobic conditions | Serine-rich, hydrophilic protein with similarity to Mbr1p; overexpression suppresses growth defects of hap2, hap3, and hap4 mutants; expression is under glucose control; cotranscribed with NAM7 in a cyp1 mutant | Putative protein of unknown function; shares a promoter with YNL194C; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Mitochondrial protein required for degradation of unstable forms of cytochrome c |
| 12 | 13 | 445 | -2.002 | 0.398 |  |  | 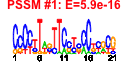 |  |  | YBR006W YBR026C YDL169C YDR258C YFL014W YHR209W YML128C YNL015W YOL083W YOR173W YOR285W YOR289W YPL196W | UGA2 ETR1 UGX2 HSP78 HSP12 CRG1 MSC1 TRX2 DCS2 OXR1 | Succinate semialdehyde dehydrogenase involved in the utilization of gamma-aminobutyrate (GABA) as a nitrogen source; part of the 4-aminobutyrate and glutamate degradation pathways; localized to the cytoplasm | 2-enoyl thioester reductase, member of the medium chain dehydrogenase/reductase family; localized to in mitochondria, where it has a probable role in fatty acid synthesis | Protein of unknown function, transcript accumulates in response to any combination of stress conditions | Oligomeric mitochondrial matrix chaperone that cooperates with Ssc1p in mitochondrial thermotolerance after heat shock; prevents the aggregation of misfolded matrix proteins; component of the mitochondrial proteolysis system | Plasma membrane localized protein that protects membranes from desiccation; induced by heat shock, oxidative stress, osmostress, stationary phase entry, glucose depletion, oleate and alcohol; regulated by the HOG and Ras-Pka pathways | Putative S-adenosylmethionine-dependent methyltransferase; mediates cantharidin resistance | Protein of unknown function; mutant is defective in directing meiotic recombination events to homologous chromatids; the authentic, non-tagged protein is detected in highly purified mitochondria and is phosphorylated | Cytoplasmic thioredoxin isoenzyme of the thioredoxin system which protects cells against oxidative and reductive stress, forms LMA1 complex with Pbi2p, acts as a cofactor for Tsa1p, required for ER-Golgi transport and vacuole inheritance | hypothetical protein | Non-essential, stress induced regulatory protein containing a HIT (histidine triad) motif; modulates m7G-oligoribonucleotide metabolism; inhibits Dcs1p; regulated by Msn2p, Msn4p, and the Ras-cAMP-cAPK signaling pathway, similar to Dcs1p. | Protein of unknown function, localized to the mitochondrial outer membrane | Putative protein of unknown function; transcription induced by the unfolded protein response; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Protein of unknown function required for normal levels of resistance to oxidative damage, null mutants are sensitive to hydrogen peroxide; member of a conserved family of proteins found in eukaryotes but not in prokaryotes |
| 197 | 13 | 460 | -1.961 | 0.466 |  |  |  | 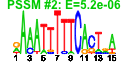 |  | YLR002C YNL001W YOR341W YDR365C YHR065C YJL010C YJL033W YKL078W YLR017W YLR435W YOR119C YPL012W YPR144C | NOC3 DOM34 RPA190 ESF1 RRP3 NOP9 HCA4 DHR2 MEU1 TSR2 RIO1 RRP12 NOC4 | Protein that forms a nuclear complex with Noc2p that binds to 66S ribosomal precursors to mediate their intranuclear transport; also binds to chromatin to promote the association of DNA replication factors and replication initiation | Endoribonuclease; functions in no-go mRNA decay, protein translation to promote G1 progression and differentiation, required for meiotic cell division; similar to the eukaryotic Pelota | RNA polymerase I subunit; largest subunit of RNA polymerase I | Nucleolar protein involved in pre-rRNA processing; depletion causes severely decreased 18S rRNA levels | Protein involved in rRNA processing; required for maturation of the 35S primary transcript of pre-rRNA and for cleavage leading to mature 18S rRNA; homologous to eIF-4a, which is a DEAD box RNA-dependent ATPase with helicase activity | Essential component of pre-40S ribosomes that is required for early cleavages of 35S pre-rRNA and hence formation of 18S rRNA; binds RNA in vitro and contains multiple pumilio-like repeats | Putative nucleolar DEAD box RNA helicase; high-copy number suppression of a U14 snoRNA processing mutant suggests an involvement in 18S rRNA synthesis | Predominantly nucleolar DEAH-box ATP-dependent RNA helicase, required for 18S rRNA synthesis | Methylthioadenosine phosphorylase (MTAP), catalyzes the initial step in the methionine salvage pathway; affects polyamine biosynthesis through regulation of ornithine decarboxylase (Spe1p) activity; regulates ADH2 gene expression | Protein with a potential role in pre-rRNA processing | Essential serine kinase involved in cell cycle progression and processing of the 20S pre-rRNA into mature 18S rRNA | Protein required for export of the ribosomal subunits; associates with the RNA components of the pre-ribosomes; contains HEAT-repeats | Nucleolar protein, forms a complex with Nop14p that mediates maturation and nuclear export of 40S ribosomal subunits |
| 290 | 19 | 443 | -1.832 | 0.438 |  |  | 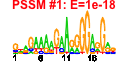 | 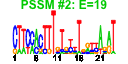 |  | YER035W YHR195W YKR009C YDR202C YDR204W YDR272W YER079W YER121W YGR130C YIL097W YIR014W YJL103C YJL141C YJL144W YJL161W YJL163C YLR149C YLR267W YML100W | EDC2 NVJ1 FOX2 RAV2 COQ4 GLO2 FYV10 GSM1 YAK1 FMP33 BOP2 BEM2 | RNA-binding protein, activates mRNA decapping directly by binding to the mRNA substrate and enhancing the activity of the decapping proteins Dcp1p and Dcp2p | Nuclear envelope protein that interacts with the vacuolar membrane protein Vac8p to promote formation of nucleus-vacuole junctions during piecemeal microautophagy of the nucleus (PMN) | Multifunctional enzyme of the peroxisomal fatty acid beta-oxidation pathway; has 3-hydroxyacyl-CoA dehydrogenase and enoyl-CoA hydratase activities | Subunit of RAVE (Rav1p, Rav2p, Skp1p), a complex that associates with the V1 domain of the vacuolar membrane (H+)-ATPase (V-ATPase) and promotes assembly and reassembly of the holoenzyme | Protein with a role in ubiquinone (Coenzyme Q) biosynthesis, possibly functioning in stabilization of Coq7p; located on the matrix face of the mitochondrial inner membrane; component of a mitochondrial ubiquinone-synthesizing complex | Cytoplasmic glyoxalase II, catalyzes the hydrolysis of S-D-lactoylglutathione into glutathione and D-lactate | Putative protein of unknown function | Putative protein of unknown function; may be involved in phosphatase regulation and/or generation of precursor metabolites and energy | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; specifically phosphorylated in vitro by mammalian diphosphoinositol pentakisphosphate (IP7) | Protein of unknown function, required for survival upon exposure to K1 killer toxin; involved in proteasome-dependent catabolite inactivation of fructose-1,6-bisphosphatase; contains CTLH domain | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; expression directly regulated by the metabolic and meiotic transcriptional regulator Ume6p; YIR014W is a non-essential gene | Putative zinc cluster protein of unknown function; proposed to be involved in the regulation of energy metabolism, based on patterns of expression and sequence analysis | Serine-threonine protein kinase that is part of a glucose-sensing system involved in growth control in response to glucose availability; translocates from the cytoplasm to the nucleus and phosphorylates Pop2p in response to a glucose signal | Cytoplasmic hydrophilin of unknown function, possibly involved in the dessication response; expression induced by osmotic stress, starvation and during stationary phase; GFP-fusion protein is induced by the DNA-damaging agent MMS | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; YLR149C is not an essential gene | Protein of unknown function, overproduction suppresses a pam1 slv3 double null mutation | Rho GTPase activating protein (RhoGAP) involved in the control of cytoskeleton organization and cellular morphogenesis; required for bud emergence |
| 38 | 15 | 439 | -1.804 | 0.356 |  |  |  |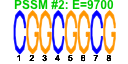  |  | YBL078C YER079W YGR008C YHR138C YIR038C YKL065C YKL142W YLR251W YMR090W YMR110C YBR230C YJR096W YLR345W YPL004C YPL203W | ATG8 STF2 GTT1 YET1 MRP8 SYM1 HFD1 OM14 LSP1 TPK2 | Protein required for autophagy; modified by the serial action of Atg4p, Atg7p, and Atg3p, and conjugated at the C terminus with phosphatidylethanolamine, to become the form essential for generation of autophagosomes | Putative protein of unknown function | Protein involved in regulation of the mitochondrial F1F0-ATP synthase; Stf1p and Stf2p act as stabilizing factors that enhance inhibitory action of the Inh1p protein | Putative protein of unknown function; has similarity to Pbi2p; double null mutant lacking Pbi2p and Yhr138p exhibits highly fragmented vacuoles | ER associated glutathione S-transferase capable of homodimerization; expression induced during the diauxic shift and throughout stationary phase; functional overlap with Gtt2p, Grx1p, and Grx2p | Endoplasmic reticulum transmembrane protein; may interact with ribosomes, based on co-purification experiments; homolog of human BAP31 protein | Putative mitochondrial ribosomal protein, has similarity to E. coli ribosomal protein S2 | Protein required for ethanol metabolism; induced by heat shock and localized to the inner mitochondrial membrane; homologous to mammalian peroxisomal membrane protein Mpv17 | Putative protein of unknown function with similarity to DTDP-glucose 4,6-dehydratases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YMR090W is not an essential gene | Putative fatty aldehyde dehydrogenase, located in the mitochondrial outer membrane and also in lipid particles; has similarity to human fatty aldehyde dehydrogenase (FALDH) which is implicated in Sjogren-Larsson syndrome | Integral mitochondrial outer membrane protein; abundance is decreased in cells grown in glucose relative to other carbon sources; appears to contain 3 alpha-helical transmembrane segments; ORF encodes a 97-basepair intron | Putative xylose and arabinose reductase; member of the aldo-keto reductase (AKR) family; GFP-fusion protein is induced in response to the DNA-damaging agent MMS | Similar to 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase enzymes responsible for the metabolism of fructoso-2,6-bisphosphate; mRNA expression is repressed by the Rfx1p-Tup1p-Ssn6p repressor complex; YLR345W is not an essential gene | Primary component of eisosomes, which are large immobile patch structures at the cell cortex associated with endocytosis, along with Pil1p and Sur7p; null mutants show activation of Pkc1p/Ypk1p stress resistance pathways | cAMP-dependent protein kinase catalytic subunit; promotes vegetative growth in response to nutrients via the Ras-cAMP signaling pathway; inhibited by regulatory subunit Bcy1p in the absence of cAMP; partially redundant with Tpk1p and Tpk3p |
| 148 | 20 | 438 | -1.764 | 0.409 |  |  |  |  |  | YDR272W YHR112C YIL111W YJL161W YKL150W YLR270W YOR035C YOR120W YBL078C YDL021W YJL155C YKL091C YLR251W YLR252W YML004C YML070W YOR036W YOR121C YOR289W YPL123C | GLO2 COX5B FMP33 MCR1 DCS1 SHE4 GCY1 ATG8 GPM2 FBP26 SYM1 GLO1 DAK1 PEP12 RNY1 | Cytoplasmic glyoxalase II, catalyzes the hydrolysis of S-D-lactoylglutathione into glutathione and D-lactate | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Subunit Vb of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; predominantly expressed during anaerobic growth while its isoform Va (Cox5Ap) is expressed during aerobic growth | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Mitochondrial NADH-cytochrome b5 reductase, involved in ergosterol biosynthesis | Non-essential hydrolase involved in mRNA decapping, may function in a feedback mechanism to regulate deadenylation, contains pyrophosphatase activity and a HIT (histidine triad) motif; interacts with neutral trehalase Nth1p | Protein containing a UCS (UNC-45/CRO1/SHE4) domain, binds to myosin motor domains to regulate myosin function; involved in endocytosis, polarization of the actin cytoskeleton, and asymmetric mRNA localization | Putative NADP(+) coupled glycerol dehydrogenase, proposed to be involved in an alternative pathway for glycerol catabolism; member of the aldo-keto reductase (AKR) family | Protein required for autophagy; modified by the serial action of Atg4p, Atg7p, and Atg3p, and conjugated at the C terminus with phosphatidylethanolamine, to become the form essential for generation of autophagosomes | Homolog of Gpm1p phosphoglycerate mutase which converts 3-phosphoglycerate to 2-phosphoglycerate in glycolysis; may be non-functional derivative of a gene duplication event | Fructose-2,6-bisphosphatase, required for glucose metabolism | Putative homolog of Sec14p, which is a phosphatidylinositol/phosphatidylcholine transfer protein involved in lipid metabolism; localizes to the nucleus | Protein required for ethanol metabolism; induced by heat shock and localized to the inner mitochondrial membrane; homologous to mammalian peroxisomal membrane protein Mpv17 | Monomeric glyoxalase I, catalyzes the detoxification of methylglyoxal (a by-product of glycolysis) via condensation with glutathione to produce S-D-lactoylglutathione; expression regulated by methylglyoxal levels and osmotic stress | Dihydroxyacetone kinase, required for detoxification of dihydroxyacetone (DHA); involved in stress adaptation | Target membrane receptor (t-SNARE) for vesicular intermediates traveling between the Golgi apparatus and the vacuole; controls entry of biosynthetic, endocytic, and retrograde traffic into the prevacuolar compartment; syntaxin | Putative protein of unknown function; transcription induced by the unfolded protein response; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | RNAse; member of the T(2) family of endoribonucleases |
| 278 | 15 | 443 | -1.722 | 0.394 |  |  |  | 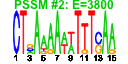 |  | YDL051W YDR120C YDR399W YDR412W YDR413C YDR449C YGL169W YGR158C YGR200C YIR026C YJR063W YKL144C YLR073C YMR093W YOR021C | LHP1 TRM1 HPT1 RRP17 UTP6 SUA5 MTR3 ELP2 YVH1 RPA12 RPC25 UTP15 | RNA binding protein required for maturation of tRNA and snRNA precursors; acts as a molecular chaperone for RNAs transcribed by polymerase III; homologous to human La (SS-B) autoantigen | tRNA methyltransferase, localizes to both the nucleus and mitochondrion to produce the modified base N2,N2-dimethylguanosine in tRNAs in both compartments | Dimeric hypoxanthine-guanine phosphoribosyltransferase, catalyzes the formation of both inosine monophosphate and guanosine monophosphate; mutations in the human homolog HPRT1 can cause Lesch-Nyhan syndrome and Kelley-Seegmiller syndrome | Protein required for cell viability | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Protein required for respiratory growth; null mutation suppresses the Cyc1p translation defect caused by the presence of an aberrant ATG codon upstream of the correct start | 3'5' exoribonuclease, exosome subunit; nucleolar protein involved in export of mRNA and ribosomal subunits; homologous to the E. coli exonuclease RNase PH | Subunit of Elongator complex, which is required for modification of wobble nucleosides in tRNA; target of Kluyveromyces lactis zymocin | Protein phosphatase involved in vegetative growth at low temperatures, sporulation, and glycogen accumulation; transcription induced by low temperature and nitrogen starvation; member of the dual-specificity family of protein phosphatases | RNA polymerase I subunit A12.2; contains two zinc binding domains, and the N terminal domain is responsible for anchoring to the RNA pol I complex | RNA polymerase III subunit C25 | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to endosomes; YLR073C is not an esssential gene | Putative protein of unknown function; YOR021C is not an essential gene |
| 198 | 23 | 436 | -1.705 | 0.410 |  |  | 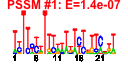 | 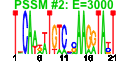 |  | YBR078W YEL042W YFL045C YJL138C YKL081W YKR059W YLR372W YMR079W YBR162C YBR283C YCR034W YEL040W YER043C YGL008C YGL225W YHR068W YJR143C YLR083C YMR215W YMR305C YMR307W YPR074C YPR119W | ECM33 GDA1 SEC53 TIF2 TEF4 TIF1 APA1 SEC14 TOS1 SSH1 FEN1 UTR2 SAH1 PMA1 VRG4 DYS1 PMT4 EMP70 GAS3 SCW10 GAS1 TKL1 CLB2 | GPI-anchored protein of unknown function, has a possible role in apical bud growth; GPI-anchoring on the plasma membrane crucial to function; phosphorylated in mitochondria; similar to Sps2p and Pst1p | Guanosine diphosphatase located in the Golgi, involved in the transport of GDP-mannose into the Golgi lumen by converting GDP to GMP after mannose is transferred its substrate | Phosphomannomutase, involved in synthesis of GDP-mannose and dolichol-phosphate-mannose; required for folding and glycosylation of secretory proteins in the ER lumen | Translation initiation factor eIF4A, identical to Tif1p; DEA(D/H)-box RNA helicase that couples ATPase activity to RNA binding and unwinding; forms a dumbbell structure of two compact domains connected by a linker; interacts with eIF4G | Translation elongation factor EF-1 gamma | Translation initiation factor eIF4A, identical to Tif2p; DEA(D/H)-box RNA helicase that couples ATPase activity to RNA binding and unwinding; forms a dumbbell structure of two compact domains connected by a linker; interacts with eIF4G | Diadenosine 5',5''-P1,P4-tetraphosphate phosphorylase I (AP4A phosphorylase), involved in catabolism of bis(5'-nucleosidyl) tetraphosphates; has similarity to Apa2p | Phosphatidylinositol/phosphatidylcholine transfer protein involved in coordinate regulation of PtdIns and PtdCho metabolism, products of which are regulators in Golgi to plasma membrane transport; functionally homologous to mammalian PITPs | Covalently-bound cell wall protein of unknown function; identified as a cell cycle regulated SBF target gene; deletion mutants are highly resistant to treatment with beta-1,3-glucanase; has sequence similarity to YJL171C | Subunit of the Ssh1 translocon complex; Sec61p homolog involved in co-translational pathway of protein translocation; not essential | Fatty acid elongase, involved in sphingolipid biosynthesis; acts on fatty acids of up to 24 carbons in length; mutations have regulatory effects on 1,3-beta-glucan synthase, vacuolar ATPase, and the secretory pathway | Cell wall protein that functions in the transfer of chitin to beta(1-6)glucan; putative chitin transglycosidase; glycosylphosphatidylinositol (GPI)-anchored protein localized to the bud neck; has a role in cell wall maintenance | S-adenosyl-L-homocysteine hydrolase, catabolizes S-adenosyl-L-homocysteine which is formed after donation of the activated methyl group of S-adenosyl-L-methionine (AdoMet) to an acceptor | Plasma membrane H+-ATPase, pumps protons out of the cell; major regulator of cytoplasmic pH and plasma membrane potential; part of the P2 subgroup of cation-transporting ATPases | Golgi GDP-mannose transporter; regulates Golgi function and glycosylation in Golgi | Deoxyhypusine synthase, catalyzes formation of deoxyhypusine, the first step in hypusine biosynthesis; triggers posttranslational hypusination of translation elongation factor eIF-5A and regulates its intracellular levels; tetrameric | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; appears to form homodimers in vivo and does not complex with other Pmt proteins; target for new antifungals | Protein whose 24kDa cleavage product is found in endosome-enriched membrane fractions, predicted to be a transmembrane protein | Putative 1,3-beta-glucanosyltransferase, has similarity to Gas1p; localizes to the cell wall | Cell wall protein with similarity to glucanases; may play a role in conjugation during mating based on mutant phenotype and its regulation by Ste12p | Beta-1,3-glucanosyltransferase, required for cell wall assembly; localizes to the cell surface via a glycosylphosphatidylinositol (GPI) anchor | Transketolase, similar to Tkl2p; catalyzes conversion of xylulose-5-phosphate and ribose-5-phosphate to sedoheptulose-7-phosphate and glyceraldehyde-3-phosphate in the pentose phosphate pathway; needed for synthesis of aromatic amino acids | B-type cyclin involved in cell cycle progression; activates Cdc28p to promote the transition from G2 to M phase; accumulates during G2 and M, then targeted via a destruction box motif for ubiquitin-mediated degradation by the proteasome |
| 101 | 35 | 450 | -1.702 | 0.536 |  |  | 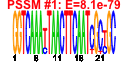 | 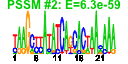 |  | YAR020C YBR182C YDR076W YDR542W YER185W YFL063W YGL249W YHL043W YHR210C YIL174W YIL175W YJR162C YLL065W YLR155C YLR157C YLR158C YLR160C YMR325W YNL337W YOR177C YBR076W YDR540C YEL049W YFL020C YGL230C YGR294W YHL046C YIL176C YIR041W YJL223C YLL064C YLR037C YOL161C YOR044W YPL278C | PAU7 SMP1 RAD55 PAU10 PUG1 ZIP2 ECM34 ASP3-1 PAU19 MPC54 ECM8 IRC4 PAU2 PAU5 PAU12 PAU13 PAU14 PAU15 PAU1 PAU18 DAN2 PAU20 IRC23 | Part of 23-member seripauperin multigene family, active during alcoholic fermentation, regulated by anaerobiosis, inhibited by oxygen, repressed by heme | Putative transcription factor involved in regulating the response to osmotic stress; member of the MADS-box family of transcription factors | Protein that stimulates strand exchange by stabilizing the binding of Rad51p to single-stranded DNA; involved in the recombinational repair of double-strand breaks in DNA during vegetative growth and meiosis; forms heterodimer with Rad57p | hypothetical protein | Plasma membrane protein involved in protoporphyrin uptake; similar in sequence to RSB1, which is involved in sphingoid long-chain base release | Meiosis-specific protein involved in normal synaptonemal complex formation and pairing between homologous chromosomes during meiosis | Putative protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Putative protein of unknown function; non-essential gene; highly expressed under anaeorbic conditions; sequence similarity to aldose 1-epimerases such as GAL10 | Cell-wall L-asparaginase II, involved in asparagine catabolism; expression is induced during nitrogen starvation; four copies of ASP3 are present in the genome reference strain S288C | Component of the meiotic outer plaque, a membrane-organizing center which is assembled on the cytoplasmic face of the spindle pole body during meiosis II and triggers the formation of the prospore membrane; potential Cdc28p substrate | Non-essential protein of unknown function | hypothetical protein; null mutant displays increased levels of spontaneous Rad52 foci | Part of 23-member seripauperin multigene family encoded mainly in subtelomeric regions, active during alcoholic fermentation, regulated by anaerobiosis, negatively regulated by oxygen, repressed by heme | Putative protein of unknown function; non-essential gene | Putative protein of unknown function; not an essential gene | Cell wall mannoprotein with similarity to Tir1p, Tir2p, Tir3p, and Tir4p; expressed under anaerobic conditions, completely repressed during aerobic growth | Putative protein of unknown function; green fluorescent protein (GFP)-fusion localizes to the ER; null mutant displays increased levels of spontaneous Rad52 foci | Putative protein of unknown function; gene expression regulated by copper levels |
| 257 | 17 | 444 | -1.688 | 0.391 |  |  |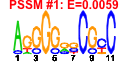  | 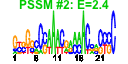 |  | YBR056W YBR149W YDL022W YMR251W YBR126C YBR214W YCL040W YCL042W YDL023C YDR074W YDR513W YDR516C YGL037C YGR194C YHR087W YNL305C YOR120W | ARA1 GPD1 GTO3 TPS1 SDS24 GLK1 TPS2 GRX2 EMI2 PNC1 XKS1 GCY1 | Putative cytoplasmic protein of unknown function | NADP+ dependent arabinose dehydrogenase, involved in carbohydrate metabolism; purified as homodimer; naturally occurs with a N-terminus degradation product | NAD-dependent glycerol-3-phosphate dehydrogenase, key enzyme of glycerol synthesis, essential for growth under osmotic stress; expression regulated by high-osmolarity glycerol response pathway; homolog of Gpd2p | Omega class glutathione transferase; putative cytosolic localization | Synthase subunit of trehalose-6-phosphate synthase/phosphatase complex, which synthesizes the storage carbohydrate trehalose; also found in a monomeric form; expression is induced by the stress response and repressed by the Ras-cAMP pathway | One of two S. cerevisiae homologs (Sds23p and Sds24p) of the Schizosaccharomyces pombe Sds23 protein, which genetic studies have implicated in APC/cyclosome regulation; may play an indirect role in fluid-phase endocytosis | Glucokinase, catalyzes the phosphorylation of glucose at C6 in the first irreversible step of glucose metabolism; one of three glucose phosphorylating enzymes; expression regulated by non-fermentable carbon sources | Putative protein of unknown function; epitope-tagged protein localizes to the cytoplasm | Phosphatase subunit of the trehalose-6-phosphate synthase/phosphatase complex, which synthesizes the storage carbohydrate trehalose; expression is induced by stress conditions and repressed by the Ras-cAMP pathway | Cytoplasmic glutaredoxin, thioltransferase, glutathione-dependent disulfide oxidoreductase involved in maintaining redox state of target proteins, also exhibits glutathione peroxidase activity, expression induced in response to stress | Non-essential protein of unknown function required for transcriptional induction of the early meiotic-specific transcription factor IME1; required for sporulation; expression is regulated by glucose-repression transcription factors Mig1/2p | Nicotinamidase that converts nicotinamide to nicotinic acid as part of the NAD(+) salvage pathway, required for life span extension by calorie restriction; PNC1 expression responds to all known stimuli that extend replicative life span | Xylulokinase, converts D-xylulose and ATP to xylulose 5-phosphate and ADP; rate limiting step in fermentation of xylulose; required for xylose fermentation by recombinant S. cerevisiae strains | Protein of unknown function involved in RNA metabolism; this single domain protein has structural similarity to the N-terminal domain of SBDS, the human protein mutated in Shwachman-Diamond Syndrome (the yeast SBDS ortholog = SDO1/YLR022C) | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; YNL305C is not an essential gene | Putative NADP(+) coupled glycerol dehydrogenase, proposed to be involved in an alternative pathway for glycerol catabolism; member of the aldo-keto reductase (AKR) family |
| 62 | 46 | 439 | -1.646 | 0.539 |  |  | 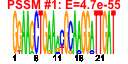 |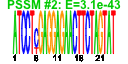  |  | YAR009C YAR010C YBL005W YBR012C YCL019W YCL020W YCL021W YDR089W YDR149C YDR340W YDR366C YER138C YER160C YGL207W YGL239C YGR039W YGR065C YGR114C YHR114W YHR145C YIL060W YJL110C YJR026W YJR027W YJR028W YJR029W YML039W YML040W YML045W YML113W YMR045C YMR046C YMR050C YMR051C YMR083W YOL056W YOL106W YOR262W YOR282W YPL075W YPR197C YBR013C YDL128W YFR056C YIL059C YOL104C | PDR3 YCL021W-A SPT16 VHT1 BZZ1 GZF3 DAT1 ADH3 GPM3 GCR1 VCX1 TMT1 | Retrotransposon TYA Gag and TYB Pol genes; in YARCTY1-1 TYB is mutant and probably non-functional | Retrotransposon TYA Gag gene co-transcribed with TYB Pol; in YARCTY1-1 TYB is mutant and probably non-functional | Transcriptional activator of the pleiotropic drug resistance network, regulates expression of ATP-binding cassette (ABC) transporters through binding to cis-acting sites known as PDREs (PDR responsive elements) | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Retrotransposon TYA Gag gene co-transcribed with TYB Pol; translated as TYA or TYA-TYB polyprotein; Gag is a nucleocapsid protein that is the structural constituent of virus-like particles (VLPs); similar to retroviral Gag | Putative protein of unknown function | Protein of unknown function; deletion confers resistance to Nickel | hypothetical protein | Subunit of the heterodimeric FACT complex (Spt16p-Pob3p), facilitates RNA Polymerase II transcription elongation through nucleosomes by destabilizing and then reassembling nucleosome structure | High-affinity plasma membrane H+-biotin (vitamin H) symporter; mutation results in fatty acid auxotrophy; 12 transmembrane domain containing major facilitator subfamily member; mRNA levels negatively regulated by iron deprivation and biotin | SH3 domain protein implicated in the regulation of actin polymerization, able to recruit actin polymerization machinery through its SH3 domains, colocalizes with cortical actin patches and Las17p, interacts with type I myosins | Putative protein of unknown function; mutant accumulates less glycogen than does wild type; YIL060W is not an essential gene | GATA zinc finger protein and Dal80p homolog that negatively regulates nitrogen catabolic gene expression by competing with Gat1p for GATA site binding; function requires a repressive carbon source; dimerizes with Dal80p and binds to Tor1p | DNA binding protein that recognizes oligo(dA).oligo(dT) tracts; Arg side chain in its N-terminal pentad Gly-Arg-Lys-Pro-Gly repeat is required for DNA-binding; not essential for viability | Mitochondrial alcohol dehydrogenase isozyme III; involved in the shuttling of mitochondrial NADH to the cytosol under anaerobic conditions and ethanol production | Homolog of Gpm1p phosphoglycerate mutase, which converts 3-phosphoglycerate to 2-phosphoglycerate in glycolysis; may be non-functional derivative of a gene duplication event | Cytoplasmic protein of unknown function; essential gene with similarity to YLR243W; contains an ATP/GTP binding site motif | Transcriptional activator of genes involved in glycolysis; DNA-binding protein that interacts and functions with the transcriptional activator Gcr2p | Putative protein of unknown function, haploid deletion mutant exhibits synthetic phenotype with alpha-synuclein | Vacuolar H+/Ca2+ exchanger involved in control of cytosolic Ca2+ concentration; has similarity to sodium/calcium exchangers, including the bovine Na+/Ca2+,K+ antiporter | Trans-aconitate methyltransferase, cytosolic enzyme that catalyzes the methyl esterification of 3-isopropylmalate, an intermediate of the leucine biosynthetic pathway, and trans-aconitate, which inhibits the citric acid cycle |
| 176 | 16 | 445 | -1.646 | 0.388 |  |  | 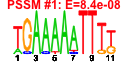 | 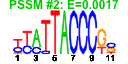 |  | YJR007W YMR146C YPL237W YBR143C YGL097W YJL111W YJR064W YLR293C YOL139C YOR046C YOR260W YOR276W YOR361C YPL037C YPL081W YPR041W | SUI2 TIF34 SUI3 SUP45 SRM1 CCT7 CCT5 GSP1 CDC33 DBP5 GCD1 CAF20 PRT1 EGD1 RPS9A TIF5 | Alpha subunit of the translation initiation factor eIF2, involved in the identification of the start codon; phosphorylation of Ser51 is required for regulation of translation by inhibiting the exchange of GDP for GTP | Subunit of the core complex of translation initiation factor 3(eIF3), which is essential for translation | Beta subunit of the translation initiation factor eIF2, involved in the identification of the start codon; proposed to be involved in mRNA binding | Polypeptide release factor involved in translation termination; mutant form acts as a recessive omnipotent suppressor | Nucleotide exchange factor for Gsp1p, localizes to the nucleus, required for nucleocytoplasmic trafficking of macromolecules; suppressor of the pheromone response pathway; potentially phosphorylated by Cdc28p | Subunit of the cytosolic chaperonin Cct ring complex, related to Tcp1p, required for the assembly of actin and tubulins in vivo | GTP binding protein (mammalian Ranp homolog) involved in the maintenance of nuclear organization, RNA processing and transport; regulated by Prp20p, Rna1p, Yrb1p, Yrb2p, Yrp4p, Yrb30p, Cse1p and Kap95p; yeast Gsp2p homolog | Cytoplasmic mRNA cap binding protein; the eIF4E-cap complex is responsible for mediating cap-dependent mRNA translation via interactions with the translation initiation factor eIF4G (Tif4631p or Tif4632p) | Cytoplasmic ATP-dependent RNA helicase of the DEAD-box family involved in mRNA export from the nucleus; involved in translation termination | Gamma subunit of the translation initiation factor eIF2B, the guanine-nucleotide exchange factor for eIF2; activity subsequently regulated by phosphorylated eIF2; first identified as a negative regulator of GCN4 expression | Phosphoprotein of the mRNA cap-binding complex involved in translational control, repressor of cap-dependent translation initiation, competes with eIF4G for binding to eIF4E | Subunit of the core complex of translation initiation factor 3(eIF3), essential for translation; part of a subcomplex (Prt1p-Rpg1p-Nip1p) that stimulates binding of mRNA and tRNA(i)Met to ribosomes | Subunit beta1 of the nascent polypeptide-associated complex (NAC) involved in protein targeting, associated with cytoplasmic ribosomes; enhances DNA binding of the Gal4p activator; homolog of human BTF3b | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps9Bp and has similarity to E. coli S4 and rat S9 ribosomal proteins | Translation initiation factor eIF-5; N-terminal domain functions as a GTPase-activating protein to mediate hydrolysis of ribosome-bound GTP; C-terminal domain is the core of ribosomal preinitiation complex formation |
| 295 | 20 | 438 | -1.630 | 0.464 |  |  | 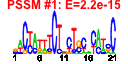 | 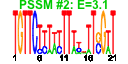 |  | YFR017C YGL259W YGR250C YGR287C YIL099W YIL101C YIL172C YIR039C YJL221C YJR066W YKL151C YKL187C YLR327C YMR104C YMR169C YOL047C YOL084W YOL157C YOR374W YPR184W | YPS5 SGA1 XBP1 YPS6 FSP2 TOR1 TMA10 YPK2 ALD3 PHM7 ALD4 GDB1 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and is induced in response to the DNA-damaging agent MMS; YFR017C is not an essential gene | Protein with similarity to GPI-anchored aspartic proteases such as Yap1p and Yap3p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; has similarity to alpha-D-glucosidase (maltase); authentic, non-tagged protein detected in purified mitochondria in high-throughput studies | Intracellular sporulation-specific glucoamylase involved in glycogen degradation; induced during starvation of a/a diploids late in sporulation, but dispensable for sporulation | Transcriptional repressor that binds to promoter sequences of the cyclin genes, CYS3, and SMF2; expression is induced by stress or starvation during mitosis, and late in meiosis; member of the Swi4p/Mbp1p family; potential Cdc28p substrate | Putative protein of unknown function with similarity to glucosidases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative GPI-anchored aspartic protease | Protein of unknown function, expression is induced during nitrogen limitation | PIK-related protein kinase and rapamycin target; subunit of TORC1, a complex that controls growth in response to nutrients by regulating translation, transcription, ribosome biogenesis, nutrient transport and autophagy; involved in meiosis | Putative protein of unknown function; the authentic, non-tagged protein is detected in a phosphorylated state in highly purified mitochondria in high-throughput studies | Protein of unknown function that associates with ribosomes | Protein kinase with similarityto serine/threonine protein kinase Ypk1p; functionally redundant with YPK1 at the genetic level; participates in a signaling pathway required for optimal cell wall integrity; homolog of mammalian kinase SGK | Cytoplasmic aldehyde dehydrogenase, involved in beta-alanine synthesis; uses NAD+ as the preferred coenzyme; very similar to Ald2p; expression is induced by stress and repressed by glucose | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Protein of unknown function, expression is regulated by phosphate levels; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery and vacuole | Putative protein of unknown function | Mitochondrial aldehyde dehydrogenase, required for growth on ethanol and conversion of acetaldehyde to acetate; phosphorylated; activity is K+ dependent; utilizes NADP+ or NAD+ equally as coenzymes; expression is glucose repressed | Glycogen debranching enzyme containing glucanotranferase and alpha-1,6-amyloglucosidase activities, required for glycogen degradation; phosphorylated in mitochondria |
| 104 | 31 | 438 | -1.591 | 0.557 | 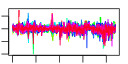 |  | 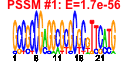 | 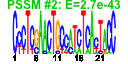 |  | YBR302C YCL049C YDL237W YDL248W YDR055W YEL060C YFL062W YGR149W YGR295C YIR044C YJL015C YJL031C YJR073C YJR161C YML058W YML131W YML132W YMR191W YNL073W YNL074C YNL208W YNL322C YNL336W YDR077W YDR129C YDR435C YGR244C YHL048W YIL034C YIR043C YKL219W | COS2 LRC1 COS7 PST1 PRB1 COS4 COS6 BET4 OPI3 COS5 HUG1 COS3 SPG5 MSK1 MLF3 KRE1 COS1 SED1 SAC6 PPM1 LSC2 COS8 CAP2 COS9 | Protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Protein of unknown function; localizes to membrane fraction; YCL049C is not an essential gene | Putative protein of unknown function; YDL237W is not an essential gene; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Cell wall protein that contains a putative GPI-attachment site; secreted by regenerating protoplasts; up-regulated by activation of the cell integrity pathway, as mediated by Rlm1p; upregulated by cell wall damage via disruption of FKS1 | Vacuolar proteinase B (yscB), a serine protease of the subtilisin family; involved in protein degradation in the vacuole and required for full protein degradation during sporulation | Putative protein of unknown function; predicted to be an integal membrane protein | Alpha subunit of Type II geranylgeranyltransferase required for vesicular transport between the endoplasmic reticulum and the Golgi; provides a membrane attachment moiety to Rab-like proteins Ypt1p and Sec4p | Phospholipid methyltransferase (methylene-fatty-acyl-phospholipid synthase), catalyzes the last two steps in phosphatidylcholine biosynthesis | Protein of unknown function, member the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Protein involved in the Mec1p-mediated checkpoint pathway that responds to DNA damage or replication arrest, transcription is induced by DNA damage | Putative protein of unknown function with similarity to oxidoreductases; HOG1 and SKO1-dependent mRNA expression is induced after osmotic shock; GFP-fusion protein localizes to the cytoplasm and is induced by the DNA-damaging agent MMS | Protein involved in salt resistance; interacts with sodium:hydrogen antiporter Nha1p; member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Protein required for survival at high temperature during stationary phase; not required for growth on nonfermentable carbon sources | Mitochondrial lysine-tRNA synthetase, required for import of both aminoacylated and deacylated forms of tRNA(Lys) into mitochondria | Serine-rich protein of unknown function; overproduction suppresses the growth inhibition caused by exposure to the immunosuppressant leflunomide | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; authentic, non-tagged protein is detected in purified mitochondria in high-throughput studies; potential orthologs found in other fungi | Cell wall glycoprotein involved in beta-glucan assembly; serves as a K1 killer toxin membrane receptor | Major stress-induced structural GPI-cell wall glycoprotein in stationary-phase cells, associates with translating ribosomes, possible role in mitochondrial genome maintenance; ORF contains two distinct variable minisatellites | Fimbrin, actin-bundling protein; cooperates with Scp1p (calponin/transgelin) in the organization and maintenance of the actin cytoskeleton | Carboxyl methyl transferase, methylates the C terminus of the protein phosphatase 2A catalytic subunit (Pph21p or Pph22p), which is important for complex formation with regulatory subunits | Beta subunit of succinyl-CoA ligase, which is a mitochondrial enzyme of the TCA cycle that catalyzes the nucleotide-dependent conversion of succinyl-CoA to succinate | Nuclear membrane protein, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins; regulation suggests a potential role in the unfolded protein response | Beta subunit of the capping protein (CP) heterodimer (Cap1p and Cap2p) which binds to the barbed ends of actin filaments preventing further polymerization; localized predominantly to cortical actin patches |
| 177 | 29 | 435 | -1.511 | 0.440 | 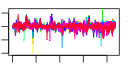 |  | 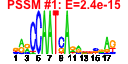 | 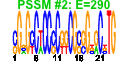 |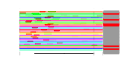  | YDL004W YDL181W YDR529C YGL191W YJL166W YKL016C YKL141W YBR039W YDL067C YDR298C YDR377W YEL024W YFR033C YGL187C YGL188C YGR182C YGR183C YHR051W YJR077C YJR121W YLR038C YLR294C YLR295C YLR395C YMR256C YNL052W YPL078C YPL271W YPR020W | ATP16 INH1 QCR7 COX13 QCR8 ATP7 SDH3 ATP3 COX9 ATP5 ATP17 NUP42 QCR6 COX4 YGL188C-A QCR9 COX6 MIR1 ATP2 COX12 ATP14 COX8 COX7 COX5A ATP4 ATP15 ATP20 | Delta subunit of the central stalk of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Protein that inhibits ATP hydrolysis by the F1F0-ATP synthase, inhibitory function is enhanced by stabilizing proteins Stf1p and Stf2p; has similarity to Stf1p and both Inh1p and Stf1p exhibit the potential to form coiled-coil structures | Subunit 7 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; oriented facing the mitochondrial matrix; N-terminus appears to play a role in complex assembly | Subunit VIa of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; not essential for cytochrome c oxidase activity but may modulate activity in response to ATP | Subunit 8 of ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; oriented facing the intermembrane space; expression is regulated by Abf1p and Cpf1p | Subunit d of the stator stalk of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | Cytochrome b subunit of succinate dehydrogenase (Sdh1p, Sdh2p, Sdh3p, Sdh4p), which couples the oxidation of succinate to the transfer of electrons to ubiquinone | Gamma subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | Subunit VIIa of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain | Subunit 5 of the stator stalk of mitochondrial F1F0 ATP synthase, which is an evolutionarily conserved enzyme complex required for ATP synthesis; homologous to bovine subunit OSCP (oligomycin sensitivity-conferring protein); phosphorylated | Subunit f of the F0 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | Subunit of the nuclear pore complex (NPC) that localizes exclusively to the cytoplasmic side; involved in RNA export, most likely at a terminal step; interacts with Gle1p | Subunit 6 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; highly acidic protein; required for maturation of cytochrome c1 | Subunit IV of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; N-terminal 25 residues of precursor are cleaved during mitochondrial import; phosphorylated | Putative protein of unknown function | Subunit 9 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; required for electron transfer at the ubiquinol oxidase site of the complex | Subunit VI of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; expression is regulated by oxygen levels | Mitochondrial phosphate carrier, imports inorganic phosphate into mitochondria; functionally redundant with Pic2p but more abundant than Pic2p under normal conditions; phosphorylated | Beta subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Subunit VIb of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; required for assembly of cytochrome c oxidase but not required for activity after assembly; phosphorylated | Subunit h of the F0 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | Subunit VIII of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain | Subunit VII of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain | Subunit Va of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; predominantly expressed during aerobic growth while its isoform Vb (Cox5Bp) is expressed during anaerobic growth | Subunit b of the stator stalk of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Epsilon subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Subunit g of the mitochondrial F1F0 ATP synthase; reversibly phosphorylated on two residues; unphosphorylated form is required for dimerization of the ATP synthase complex |
| 188 | 17 | 447 | -1.397 | 0.384 |  |  |  |  |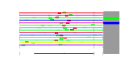  | YBL028C YBR266C YCL053C YFR001W YIL096C YKL099C YLR068W YLR221C YMR269W YOR252W YOR287C YPL146C YPR142C YPR143W YBR267W YBR271W YGL029W | LOC1 UTP11 FYV7 RSA3 TMA23 TMA16 NOP53 RRP15 REI1 CGR1 | Protein of unknown function that may interact with ribosomes; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus; predicted to be involved in ribosome biogenesis | Nuclear protein involved in asymmetric localization of ASH1 mRNA; binds double-stranded RNA in vitro; constituent of 66S pre-ribosomal particles | Putative protein of unknown function; associates with precursors of the 60S ribosomal subunit | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Protein of unknown function, required for survival upon exposure to K1 killer toxin; involved in processing the 35S rRNA primary transcript to generate the 20S and 27SA2 pre-rRNA transcripts | Protein with a likely role in ribosomal maturation, required for accumulation of wild-type levels of large (60S) ribosomal subunits; binds to the helicase Dbp6p in pre-60S ribosomal particles in the nucleolus | Nucleolar protein of unknown function implicated in ribosome biogenesis; TMA23 may be a fungal-specific gene as no homologs have been yet identified in higher eukaryotes | Protein of unknown function that associates with ribosomes | Putative protein of unknown function; may play a role in the ribosome and rRNA biosynthesis based on expression profiles and mutant phenotype | Nucleolar protein; involved in biogenesis of the 60S subunit of the ribosome; interacts with rRNA processing factors Cbf5p and Nop2p; null mutant is viable but growth is severely impaired | Nucleolar protein, constituent of pre-60S ribosomal particles; required for processing of the 27S pre-rRNA at the A2 site to yield 5.8S and 25S rRNA | Cytoplasmic pre-60S factor; required for the correct recycling of shuttling factors Alb1, Arx1 and Tif6 at the end of the ribosomal large subunit biogenesis; involved in bud growth in the mitotic signaling network | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YBR271W is not as essential gene | Protein involved in nucleolar integrity and processing of the pre-rRNA for the 60S ribosome subunit; transcript is induced in response to cytotoxic stress but not genotoxic stress |
| 113 | 26 | 432 | -1.384 | 0.460 |  |  |  |  |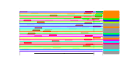  | YBL002W YBL003C YBL032W YBR009C YBR010W YBR162C YBR187W YBR283C YDL055C YDR224C YDR225W YDR245W YGL225W YJL158C YMR215W YMR305C YMR307W YNL030W YNL031C YOR247W YOR248W YPL163C YDL084W YGL103W YLR180W YNL010W | HTB2 HTA2 HEK2 HHF1 HHT1 TOS1 GDT1 SSH1 PSA1 HTB1 HTA1 MNN10 VRG4 CIS3 GAS3 SCW10 GAS1 HHF2 HHT2 SRL1 SVS1 SUB2 RPL28 SAM1 | One of two nearly identical (see HTB1) histone H2B subtypes required for chromatin assembly and chromosome function; Rad6p-Bre1p-Lge1p mediated ubiquitination regulates transcriptional activation, meiotic DSB formation and H3 methylation | One of two nearly identical (see also HTA1) histone H2A subtypes; core histone required for chromatin assembly and chromosome function; DNA damage-dependent phosphorylation by Mec1p facilitates DNA repair; acetylated by Nat4p | RNA binding protein with similarity to hnRNP-K that localizes to the cytoplasm and to subtelomeric DNA; required for the proper localization of ASH1 mRNA; involved in the regulation of telomere position effect and telomere length | One of two identical histone H4 proteins (see also HHF2); core histone required for chromatin assembly and chromosome function; contributes to telomeric silencing; N-terminal domain involved in maintaining genomic integrity | One of two identical histone H3 proteins (see also HHT2); core histone required for chromatin assembly, involved in heterochromatin-mediated telomeric and HM silencing; regulated by acetylation, methylation, and mitotic phosphorylation | Covalently-bound cell wall protein of unknown function; identified as a cell cycle regulated SBF target gene; deletion mutants are highly resistant to treatment with beta-1,3-glucanase; has sequence similarity to YJL171C | Putative protein of unknown function; expression is reduced in a gcr1 null mutant; GFP-fusion protein localizes to the vacuole; expression pattern and physical interactions suggest a possible role in ribosome biogenesis | Subunit of the Ssh1 translocon complex; Sec61p homolog involved in co-translational pathway of protein translocation; not essential | GDP-mannose pyrophosphorylase (mannose-1-phosphate guanyltransferase), synthesizes GDP-mannose from GTP and mannose-1-phosphate in cell wall biosynthesis; required for normal cell wall structure | One of two nearly identical (see HTB2) histone H2B subtypes required for chromatin assembly and chromosome function; Rad6p-Bre1p-Lge1p mediated ubiquitination regulates transcriptional activation, meiotic DSB formation and H3 methylation | One of two nearly identical (see also HTA2) histone H2A subtypes; core histone required for chromatin assembly and chromosome function; DNA damage-dependent phosphorylation by Mec1p facilitates DNA repair; acetylated by Nat4p | Subunit of a Golgi mannosyltransferase complex also containing Anp1p, Mnn9p, Mnn11p, and Hoc1p that mediates elongation of the polysaccharide mannan backbone; membrane protein of the mannosyltransferase family | Golgi GDP-mannose transporter; regulates Golgi function and glycosylation in Golgi | Mannose-containing glycoprotein constituent of the cell wall; member of the PIR (proteins with internal repeats) family | Putative 1,3-beta-glucanosyltransferase, has similarity to Gas1p; localizes to the cell wall | Cell wall protein with similarity to glucanases; may play a role in conjugation during mating based on mutant phenotype and its regulation by Ste12p | Beta-1,3-glucanosyltransferase, required for cell wall assembly; localizes to the cell surface via a glycosylphosphatidylinositol (GPI) anchor | One of two identical histone H4 proteins (see also HHF1); core histone required for chromatin assembly and chromosome function; contributes to telomeric silencing; N-terminal domain involved in maintaining genomic integrity | One of two identical histone H3 proteins (see also HHT1); core histone required for chromatin assembly, involved in heterochromatin-mediated telomeric and HM silencing; regulated by acetylation, methylation, and mitotic phosphorylation | Mannoprotein that exhibits a tight association with the cell wall, required for cell wall stability in the absence of GPI-anchored mannoproteins; has a high serine-threonine content; expression is induced in cell wall mutants | Cell wall and vacuolar protein, required for wild-type resistance to vanadate | Component of the TREX complex required for nuclear mRNA export; member of the DEAD-box RNA helicase superfamily and is involved in early and late steps of spliceosome assembly; homolog of the human splicing factor hUAP56 | Ribosomal protein of the large (60S) ribosomal subunit, has similarity to E. coli L15 and rat L27a ribosomal proteins; may have peptidyl transferase activity; can mutate to cycloheximide resistance | S-adenosylmethionine synthetase, catalyzes transfer of the adenosyl group of ATP to the sulfur atom of methionine; one of two differentially regulated isozymes (Sam1p and Sam2p) | Putative protein of unknown function with similarity to phosphoserine phosphatases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; homozygous diploid mutant shows an increase in glycogen accumulation |
| 16 | 15 | 451 | -1.384 | 0.393 | 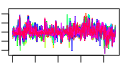 |  | 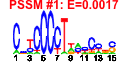 |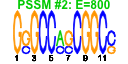  |  | YAL034C YBR169C YBR230C YBR241C YDL222C YDR204W YDR231C YFR017C YGR248W YHR087W YIL136W YKL151C YPL247C YDL110C YLR177W | FUN19 SSE2 OM14 FMP45 COQ4 COX20 SOL4 OM45 TMA17 | Non-essential protein of unknown function | Member of the heat shock protein 70 (HSP70) family; may be involved in protein folding; localized to the cytoplasm; highly homologous to the heat shock protein Sse1p | Integral mitochondrial outer membrane protein; abundance is decreased in cells grown in glucose relative to other carbon sources; appears to contain 3 alpha-helical transmembrane segments; ORF encodes a 97-basepair intron | Putative transporter, member of the sugar porter family; green fluorescent protein (GFP)-fusion protein localizes to the vacuolar membrane; YBR241C is not an essential gene | Integral membrane protein localized to mitochondria (untagged protein) and eisosomes, immobile patches at the cortex associated with endocytosis; sporulation and sphingolipid content are altered in mutants; has homologs SUR7 and YNL194C | Protein with a role in ubiquinone (Coenzyme Q) biosynthesis, possibly functioning in stabilization of Coq7p; located on the matrix face of the mitochondrial inner membrane; component of a mitochondrial ubiquinone-synthesizing complex | Mitochondrial inner membrane protein, required for proteolytic processing of Cox2p and its assembly into cytochrome c oxidase | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and is induced in response to the DNA-damaging agent MMS; YFR017C is not an essential gene | 6-phosphogluconolactonase with similarity to Sol3p | Protein of unknown function involved in RNA metabolism; this single domain protein has structural similarity to the N-terminal domain of SBDS, the human protein mutated in Shwachman-Diamond Syndrome (the yeast SBDS ortholog = SDO1/YLR022C) | Protein of unknown function, major constituent of the mitochondrial outer membrane; located on the outer (cytosolic) face of the outer membrane | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; similar to the petunia WD repeat protein an11; YPL247C is not an essential gene | Protein of unknown function that associates with ribosomes | Putative protein of unknown function; phosphorylated by Dbf2p-Mob1p in vitro; some strains contain microsatellite polymophisms at this locus; YLR177W is not an essential gene |
| 253 | 46 | 456 | -1.342 | 0.590 | 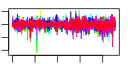 |  | 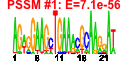 | 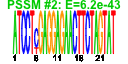 |  | YDR461W YGR259C YLR228C YLR440C YNL040W YNL095C YPR035W YAR009C YAR010C YBL005W YBR012C YCL019W YCL020W YCR018C YDR264C YDR340W YDR366C YER064C YER138C YER160C YGL204C YGR180C YGR260W YHR145C YIL060W YJR026W YJR027W YJR028W YJR029W YKL043W YLR023C YLR138W YLR237W YLR331C YLR463C YML039W YML040W YML045W YML113W YMR045C YMR046C YMR050C YMR051C YOL106W YOR153W YPR197C | MFA1 ECM22 SEC39 GLN1 PDR3 SRD1 AKR1 RNR4 TNA1 PHD1 IZH3 NHA1 THI7 DAT1 NCB2 | Mating pheromone a-factor, made by a cells; interacts with alpha cells to induce cell cycle arrest and other responses leading to mating; biogenesis involves C-terminal modification, N-terminal proteolysis, and export; also encoded by MFA2 | Sterol regulatory element binding protein, regulates transcription of the sterol biosynthetic genes ERG2 and ERG3; member of the fungus-specific Zn[2]-Cys[6] binuclear cluster family of transcription factors; homologous to Upc2p | Protein of unknown function proposed to be involved in protein secretion; interacts with Dsl1p and localizes to the ER and nuclear envelope | Putative protein of unknown function with strong similarity to alanyl-tRNA synthases from Eubacteria; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YNL040W is not an essential gene | Putative protein of unknown function predicted to contain a transmembrane domain; YNL095C is not an essential gene | Glutamine synthetase (GS), synthesizes glutamine from glutamate and ammonia; with Glt1p, forms the secondary pathway for glutamate biosynthesis from ammonia; expression regulated by nitrogen source and by amino acid limitation | Retrotransposon TYA Gag and TYB Pol genes; in YARCTY1-1 TYB is mutant and probably non-functional | Retrotransposon TYA Gag gene co-transcribed with TYB Pol; in YARCTY1-1 TYB is mutant and probably non-functional | Transcriptional activator of the pleiotropic drug resistance network, regulates expression of ATP-binding cassette (ABC) transporters through binding to cis-acting sites known as PDREs (PDR responsive elements) | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Retrotransposon TYA Gag gene co-transcribed with TYB Pol; translated as TYA or TYA-TYB polyprotein; Gag is a nucleocapsid protein that is the structural constituent of virus-like particles (VLPs); similar to retroviral Gag | Protein involved in the processing of pre-rRNA to mature rRNA; contains a C2/C2 zinc finger motif; srd1 mutation suppresses defects caused by the rrp1-1 mutation | Palmitoyl transferase involved in protein palmitoylation; acts as a negative regulator of pheromone response pathway; required for endocytosis of pheromone receptors; involved in cell shape control; contains ankyrin repeats | hypothetical protein | Non-essential nuclear protein; null mutation has global effects on transcription | Ribonucleotide-diphosphate reductase (RNR), small subunit; the RNR complex catalyzes the rate-limiting step in dNTP synthesis and is regulated by DNA replication and DNA damage checkpoint pathways via localization of the small subunits | High affinity nicotinic acid plasma membrane permease, responsible for uptake of low levels of nicotinic acid; expression of the gene increases in the absence of extracellular nicotinic acid or para-aminobenzoate (PABA) | Putative protein of unknown function; mutant accumulates less glycogen than does wild type; YIL060W is not an essential gene | Transcriptional activator that enhances pseudohyphal growth; regulates expression of FLO11, an adhesin required for pseudohyphal filament formation; similar to StuA, an A. nidulans developmental regulator; potential Cdc28p substrate | Membrane protein involved in zinc metabolism, member of the four-protein IZH family, expression induced by zinc deficiency; deletion reduces sensitivity to elevated zinc and shortens lag phase, overexpression reduces Zap1p activity | Na+/H+ antiporter involved in sodium and potassium efflux through the plasma membrane; required for alkali cation tolerance at acidic pH | Plasma membrane transporter responsible for the uptake of thiamine, member of the major facilitator superfamily of transporters; mutation of human ortholog causes thiamine-responsive megaloblastic anemia | DNA binding protein that recognizes oligo(dA).oligo(dT) tracts; Arg side chain in its N-terminal pentad Gly-Arg-Lys-Pro-Gly repeat is required for DNA-binding; not essential for viability | Subunit of a heterodimeric NC2 transcription regulator complex with Bur6p; complex binds to TBP and can repress transcription by preventing preinitiation complex assembly or stimulate activated transcription; homologous to human NC2beta |
| 127 | 22 | 436 | -1.336 | 0.450 |  |  | 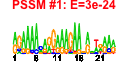 | 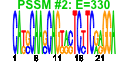 |  | YCR091W YDL025C YDL199C YGL259W YIL107C YIR039C YJR091C YKL140W YKL162C YKR018C YKR076W YOL153C YPL123C YPL166W YPR184W YGL104C YKR017C YLR080W YOL048C YOL083W YOR152C YOR386W | KIN82 YPS5 PFK26 YPS6 JSN1 TGL1 ECM4 RNY1 ATG29 GDB1 VPS73 EMP46 PHR1 | Putative serine/threonine protein kinase, most similar to cyclic nucleotide-dependent protein kinase subfamily and the protein kinase C subfamily | Putative protein kinase, potentially phosphorylated by Cdc28p; YDL025C is not an essential gene | Putative transporter, member of the sugar porter family | Protein with similarity to GPI-anchored aspartic proteases such as Yap1p and Yap3p | 6-phosphofructo-2-kinase, inhibited by phosphoenolpyruvate and sn-glycerol 3-phosphate, has negligible fructose-2,6-bisphosphatase activity, transcriptional regulation involves protein kinase A | Putative GPI-anchored aspartic protease | Member of the Puf family of RNA-binding proteins, interacts with mRNAs encoding membrane-associated proteins; overexpression suppresses a tub2-150 mutation and causes increased sensitivity to benomyl in wild-type cells | Steryl ester hydrolase, one of three gene products (Yeh1p, Yeh2p, Tgl1p) responsible for steryl ester hydrolase activity and involved in sterol homeostasis; localized to lipid particle membranes | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the mitochondrion | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Omega class glutathione transferase; not essential; similar to Ygr154cp; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | RNAse; member of the T(2) family of endoribonucleases | Protein specifically required for autophagy; may function in autophagosome formation at the pre-autophagosomal structure in collaboration with other autophagy proteins | Glycogen debranching enzyme containing glucanotranferase and alpha-1,6-amyloglucosidase activities, required for glycogen degradation; phosphorylated in mitochondria | Mitochondrial protein; mutation affects vacuolar protein sorting; putative transporter; member of the sugar porter family | Putative protein of unknown function; contains a RING finger motif | Integral membrane component of endoplasmic reticulum-derived COPII-coated vesicles, which function in ER to Golgi transport | Putative protein of unknown function | hypothetical protein | Putative protein of unknown function; has no similarity to any known protein; YOR152C is not an essential gene | DNA photolyase involved in photoreactivation, repairs pyrimidine dimers in the presence of visible light; induced by DNA damage; regulated by transcriptional repressor Rph1p |
| 175 | 28 | 440 | -1.332 | 0.447 | 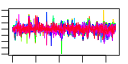 |  | 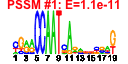 | 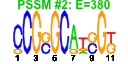 |  | YBL045C YBR039W YDL067C YDR298C YDR377W YEL024W YFR033C YGL187C YGL188C YGR183C YHR051W YJR077C YJR121W YLL041C YLR038C YLR294C YLR295C YLR395C YMR256C YNL052W YOR065W YPL078C YPL271W YPR020W YPR191W YBL099W YDL004W YLR168C | COR1 ATP3 COX9 ATP5 ATP17 NUP42 QCR6 COX4 YGL188C-A QCR9 COX6 MIR1 ATP2 SDH2 COX12 ATP14 COX8 COX7 COX5A CYT1 ATP4 ATP15 ATP20 QCR2 ATP1 ATP16 AIM30 | Core subunit of the ubiquinol-cytochrome c reductase complex (bc1 complex), which is a component of the mitochondrial inner membrane electron transport chain | Gamma subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | Subunit VIIa of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain | Subunit 5 of the stator stalk of mitochondrial F1F0 ATP synthase, which is an evolutionarily conserved enzyme complex required for ATP synthesis; homologous to bovine subunit OSCP (oligomycin sensitivity-conferring protein); phosphorylated | Subunit f of the F0 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | Subunit of the nuclear pore complex (NPC) that localizes exclusively to the cytoplasmic side; involved in RNA export, most likely at a terminal step; interacts with Gle1p | Subunit 6 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; highly acidic protein; required for maturation of cytochrome c1 | Subunit IV of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; N-terminal 25 residues of precursor are cleaved during mitochondrial import; phosphorylated | Putative protein of unknown function | Subunit 9 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; required for electron transfer at the ubiquinol oxidase site of the complex | Subunit VI of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; expression is regulated by oxygen levels | Mitochondrial phosphate carrier, imports inorganic phosphate into mitochondria; functionally redundant with Pic2p but more abundant than Pic2p under normal conditions; phosphorylated | Beta subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Iron-sulfur protein subunit of succinate dehydrogenase (Sdh1p, Sdh2p, Sdh3p, Sdh4p), which couples the oxidation of succinate to the transfer of electrons to ubiquinone | Subunit VIb of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; required for assembly of cytochrome c oxidase but not required for activity after assembly; phosphorylated | Subunit h of the F0 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | Subunit VIII of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain | Subunit VII of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain | Subunit Va of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; predominantly expressed during aerobic growth while its isoform Vb (Cox5Bp) is expressed during anaerobic growth | Cytochrome c1, component of the mitochondrial respiratory chain; expression is regulated by the heme-activated, glucose-repressed Hap2p/3p/4p/5p CCAAT-binding complex | Subunit b of the stator stalk of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Epsilon subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Subunit g of the mitochondrial F1F0 ATP synthase; reversibly phosphorylated on two residues; unphosphorylated form is required for dimerization of the ATP synthase complex | Subunit 2 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; phosphorylated; transcription is regulated by Hap1p, Hap2p/Hap3p, and heme | Alpha subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Delta subunit of the central stalk of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Putative protein of unknown function that may be involved in intramitochondrial sorting; similar to Ups1p and to human PRELI; GFP-tagged protein localizes to mitochondria; required for wild-type respiratory growth |
| 171 | 15 | 442 | -1.321 | 0.420 | 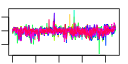 |  | 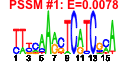 | 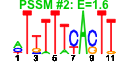 |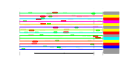  | YDL014W YHL011C YHR144C YKL181W YLR413W YNL087W YOL041C YBL024W YEL026W YGR123C YGR162W YHL039W YLR198C YNL256W YNR012W | NOP1 PRE7 DCD1 PRS1 TCB2 NOP12 NCL1 SNU13 PPT1 TIF4631 FOL1 URK1 | Nucleolar protein, component of the small subunit processome complex, which is required for processing of pre-18S rRNA; has similarity to mammalian fibrillarin | 20S proteasome beta-type subunit | Deoxycytidine monophosphate (dCMP) deaminase required for dCTP and dTTP synthesis; expression is NOT cell cycle regulated | 5-phospho-ribosyl-1(alpha)-pyrophosphate synthetase, involved in nucleotide, histidine, and tryptophan biosynthesis; one of five related enzymes, which are active as heteromultimeric complexes | Putative protein of unknown function; YLR413W is not an essential gene | Bud-specific protein with a potential role in membrane trafficking; GFP-fusion protein migrates from the cell surface to intracellular vesicles near vacuole; contains 3 calcium and lipid binding domains; mRNA is targeted to the bud | Nucleolar protein, required for pre-25S rRNA processing; contains an RNA recognition motif (RRM) and has similarity to Nop13p, Nsr1p, and putative orthologs in Drosophila and S. pombe | S-adenosyl-L-methionine-dependent tRNA: m5C-methyltransferase, methylates cytosine to m5C at several positions in tRNAs and intron-containing pre-tRNAs; similar to Nop2p and human proliferation associated nucleolar protein p120 | RNA binding protein, part of U3 snoRNP involved in rRNA processing, part of U4/U6-U5 tri-snRNP involved in mRNA splicing, similar to human 15.5K protein | Protein serine/threonine phosphatase with similarity to human phosphatase PP5; present in both the nucleus and cytoplasm; expressed during logarithmic growth; computational analyses suggest roles in phosphate metabolism and rRNA processing | Translation initiation factor eIF4G, subunit of the mRNA cap-binding protein complex (eIF4F) that also contains eIF4E (Cdc33p); associates with the poly(A)-binding protein Pab1p, also interacts with eIF4A (Tif1p); homologous to Tif4632p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Multifunctional enzyme of the folic acid biosynthesis pathway, has dihydropteroate synthetase, dihydro-6-hydroxymethylpterin pyrophosphokinase, and dihydroneopterin aldolase activities | Uridine/cytidine kinase, component of the pyrimidine ribonucleotide salvage pathway that converts uridine into UMP and cytidine into CMP; involved in the pyrimidine deoxyribonucleotide salvage pathway, converting deoxycytidine into dCMP |
| 258 | 18 | 446 | -1.311 | 0.384 |  |  | 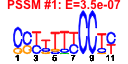 |  |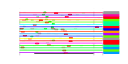  | YBR139W YLR258W YML100W YPL004C YBR052C YBR053C YBR149W YDL022W YHL021C YHR104W YIL124W YKL035W YKL103C YKL150W YMR110C YMR250W YMR297W YOL082W | GSY2 BEM2 LSP1 RFS1 ARA1 GPD1 AIM17 GRE3 AYR1 UGP1 LAP4 MCR1 HFD1 GAD1 PRC1 ATG19 | Putative serine type carboxypeptidase with a role in phytochelatin synthesis; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; expression induced by nitrogen limitation in a GLN3, GAT1-independent manner | Glycogen synthase, similar to Gsy1p; expression induced by glucose limitation, nitrogen starvation, heat shock, and stationary phase; activity regulated by cAMP-dependent, Snf1p and Pho85p kinases as well as by the Gac1p-Glc7p phosphatase | Rho GTPase activating protein (RhoGAP) involved in the control of cytoskeleton organization and cellular morphogenesis; required for bud emergence | Primary component of eisosomes, which are large immobile patch structures at the cell cortex associated with endocytosis, along with Pil1p and Sur7p; null mutants show activation of Pkc1p/Ypk1p stress resistance pathways | Protein of unknown function; member of a flavodoxin-like fold protein family that includes Pst2p and Ycp4p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Putative protein of unknown function; induced by cell wall perturbation | NADP+ dependent arabinose dehydrogenase, involved in carbohydrate metabolism; purified as homodimer; naturally occurs with a N-terminus degradation product | NAD-dependent glycerol-3-phosphate dehydrogenase, key enzyme of glycerol synthesis, essential for growth under osmotic stress; expression regulated by high-osmolarity glycerol response pathway; homolog of Gpd2p | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) | Aldose reductase involved in methylglyoxal, d-xylose and arabinose metabolism; stress induced (osmotic, ionic, oxidative, heat shock, starvation and heavy metals); regulated by the HOG pathway | NADPH-dependent 1-acyl dihydroxyacetone phosphate reductase found in lipid particles, ER, and mitochondrial outer membrane; involved in phosphatidic acid biosynthesis; required for spore germination; capable of metabolizing steroid hormones | UDP-glucose pyrophosphorylase (UGPase), catalyses the reversible formation of UDP-Glc from glucose 1-phosphate and UTP, involved in a wide variety of metabolic pathways, expression modulated by Pho85p through Pho4p | Vacuolar aminopeptidase, often used as a marker protein in studies of autophagy and cytosol to vacuole targeting (CVT) pathway | Mitochondrial NADH-cytochrome b5 reductase, involved in ergosterol biosynthesis | Putative fatty aldehyde dehydrogenase, located in the mitochondrial outer membrane and also in lipid particles; has similarity to human fatty aldehyde dehydrogenase (FALDH) which is implicated in Sjogren-Larsson syndrome | Glutamate decarboxylase, converts glutamate into gamma-aminobutyric acid (GABA) during glutamate catabolism; involved in response to oxidative stress | Vacuolar carboxypeptidase Y (proteinase C), broad-specificity C-terminal exopeptidase involved in non-specific protein degradation in the vacuole; member of the serine carboxypeptidase family | Protein involved in the cytoplasm-to-vacuole targeting pathway and in autophagy, recognizes cargo proteins and delivers them to the preautophagosomal structure for eventual engulfment by the autophagosome and degradation |
| 215 | 17 | 443 | -1.296 | 0.395 |  |  | 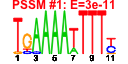 | 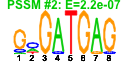 |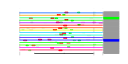  | YLR051C YMR014W YOR078W YDR152W YFR001W YLR068W YLR221C YMR239C YMR269W YNL308C YNR024W YOR056C YOR252W YOR287C YPL146C YPR142C YPR143W | FCF2 BUD22 BUD21 GIR2 LOC1 FYV7 RSA3 RNT1 TMA23 KRI1 NOB1 TMA16 NOP53 RRP15 | Essential nucleolar protein involved in the early steps of 35S rRNA processing; interacts with Faf1p; member of a transcriptionally co-regulated set of genes called the RRB regulon | Protein involved in bud-site selection; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Component of small ribosomal subunit (SSU) processosome that contains U3 snoRNA; originally isolated as bud-site selection mutant that displays a random budding pattern | Highly-acidic cytoplasmic RWD domain-containing protein of unknown function, sensitive to proteolysis, N-terminal region has high content of acidic amino acid residues, putative IUP (intrinsically unstructured protein) | Nuclear protein involved in asymmetric localization of ASH1 mRNA; binds double-stranded RNA in vitro; constituent of 66S pre-ribosomal particles | Protein of unknown function, required for survival upon exposure to K1 killer toxin; involved in processing the 35S rRNA primary transcript to generate the 20S and 27SA2 pre-rRNA transcripts | Protein with a likely role in ribosomal maturation, required for accumulation of wild-type levels of large (60S) ribosomal subunits; binds to the helicase Dbp6p in pre-60S ribosomal particles in the nucleolus | RNAase III; involved in rDNA transcription and rRNA processing; also cleaves a stem-loop structure at the 3' end of U2 snRNA to ensure formation of the correct U2 3' end | Nucleolar protein of unknown function implicated in ribosome biogenesis; TMA23 may be a fungal-specific gene as no homologs have been yet identified in higher eukaryotes | Essential nucleolar protein required for 40S ribosome biogenesis; physically and functionally interacts with Krr1p | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the nucleus | Essential nuclear protein involved in proteasome maturation and synthesis of 40S ribosomal subunits; required for cleavage of the 20S pre-rRNA to generate the mature 18S rRNA | Protein of unknown function that associates with ribosomes | Putative protein of unknown function; may play a role in the ribosome and rRNA biosynthesis based on expression profiles and mutant phenotype | Nucleolar protein; involved in biogenesis of the 60S subunit of the ribosome; interacts with rRNA processing factors Cbf5p and Nop2p; null mutant is viable but growth is severely impaired | Nucleolar protein, constituent of pre-60S ribosomal particles; required for processing of the 27S pre-rRNA at the A2 site to yield 5.8S and 25S rRNA |
| 70 | 23 | 437 | -1.274 | 0.456 |  | 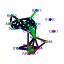 | 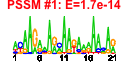 | 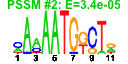 |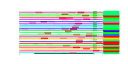  | YAL003W YBR025C YBR106W YDL192W YDR050C YER043C YGR254W YHL015W YHR128W YHR174W YIL053W YJR009C YLR044C YLR134W YLR150W YOL086C YPL131W YPL143W YPR080W YAL038W YKL060C YLR075W YMR303C | EFB1 OLA1 PHO88 ARF1 TPI1 SAH1 ENO1 YHL015W-A FUR1 ENO2 RHR2 TDH2 PDC1 PDC5 STM1 ADH1 RPL5 RPL33A TEF1 CDC19 FBA1 RPL10 ADH2 | Translation elongation factor 1 beta; stimulates nucleotide exchange to regenerate EF-1 alpha-GTP for the next elongation cycle; part of the EF-1 complex, which facilitates binding of aminoacyl-tRNA to the ribosomal A site | P-loop ATPase with similarity to human OLA1 and bacterial YchF; identified as specifically interacting with the proteasome; protein levels are induced by hydrogen peroxide | Probable membrane protein, involved in phosphate transport; pho88 pho86 double null mutant exhibits enhanced synthesis of repressible acid phosphatase at high inorganic phosphate concentrations | ADP-ribosylation factor, GTPase of the Ras superfamily involved in regulation of coated vesicle formation in intracellular trafficking within the Golgi; functionally interchangeable with Arf2p | Triose phosphate isomerase, abundant glycolytic enzyme; mRNA half-life is regulated by iron availability; transcription is controlled by activators Reb1p, Gcr1p, and Rap1p through binding sites in the 5' non-coding region | S-adenosyl-L-homocysteine hydrolase, catabolizes S-adenosyl-L-homocysteine which is formed after donation of the activated methyl group of S-adenosyl-L-methionine (AdoMet) to an acceptor | Enolase I, a phosphopyruvate hydratase that catalyzes the conversion of 2-phosphoglycerate to phosphoenolpyruvate during glycolysis and the reverse reaction during gluconeogenesis; expression is repressed in response to glucose | Putative protein of unknown function | Uracil phosphoribosyltransferase, synthesizes UMP from uracil; involved in the pyrimidine salvage pathway | Enolase II, a phosphopyruvate hydratase that catalyzes the conversion of 2-phosphoglycerate to phosphoenolpyruvate during glycolysis and the reverse reaction during gluconeogenesis; expression is induced in response to glucose | Constitutively expressed isoform of DL-glycerol-3-phosphatase; involved in glycerol biosynthesis, induced in response to both anaerobic and, along with the Hor2p/Gpp2p isoform, osmotic stress | Glyceraldehyde-3-phosphate dehydrogenase, isozyme 2, involved in glycolysis and gluconeogenesis; tetramer that catalyzes the reaction of glyceraldehyde-3-phosphate to 1,3 bis-phosphoglycerate; detected in the cytoplasm and cell-wall | Major of three pyruvate decarboxylase isozymes, key enzyme in alcoholic fermentation, decarboxylates pyruvate to acetaldehyde; subject to glucose-, ethanol-, and autoregulation; involved in amino acid catabolism | Minor isoform of pyruvate decarboxylase, key enzyme in alcoholic fermentation, decarboxylates pyruvate to acetaldehyde, regulation is glucose- and ethanol-dependent, repressed by thiamine, involved in amino acid catabolism | Protein that binds G4 quadruplex and purine motif triplex nucleic acid; acts with Cdc13p to maintain telomere structure; interacts with ribosomes and subtelomeric Y' DNA; multicopy suppressor of tom1 and pop2 mutations | Alcohol dehydrogenase, fermentative isozyme active as homo- or heterotetramers; required for the reduction of acetaldehyde to ethanol, the last step in the glycolytic pathway | Protein component of the large (60S) ribosomal subunit with similarity to E. coli L18 and rat L5 ribosomal proteins; binds 5S rRNA and is required for 60S subunit assembly | N-terminally acetylated ribosomal protein L37 of the large (60S) ribosomal subunit, nearly identical to Rpl33Bp and has similarity to rat L35a; rpl33a null mutant exhibits slow growth while rpl33a rpl33b double null mutant is inviable | Translational elongation factor EF-1 alpha; also encoded by TEF2; functions in the binding reaction of aminoacyl-tRNA (AA-tRNA) to ribosomes | Pyruvate kinase, functions as a homotetramer in glycolysis to convert phosphoenolpyruvate to pyruvate, the input for aerobic (TCA cycle) or anaerobic (glucose fermentation) respiration | Fructose 1,6-bisphosphate aldolase, required for glycolysis and gluconeogenesis; catalyzes conversion of fructose 1,6 bisphosphate to glyceraldehyde-3-P and dihydroxyacetone-P; locates to mitochondrial outer surface upon oxidative stress | Protein component of the large (60S) ribosomal subunit, responsible for joining the 40S and 60S subunits; regulates translation initiation; has similarity to rat L10 ribosomal protein and to members of the QM gene family | Glucose-repressible alcohol dehydrogenase II, catalyzes the conversion of ethanol to acetaldehyde; involved in the production of certain carboxylate esters; regulated by ADR1 |
| 88 | 22 | 444 | -1.212 | 0.515 | 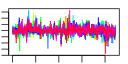 |  |  |  |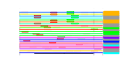  | YCR007C YDL024C YDL244W YER158C YFL058W YFL059W YFL060C YGR146C YJR156C YKL148C YLR156W YLR159W YLR161W YNL332W YOL084W YOR134W YDR059C YGL121C YGL156W YMR040W YMR271C YNL093W | DIA3 THI13 THI5 SNZ3 SNO3 THI11 SOR1 THI12 PHM7 BAG7 UBC5 GPG1 AMS1 YET2 URA10 YPT53 | Putative integral membrane protein, member of DUP240 gene family; YCR007C is not an essential gene | Protein of unknown function, involved in invasive and pseudohyphal growth | Protein involved in synthesis of the thiamine precursor hydroxymethylpyrimidine (HMP); member of a subtelomeric gene family including THI5, THI11, THI12, and THI13 | Protein of unknown function, has similarity to Afr1p; potentially phosphorylated by Cdc28p | Member of a stationary phase-induced gene family; transcription of SNZ2 is induced prior to diauxic shift, and also in the absence of thiamin in a Thi2p-dependent manner; forms a coregulated gene pair with SNO3 | Protein of unknown function, nearly identical to Sno2p; expression is induced before the diauxic shift and also in the absence of thiamin | Putative protein of unknown function; induced by iron homeostasis transcription factor Aft2p; multicopy suppressor of a temperature sensitive hsf1 mutant | Sorbitol dehydrogenase; expression is induced in the presence of sorbitol | Putative protein of unknown function; exhibits a two-hybrid interaction with Jsn1p in a large-scale analysis | Putative protein of unknown function; YLR156W, YLR159W, and YLR161W are three identical open reading frames encoded near the ribosomal DNA region of chromosome 12 | Protein of unknown function, expression is regulated by phosphate levels; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery and vacuole | Rho GTPase activating protein (RhoGAP), stimulates the intrinsic GTPase activity of Rho1p, which plays a role in actin cytoskeleton organization and control of cell wall synthesis; structurally and functionally related to Sac7p | Ubiquitin-conjugating enzyme that mediates selective degradation of short-lived and abnormal proteins, central component of the cellular stress response; expression is heat inducible | Proposed gamma subunit of the heterotrimeric G protein that interacts with the receptor Grp1p; involved in regulation of pseudohyphal growth; requires Gpb1p or Gpb2p to interact with Gpa2p | Vacuolar alpha mannosidase, involved in free oligosaccharide (fOS) degradation; delivered to the vacuole in a novel pathway separate from the secretory pathway | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; homolog of human BAP31 protein | One of two orotate phosphoribosyltransferase isozymes (see also URA5) that catalyze the fifth enzymatic step in the de novo biosynthesis of pyrimidines, converting orotate into orotidine-5'-phosphate | GTPase, similar to Ypt51p and Ypt52p and to mammalian rab5; required for vacuolar protein sorting and endocytosis |
| 67 | 20 | 456 | -1.190 | 0.429 |  |  |  | 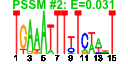 |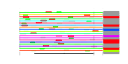  | YBR061C YDL201W YDR412W YER118C YKL021C YKR026C YKR060W YLL011W YLR003C YLR243W YLR287C YLR291C YLR449W YML080W YNL114C YOR101W YOR224C YPR187W YDL111C YNL151C | TRM7 TRM8 RRP17 SHO1 MAK11 GCN3 UTP30 SOF1 GCD7 FPR4 DUS1 RAS1 RPB8 RPO26 RRP42 RPC31 | 2'-O-ribose methyltransferase, methylates the 2'-O-ribose of nucleotides at positions 32 and 34 of the tRNA anticodon loop | Subunit of a tRNA methyltransferase complex composed of Trm8p and Trm82p that catalyzes 7-methylguanosine modification of tRNA | Protein required for cell viability | Transmembrane osmosensor, participates in activation of both the Cdc42p- and MAP kinase-dependent filamentous growth pathway and the high-osmolarity glycerol response pathway | Protein involved in an early, nucleolar step of 60S ribosomal subunit biogenesis; essential for cell growth and replication of killer M1 dsRNA virus; contains four beta-transducin repeats | Alpha subunit of the translation initiation factor eIF2B, the guanine-nucleotide exchange factor for eIF2; activity subsequently regulated by phosphorylated eIF2; first identified as a positive regulator of GCN4 expression | Possible U3 snoRNP protein involved in maturation of pre-18S rRNA, based on computational analysis of large-scale protein-protein interaction data | Essential protein required for biogenesis of 40S (small) ribosomal subunit; has similarity to the beta subunit of trimeric G-proteins and the splicing factor Prp4p | Putative protein of unknown function that may participate in DNA replication; green fluorescent protein (GFP)-fusion protein is localized to the nucleus; YLR003C is not an essential gene | Putative protein of unknown function; YLR243W is an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR287C is not an essential gene | Beta subunit of the translation initiation factor eIF2B, the guanine-nucleotide exchange factor for eIF2; activity subsequently regulated by phosphorylated eIF2; first identified as a negative regulator of GCN4 expression | Peptidyl-prolyl cis-trans isomerase (PPIase) (proline isomerase) localized to the nucleus; catalyzes isomerization of proline residues in histones H3 and H4, which affects lysine methylation of those histones | Dihydrouridine synthase, member of a widespread family of conserved proteins including Smm1p, Dus3p, and Dus4p; modifies pre-tRNA(Phe) at U17 | GTPase involved in G-protein signaling in the adenylate cyclase activating pathway, plays a role in cell proliferation; localized to the plasma membrane; homolog of mammalian RAS proto-oncogenes | RNA polymerase subunit ABC14.5, common to RNA polymerases I, II, and III | RNA polymerase subunit ABC23, common to RNA polymerases I, II, and III; part of central core; similar to bacterial omega subunit | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex with Rrp4p, Rrp41p, Rrp43p and Dis3p | RNA polymerase III subunit C31; contains HMG-like C-terminal domain |
| 265 | 27 | 437 | -1.160 | 0.455 |  |  | 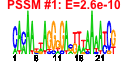 | 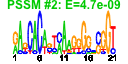 |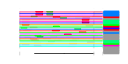  | YBL015W YKL107W YLR164W YLR284C YAL054C YBL048W YBR116C YBR117C YCR021C YDL085W YDL223C YDR256C YER039C YGR067C YGR144W YLR311C YMR118C YMR322C YMR323W YNL202W YNL237W YOR100C YOR391C YOR393W YPL185W YPL280W YPL281C | ACH1 ECI1 ACS1 TKL2 HSP30 NDE2 HBT1 CTA1 HVG1 THI4 SNO4 ERR3 SPS19 YTP1 CRC1 HSP33 ERR1 HSP32 ERR2 | Acetyl-coA hydrolase, primarily localized to mitochondria; phosphorylated; required for acetate utilization and for diploid pseudohyphal growth | Putative protein of unknown function | Mitochondrial inner membrane of unknown function; similar to Tim18p and Sdh4p; expression induced by nitrogen limitation in a GLN3, GAT1-dependent manner | Peroxisomal delta3,delta2-enoyl-CoA isomerase, hexameric protein that converts 3-hexenoyl-CoA to trans-2-hexenoyl-CoA, essential for the beta-oxidation of unsaturated fatty acids, oleate-induced | Acetyl-coA synthetase isoform which, along with Acs2p, is the nuclear source of acetyl-coA for histone acetlyation; expressed during growth on nonfermentable carbon sources and under aerobic conditions | Transketolase, similar to Tkl1p; catalyzes conversion of xylulose-5-phosphate and ribose-5-phosphate to sedoheptulose-7-phosphate and glyceraldehyde-3-phosphate in the pentose phosphate pathway; needed for synthesis of aromatic amino acids | Hydrophobic plasma membrane localized, stress-responsive protein that negatively regulates the H(+)-ATPase Pma1p; induced by heat shock, ethanol treatment, weak organic acid, glucose limitation, and entry into stationary phase | Mitochondrial external NADH dehydrogenase, catalyzes the oxidation of cytosolic NADH; Nde1p and Nde2p are involved in providing the cytosolic NADH to the mitochondrial respiratory chain | Substrate of the Hub1p ubiquitin-like protein that localizes to the shmoo tip (mating projection); mutants are defective for mating projection formation, thereby implicating Hbt1p in polarized cell morphogenesis | Catalase A, breaks down hydrogen peroxide in the peroxisomal matrix formed by acyl-CoA oxidase (Pox1p) during fatty acid beta-oxidation | Protein of unknown function, has homology to Vrg4p | Putative protein of unknown function; contains a zinc finger motif similar to that of Adr1p | Thiazole synthase, catalyzes formation of the thiazole moiety of thiamin pyrophosphate; required for thiamine biosynthesis and for mitochondrial genome stability | Protein of unknown function with similarity to succinate dehydrogenase cytochrome b subunit; YMR118C is not an essential gene | Possible chaperone and cysteine protease with similarity to E. coli Hsp31 and S. cerevisiae Hsp31p, Hsp32p, and Hsp33p; member of the DJ-1/ThiJ/PfpI superfamily; may have a role in pyridoxine metabolism | Protein of unknown function, has similarity to enolases | Peroxisomal 2,4-dienoyl-CoA reductase, auxiliary enzyme of fatty acid beta-oxidation; homodimeric enzyme required for growth and sporulation on petroselineate medium; expression induced during late sporulation and in the presence of oleate | Probable type-III integral membrane protein of unknown function, has regions of similarity to mitochondrial electron transport proteins | Mitochondrial inner membrane carnitine transporter, required for carnitine-dependent transport of acetyl-CoA from peroxisomes to mitochondria during fatty acid beta-oxidation | Possible chaperone and cysteine protease with similarity to E. coli Hsp31 and S. cerevisiae Hsp31p, Hsp32p, and Sno4p; member of the DJ-1/ThiJ/PfpI superfamily, which includes human DJ-1 involved in Parkinson's disease | Possible chaperone and cysteine protease with similarity to E. coli Hsp31 and S. cerevisiae Hsp31p, Hsp33p, and Sno4p; member of the DJ-1/ThiJ/PfpI superfamily, which includes human DJ-1 involved in Parkinson's disease |
| 135 | 14 | 446 | -1.120 | 0.423 | 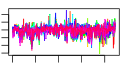 |  |  |  |  | YBR271W YJL200C YJR063W YLR073C YML018C YNL256W YNR012W YNR046W YAL036C YJL208C YKL191W YMR310C YOL124C YOR224C | ACO2 RPA12 FOL1 URK1 TRM112 RBG1 NUC1 DPH2 TRM11 RPB8 | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YBR271W is not as essential gene | Putative mitochondrial aconitase isozyme; similarity to Aco1p, an aconitase required for the TCA cycle; expression induced during growth on glucose, by amino acid starvation via Gcn4p, and repressed on ethanol | RNA polymerase I subunit A12.2; contains two zinc binding domains, and the N terminal domain is responsible for anchoring to the RNA pol I complex | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to endosomes; YLR073C is not an esssential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the membrane of the vacuole; YML018C is not an essential gene | Multifunctional enzyme of the folic acid biosynthesis pathway, has dihydropteroate synthetase, dihydro-6-hydroxymethylpterin pyrophosphokinase, and dihydroneopterin aldolase activities | Uridine/cytidine kinase, component of the pyrimidine ribonucleotide salvage pathway that converts uridine into UMP and cytidine into CMP; involved in the pyrimidine deoxyribonucleotide salvage pathway, converting deoxycytidine into dCMP | Subunit of an adoMet-dependent tRNA methyltransferase (MTase) complex (Trm11p-Trm112p), required for the methylation of the guanosine nucleotide at position 10 (m2G10) in tRNAs; putative zinc binding subunit of other MTase-related proteins | Member of the DRG family of GTP-binding proteins; interacts with translating ribosomes and with Tma46p | Major mitochondrial nuclease, has RNAse and DNA endo- and exonucleolytic activities; has roles in mitochondrial recombination, apoptosis and maintenance of polyploidy | Protein required, along with Dph1p, Kti11p, Jjj3p, and Dph5p, for synthesis of diphthamide, which is a modified histidine residue of translation elongation factor 2 (Eft1p or Eft2p); may act in a complex with Dph1p and Kti11p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; YMR310C is not an essential gene | Catalytic subunit of an adoMet-dependent tRNA methyltransferase complex (Trm11p-Trm112p), required for the methylation of the guanosine nucleotide at position 10 (m2G10) in tRNAs; contains a THUMP domain and a methyltransferase domain | RNA polymerase subunit ABC14.5, common to RNA polymerases I, II, and III |
| 141 | 37 | 437 | -1.055 | 0.546 | 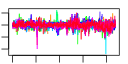 |  |  |  |  | YBL102W YCR069W YFR044C YGL199C YIL042C YIL157C YLR370C YMR099C YNL070W YNL243W YOL129W YOR089C YOR176W YPR023C YPR036W YBR222C YBR302C YDL237W YDL248W YDR105C YER130C YER178W YFL018C YFL062W YGR080W YGR132C YGR295C YHR033W YIL088C YIR044C YJR161C YLR079W YML132W YNL336W YOL155C YOR245C YPR028W | SFT2 CPR4 DUG1 PKP1 COA1 ARC18 TOM7 SLA2 VPS68 VPS21 HEM15 EAF3 VMA13 PCS60 COS2 LRC1 COS7 TMS1 PDA1 LPD1 COS4 TWF1 PHB1 COS6 AVT7 COS5 SIC1 COS3 COS1 HPF1 DGA1 YOP1 | Non-essential tetra-spanning membrane protein found mostly in the late Golgi, can suppress some sed5 alleles; may be part of the transport machinery, but precise function is unknown; similar to mammalian syntaxin 5 | Peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; has a potential role in the secretory pathway | Probable di- and tri-peptidase; forms a complex with Dug2p and Dug3p to degrade glutathione (GSH) and other peptides containing a gamma-glu-X bond in an alternative pathway to GSH degradation by gamma-glutamyl transpeptidase (Ecm38p) | Mitochondrial kinase, phosphorylates pyruvate dehydrogenase alpha subunit Pda1p | Mitochondrial inner membrane protein required for assembly of the cytochrome c oxidase complex (complex IV); interacts with complex IV assembly factor Shy1p during the early stages of assembly | Subunit of the ARP2/3 complex, which is required for the motility and integrity of cortical actin patches | Glucose-6-phosphate 1-epimerase (hexose-6-phosphate mutarotase), likely involved in carbohydrate metabolism; GFP-fusion protein localizes to both the nucleus and cytoplasm and is induced in response to the DNA-damaging agent MMS | Component of the TOM (translocase of outer membrane) complex responsible for recognition and initial import steps for all mitochondrially directed proteins; promotes assembly and stability of the TOM complex | Transmembrane actin-binding protein involved in membrane cytoskeleton assembly and cell polarization; adaptor protein that links actin to clathrin and endocytosis; present in the actin cortical patch of the emerging bud tip; dimer in vivo | Vacuolar membrane protein of unknown function involved in vacuolar protein sorting; also detected in the mitochondria | GTPase required for transport during endocytosis and for correct sorting of vacuolar hydrolases; localized in endocytic intermediates; detected in mitochondria; geranylgeranylation required for membrane association; mammalian Rab5 homolog | Ferrochelatase, a mitochondrial inner membrane protein, catalyzes the insertion of ferrous iron into protoporphyrin IX, the eighth and final step in the heme biosynthetic pathway | Esa1p-associated factor, nonessential component of the NuA4 acetyltransferase complex, homologous to Drosophila dosage compensation protein MSL3 | Subunit H of the eight-subunit V1 peripheral membrane domain of the vacuolar H+-ATPase (V-ATPase), an electrogenic proton pump found throughout the endomembrane system; serves as an activator or a structural stabilizer of the V-ATPase | Peroxisomal AMP-binding protein, localizes to both the peroxisomal peripheral membrane and matrix, expression is highly inducible by oleic acid, similar to E. coli long chain acyl-CoA synthetase | Protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Putative protein of unknown function; YDL237W is not an essential gene; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Vacuolar membrane protein of unknown function that is conserved in mammals; predicted to contain eleven transmembrane helices; interacts with Pdr5p, a protein involved in multidrug resistance | hypothetical protein | E1 alpha subunit of the pyruvate dehydrogenase (PDH) complex, catalyzes the direct oxidative decarboxylation of pyruvate to acetyl-CoA; phosphorylated; regulated by glucose | Dihydrolipoamide dehydrogenase, the lipoamide dehydrogenase component (E3) of the pyruvate dehydrogenase and 2-oxoglutarate dehydrogenase multi-enzyme complexes | Twinfilin, highly conserved actin monomer-sequestering protein involved in regulation of the cortical actin cytoskeleton, composed of two cofilin-like regions, localizes actin monomers to sites of rapid filament assembly | Subunit of the prohibitin complex (Phb1p-Phb2p), a 1.2 MDa ring-shaped inner mitochondrial membrane chaperone that stabilizes newly synthesized proteins; determinant of replicative life span; involved in mitochondrial segregation | Putative protein of unknown function; epitope-tagged protein localizes to the cytoplasm | Putative transporter, member of a family of seven S. cerevisiae genes (AVT1-7) related to vesicular GABA-glycine transporters | Protein of unknown function, member the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Inhibitor of Cdc28-Clb kinase complexes that controls G1/S phase transition, preventing premature S phase and ensuring genomic integrity; phosphorylation targets Sic1p for SCF(CDC4)-dependent turnover; functional homolog of mammalian Kip1 | Protein involved in salt resistance; interacts with sodium:hydrogen antiporter Nha1p; member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Haze-protective mannoprotein that reduces the particle size of aggregated proteins in white wines | Diacylglycerol acyltransferase, catalyzes the terminal step of triacylglycerol (TAG) formation, acylates diacylglycerol using acyl-CoA as an acyl donor, localized to lipid particles | Membrane protein that interacts with Yip1p to mediate membrane traffic; overexpression results in cell death and accumulation of internal cell membranes; regulates vesicular traffic in stressed cells |
| 231 | 23 | 443 | -1.036 | 0.430 |  |  |  |  |  | YGL012W YNR043W YAL035W YBL077W YBR106W YBR121C YBR249C YCR053W YDR037W YFL022C YGL022W YGL148W YGL202W YGR185C YGR285C YHR025W YHR208W YJR007W YLR060W YLR150W YML008C YMR146C YPR033C | ERG4 MVD1 FUN12 PHO88 GRS1 ARO4 THR4 KRS1 FRS2 STT3 ARO2 ARO8 TYS1 ZUO1 THR1 BAT1 SUI2 FRS1 STM1 ERG6 TIF34 HTS1 | C-24(28) sterol reductase, catalyzes the final step in ergosterol biosynthesis; mutants are viable, but lack ergosterol | Mevalonate pyrophosphate decarboxylase, essential enzyme involved in the biosynthesis of isoprenoids and sterols, including ergosterol; acts as a homodimer | GTPase, required for general translation initiation by promoting Met-tRNAiMet binding to ribosomes and ribosomal subunit joining; homolog of bacterial IF2 | Probable membrane protein, involved in phosphate transport; pho88 pho86 double null mutant exhibits enhanced synthesis of repressible acid phosphatase at high inorganic phosphate concentrations | Cytoplasmic and mitochondrial glycyl-tRNA synthase that ligates glycine to the cognate anticodon bearing tRNA; transcription termination factor that may interact with the 3'-end of pre-mRNA to promote 3'-end formation | 3-deoxy-D-arabino-heptulosonate-7-phosphate (DAHP) synthase, catalyzes the first step in aromatic amino acid biosynthesis and is feedback-inhibited by tyrosine or high concentrations of phenylalanine or tryptophan | Threonine synthase, conserved protein that catalyzes formation of threonine from 0-phosphohomoserine; expression is regulated by the GCN4-mediated general amino acid control pathway | Lysyl-tRNA synthetase; also identified as a negative regulator of general control of amino acid biosynthesis | Alpha subunit of cytoplasmic phenylalanyl-tRNA synthetase, forms a tetramer with Frs1p to form active enzyme; evolutionarily distant from mitochondrial phenylalanyl-tRNA synthetase based on protein sequence, but substrate binding is similar | Subunit of the oligosaccharyltransferase complex of the ER lumen, which catalyzes asparagine-linked glycosylation of newly synthesized proteins; forms a subcomplex with Ost3p and Ost4p and is directly involved in catalysis | Bifunctional chorismate synthase and flavin reductase, catalyzes the conversion of 5-enolpyruvylshikimate 3-phosphate (EPSP) to form chorismate, which is a precursor to aromatic amino acids | Aromatic aminotransferase I, expression is regulated by general control of amino acid biosynthesis | Cytoplasmic tyrosyl-tRNA synthetase, class I aminoacyl-tRNA synthetase that aminoacylates tRNA(Tyr), required for cytoplasmic protein synthesis, interacts with positions 34 and 35 of the anticodon of tRNATyr | Cytosolic ribosome-associated chaperone that acts, together with Ssz1p and the Ssb proteins, as a chaperone for nascent polypeptide chains; contains a DnaJ domain and functions as a J-protein partner for Ssb1p and Ssb2p | Homoserine kinase, conserved protein required for threonine biosynthesis; expression is regulated by the GCN4-mediated general amino acid control pathway | Mitochondrial branched-chain amino acid aminotransferase, homolog of murine ECA39; highly expressed during logarithmic phase and repressed during stationary phase | Alpha subunit of the translation initiation factor eIF2, involved in the identification of the start codon; phosphorylation of Ser51 is required for regulation of translation by inhibiting the exchange of GDP for GTP | Beta subunit of cytoplasmic phenylalanyl-tRNA synthetase, forms a tetramer with Frs2p to generate active enzyme; sequence is evolutionarily distant from mitochondrial phenylalanyl-tRNA synthetase (Msf1p), but substrate binding is similar | Protein that binds G4 quadruplex and purine motif triplex nucleic acid; acts with Cdc13p to maintain telomere structure; interacts with ribosomes and subtelomeric Y' DNA; multicopy suppressor of tom1 and pop2 mutations | Delta(24)-sterol C-methyltransferase, converts zymosterol to fecosterol in the ergosterol biosynthetic pathway by methylating position C-24; localized to both lipid particles and mitochondrial outer membrane | Subunit of the core complex of translation initiation factor 3(eIF3), which is essential for translation | Cytoplasmic and mitochondrial histidine tRNA synthetase; encoded by a single nuclear gene that specifies two messages; efficient mitochondrial localization requires both a presequence and an amino-terminal sequence |
| 7 | 28 | 444 | -0.999 | 0.437 |  |  | 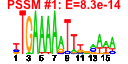 | 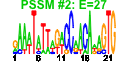 |  | YBR143C YBR154C YDL219W YDL236W YDR023W YGL070C YGL097W YGL105W YGL245W YGL246C YHL013C YHR013C YJR064W YJR069C YLR285W YLR293C YMR235C YMR260C YOL139C YOR046C YOR051C YOR253W YOR276W YOR277C YPL243W YPL273W YPR060C YPR118W | SUP45 RPB5 DTD1 PHO13 SES1 RPB9 SRM1 ARC1 GUS1 RAI1 OTU2 ARD1 CCT5 HAM1 NNT1 GSP1 RNA1 TIF11 CDC33 DBP5 NAT5 CAF20 SRP68 SAM4 ARO7 MRI1 | Polypeptide release factor involved in translation termination; mutant form acts as a recessive omnipotent suppressor | RNA polymerase subunit ABC27, common to RNA polymerases I, II, and III; contacts DNA and affects transactivation | D-Tyr-tRNA(Tyr) deacylase, functions in protein translation, may affect nonsense suppression via alteration of the protein synthesis machinery; ubiquitous among eukaryotes | Alkaline phosphatase specific for p-nitrophenyl phosphate, involved in dephosphorylation of histone II-A and casein | Cytosolic seryl-tRNA synthetase, class II aminoacyl-tRNA synthetase that aminoacylates tRNA(Ser), displays tRNA-dependent amino acid recognition which enhances discrimination of the serine substrate, interacts with peroxin Pex21p | RNA polymerase II subunit B12.6; contacts DNA; mutations affect transcription start site; involved in telomere maintenance | Nucleotide exchange factor for Gsp1p, localizes to the nucleus, required for nucleocytoplasmic trafficking of macromolecules; suppressor of the pheromone response pathway; potentially phosphorylated by Cdc28p | Protein that binds tRNA and methionyl- and glutamyl-tRNA synthetases (Mes1p and Gus1p), delivering tRNA to them, stimulating catalysis, and ensuring their localization to the cytoplasm; also binds quadruplex nucleic acids | Glutamyl-tRNA synthetase (GluRS), forms a complex with methionyl-tRNA synthetase (Mes1p) and Arc1p; complex formation increases the catalytic efficiency of both tRNA synthetases and ensures their correct localization to the cytoplasm | Nuclear protein that binds to and stabilizes the exoribonuclease Rat1p, required for pre-rRNA processing | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; member of the ovarian tumor-like (OTU) superfamily of predicted cysteine proteases; shows cytoplasmic localization | Subunit of the N-terminal acetyltransferase NatA (Nat1p, Ard1p, Nat5p); N-terminally acetylates many proteins, which influences multiple processes such as the cell cycle, heat-shock resistance, mating, sporulation, and telomeric silencing | Subunit of the cytosolic chaperonin Cct ring complex, related to Tcp1p, required for the assembly of actin and tubulins in vivo | Conserved protein with deoxyribonucleoside triphosphate pyrophosphohydrolase activity, mediates exclusion of noncanonical purines from deoxyribonucleoside triphosphate pools; mutant is sensitive to the base analog 6-N-hydroxylaminopurine | Putative nicotinamide N-methyltransferase, has a role in rDNA silencing and in lifespan determination | GTP binding protein (mammalian Ranp homolog) involved in the maintenance of nuclear organization, RNA processing and transport; regulated by Prp20p, Rna1p, Yrb1p, Yrb2p, Yrp4p, Yrb30p, Cse1p and Kap95p; yeast Gsp2p homolog | GTPase activating protein (GAP) for Gsp1p, involved in nuclear transport | Translation initiation factor eIF1A, essential protein that forms a complex with Sui1p (eIF1) and the 40S ribosomal subunit and scans for the start codon; C-terminus associates with Fun12p (eIF5B); N terminus interacts with eIF2 and eIF3 | Cytoplasmic mRNA cap binding protein; the eIF4E-cap complex is responsible for mediating cap-dependent mRNA translation via interactions with the translation initiation factor eIF4G (Tif4631p or Tif4632p) | Cytoplasmic ATP-dependent RNA helicase of the DEAD-box family involved in mRNA export from the nucleus; involved in translation termination | Nuclear protein that inhibits replication of Brome mosaic virus in S. cerevisiae, which is a model system for studying replication of positive-strand RNA viruses in their natural hosts | Phosphoprotein of the mRNA cap-binding complex involved in translational control, repressor of cap-dependent translation initiation, competes with eIF4G for binding to eIF4E | Core component of the signal recognition particle (SRP) ribonucleoprotein (RNP) complex that functions in targeting nascent secretory proteins to the endoplasmic reticulum (ER) membrane | S-adenosylmethionine-homocysteine methyltransferase, functions along with Mht1p in the conversion of S-adenosylmethionine (AdoMet) to methionine to control the methionine/AdoMet ratio | Chorismate mutase, catalyzes the conversion of chorismate to prephenate to initiate the tyrosine/phenylalanine-specific branch of aromatic amino acid biosynthesis | Methylthioribose-1-phosphate isomerase, catalyzes the isomerization of 5-methylthioribose-1-phosphate to 5-methylthioribulose-1-phosphate in the methionine salvage pathway |
| 110 | 17 | 444 | -0.958 | 0.506 |  |  |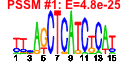  | 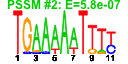 |  | YDL213C YDR365C YFL002C YGR079W YHR040W YKR025W YNR003C YNR054C YOR056C YPL212C YPR137W YDL150W YIL091C YJR041C YLL034C YML060W YOR207C | NOP6 ESF1 SPB4 BCD1 RPC37 RPC34 ESF2 NOB1 PUS1 RRP9 RPC53 URB2 RIX7 OGG1 COP1 | Putative RNA-binding protein implicated in ribosome biogenesis; contains an RNA recognition motif (RRM) and has similarity to hydrophilins; NOP6 may be a fungal-specific gene as no homologs have been yet identified in higher eukaryotes | Nucleolar protein involved in pre-rRNA processing; depletion causes severely decreased 18S rRNA levels | Putative ATP-dependent RNA helicase, nucleolar protein required for synthesis of 60S ribosomal subunits at a late step in the pathway; sediments with 66S pre-ribosomes in sucrose gradients | Putative protein of unknown function; YGR079W is not an essential gene | Essential protein required for the accumulation of box C/D snoRNA | RNA polymerase III subunit C37 | RNA polymerase III subunit C34; interacts with TFIIIB70 and is a key determinant in pol III recruitment by the preinitiation complex | Essential nucleolar protein involved in pre-18S rRNA processing; component of the small subunit (SSU) processome; has sequence similarity to mABT1, a mouse transcription activator | Essential nuclear protein involved in proteasome maturation and synthesis of 40S ribosomal subunits; required for cleavage of the 20S pre-rRNA to generate the mature 18S rRNA | tRNA:pseudouridine synthase, introduces pseudouridines at positions 26-28, 34-36, 65, and 67 of tRNA; nuclear protein that appears to be involved in tRNA export; also acts on U2 snRNA | Protein involved in pre-rRNA processing, associated with U3 snRNP; component of small ribosomal subunit (SSU) processosome; ortholog of the human U3-55k protein | RNA polymerase III subunit C53 | Protein required for cell viability | Nucleolar protein required for normal metabolism of the rRNA primary transcript, proposed to be involved in ribosome biogenesis | Putative ATPase of the AAA family, required for export of pre-ribosomal large subunits from the nucleus; distributed between the nucleolus, nucleoplasm, and nuclear periphery depending on growth conditions | Mitochondrial glycosylase/lyase that specifically excises 7,8-dihydro-8-oxoguanine residues located opposite cytosine or thymine residues in DNA, repairs oxidative damage to mitochondrial DNA | Alpha subunit of COPI vesicle coatomer complex, which surrounds transport vesicles in the early secretory pathway |
| 94 | 30 | 448 | -0.940 | 0.498 |  |  |  |  |  | YBR077C YIL036W YIL172C YJL100W YJL221C YJR021C YKL005C YML029W YMR098C YMR323W YOL044W YOL157C YOR162C YOR393W YPL281C YBR065C YBR270C YER162C YGL003C YHL024W YIL056W YIL105C YJL149W YLR189C YLR431C YOL126C YOR023C YOR380W YPL219W YPR008W | SLM4 CST6 LSB6 FSP2 REC107 BYE1 USA1 LRC4 ERR3 PEX15 YRR1 ERR1 ERR2 ECM2 BIT2 RAD4 CDH1 RIM4 VHR1 SLM1 DAS1 ATG26 ATG23 MDH2 AHC1 RDR1 PCL8 HAA1 | Component of the EGO complex, which is involved in the regulation of microautophagy, and of the GSE complex, which is required for proper sorting of amino acid permease Gap1p; gene exhibits synthetic genetic interaction with MSS4 | Basic leucine zipper (bZIP) transcription factor of the ATF/CREB family, activates transcription of genes involved in utilization of non-optimal carbon sources; involved in telomere maintenance | Putative protein of unknown function with similarity to glucosidases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Phosphatidylinositol 4-kinase that binds Las17p, which is a homolog of human Wiskott-Aldrich Syndrome protein involved in actin patch assembly and actin polymerization | Protein of unknown function, expression is induced during nitrogen limitation | Protein involved in early stages of meiotic recombination; involved in coordination between the initiation of recombination and the first division of meiosis; part of a complex (Rec107p-Mei4p-Rec114p) required for ds break formation | Negative regulator of transcription elongation, contains a TFIIS-like domain and a PHD finger, multicopy suppressor of temperature-sensitive ess1 mutations, probably binds RNA polymerase II large subunit | Protein involved in ER-associated protein degradation (ERAD); component of the Hrd1p complex; interacts with the U1 snRNP-specific protein, Snp1p | Protein of unknown function; non-tagged protein is detected in highly purified mitochondria; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Protein of unknown function, has similarity to enolases | Phosphorylated tail-anchored type II integral peroxisomal membrane protein required for peroxisome biogenesis, cells lacking Pex15p mislocalize peroxisomal matrix proteins to cytosol, overexpression results in impaired peroxisome assembly | Putative protein of unknown function | Zn2-Cys6 zinc-finger transcription factor that activates genes involved in multidrug resistance; paralog of Yrm1p, acting on an overlapping set of target genes | Pre-mRNA splicing factor, facilitates the cooperative formation of U2/U6 helix II in association with stem II in the spliceosome, function may be regulated by Slu7p | hypothetical protein | Protein that recognizes and binds damaged DNA (with Rad23p) during nucleotide excision repair; subunit of Nuclear Excision Repair Factor 2 (NEF2); homolog of human XPC protein | Cell-cycle regulated activator of the anaphase-promoting complex/cyclosome (APC/C), which directs ubiquitination of cyclins resulting in mitotic exit; targets the APC/C to specific substrates including CDC20, ASE1, CIN8 and FIN1 | Putative RNA-binding protein required for the expression of early and middle sporulation genes | Transcriptional activator, required for the vitamin H-responsive element (VHRE) mediated induction of VHT1 (Vitamin H transporter) and BIO5 (biotin biosynthesis intermediate transporter) in response to low biotin concentrations | Phosphoinositide PI4,5P(2) binding protein, forms a complex with Slm2p; acts downstream of Mss4p in a pathway regulating actin cytoskeleton organization in response to stress; subunit of and phosphorylated by the TORC2 complex | Putative SCF ubiquitin ligase F-box protein; interacts physically with both Cdc53p and Skp1 and genetically with CDC34; similar to putative F-box protein YDR131C; null mutant suppresses dst1delta sensitivity for 6-azauracil | UDP-glucose:sterol glucosyltransferase, conserved enzyme involved in synthesis of sterol glucoside membrane lipids, involved in autophagy | Peripheral membrane protein, required for autophagy and for the cytoplasm-to-vacuole targeting (Cvt) pathway | Cytoplasmic malate dehydrogenase, one of the three isozymes that catalyze interconversion of malate and oxaloacetate; involved in gluconeogenesis during growth on ethanol or acetate as carbon source; interacts with Pck1p and Fbp1p | Subunit of the Ada histone acetyltransferase complex, required for structural integrity of the complex | Transcriptional repressor involved in the control of multidrug resistance; negatively regulates expression of the PDR5 gene; member of the Gal4p family of zinc cluster proteins | Cyclin, interacts with Pho85p cyclin-dependent kinase (Cdk) to phosphorylate and regulate glycogen synthase, also activates Pho85p for Glc8p phosphorylation | Transcriptional activator involved in the transcription of TPO2, HSP30 and other genes encoding membrane stress proteins; involved in adaptation to weak acid stress |
| 164 | 19 | 447 | -0.930 | 0.502 |  |  |  |  |  | YGL211W YHR065C YJR003C YJR132W YKL014C YLR015W YMR211W YOR119C YPL245W YGL111W YGR081C YGR095C YIL079C YIL103W YIL104C YKR024C YKR060W YLR014C YNL124W | NCS6 RRP3 NMD5 URB1 BRE2 DML1 RIO1 NSA1 SLX9 RRP46 AIR1 DPH1 SHQ1 DBP7 UTP30 PPR1 NAF1 | Protein required for thiolation of the uridine at the wobble position of Gln, Lys, and Glu tRNAs; has a role in urmylation and in invasive and pseudohyphal growth; inhibits replication of Brome mosaic virus in S. cerevisiae | Protein involved in rRNA processing; required for maturation of the 35S primary transcript of pre-rRNA and for cleavage leading to mature 18S rRNA; homologous to eIF-4a, which is a DEAD box RNA-dependent ATPase with helicase activity | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Karyopherin, a carrier protein involved in nuclear import of proteins; importin beta homolog | Nucleolar protein required for the normal accumulation of 25S and 5.8S rRNAs, associated with the 27SA2 pre-ribosomal particle; proposed to be involved in the biogenesis of the 60S ribosomal subunit | Subunit of the COMPASS (Set1C) complex, which methylates histone H3 on lysine 4 and is required in transcriptional silencing near telomeres; involved in telomere maintenance; similar to trithorax-group protein ASH2L | Essential protein involved in mtDNA inheritance, may also function in the partitioning of the mitochondrial organelle or in the segregation of chromosomes, exhibits regions similar to members of a GTPase family | Essential serine kinase involved in cell cycle progression and processing of the 20S pre-rRNA into mature 18S rRNA | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the nucleus and the cytoplasm | Constituent of 66S pre-ribosomal particles, involved in 60S ribosomal subunit biogenesis | Protein required for pre-rRNA processing; associated with the 90S pre-ribosome and 43S small ribosomal subunit precursor; interacts with U3 snoRNA; deletion mutant has synthetic fitness defect with an sgs1 deletion mutant | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex | RING finger protein that interacts with the arginine methyltransferase Hmt1p to regulate methylation of Npl3p, which modulates Npl3p function in mRNA processing and export; has similarity to Air2p | Protein required, along with Dph2p, Kti11p, Jjj3p, and Dph5p, for synthesis of diphthamide, which is a modified histidine residue of translation elongation factor 2 (Eft1p or Eft2p); may act in a complex with Dph2p and Kti11p | Essential nuclear protein, required for accumulation of box H/ACA snoRNAs and for rRNA processing; interacts with Naf1p | Putative ATP-dependent RNA helicase of the DEAD-box family involved in ribosomal biogenesis | Possible U3 snoRNP protein involved in maturation of pre-18S rRNA, based on computational analysis of large-scale protein-protein interaction data | Zinc finger transcription factor containing a Zn(2)-Cys(6) binuclear cluster domain, positively regulates transcription of genes involved in uracil biosynthesis; activity may be modulated by interaction with Tup1p | Protein required for the assembly of box H/ACA snoRNPs and for pre-rRNA processing, forms a complex with Shq1p and interacts with H/ACA snoRNP components Nhp2p and Cbf5p; has similarity to Gar1p and other RNA-binding proteins |
| 255 | 30 | 447 | -0.895 | 0.438 |  |  |  |  |  | YBR154C YDR211W YDR212W YDR245W YDR331W YHR193C YIL110W YJL145W YJL183W YJR124C YKL154W YKR026C YMR033W YMR125W YMR212C YMR235C YMR260C YNL001W YNL255C YOR151C YOR168W YOR169C YOR212W YOR217W YPL050C YPL227C YPL235W YPL237W YPL238C YPR118W | RPB5 GCD6 TCP1 MNN10 GPI8 EGD2 MNI1 SFH5 MNN11 SRP102 GCN3 ARP9 SUT1 EFR3 RNA1 TIF11 DOM34 GIS2 RPB2 GLN4 STE4 RFC1 MNN9 ALG5 RVB2 SUI3 MRI1 | RNA polymerase subunit ABC27, common to RNA polymerases I, II, and III; contacts DNA and affects transactivation | Catalytic epsilon subunit of the translation initiation factor eIF2B, the guanine-nucleotide exchange factor for eIF2; activity subsequently regulated by phosphorylated eIF2; first identified as a negative regulator of GCN4 expression | Alpha subunit of chaperonin-containing T-complex, which mediates protein folding in the cytosol; involved in maintenance of actin cytoskeleton; homolog to Drosophila melanogaster and mouse tailless complex polypeptide | Subunit of a Golgi mannosyltransferase complex also containing Anp1p, Mnn9p, Mnn11p, and Hoc1p that mediates elongation of the polysaccharide mannan backbone; membrane protein of the mannosyltransferase family | ER membrane glycoprotein subunit of the glycosylphosphatidylinositol transamidase complex that adds glycosylphosphatidylinositol (GPI) anchors to newly synthesized proteins; human PIG-K protein is a functional homolog | Alpha subunit of the heteromeric nascent polypeptide-associated complex (NAC) involved in protein sorting and translocation, associated with cytoplasmic ribosomes | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; deletion mutant exhibits a weak vacuolar protein sorting defect, enhanced resistance to caspofungin, and is synthetically lethal with MEN mutants | Putative phosphatidylinositol transfer protein (PITP), exhibits phosphatidylinositol- but not phosphatidylcholine-transfer activity, localized to cytosol and microsomes, similar to Sec14p; may be PITP regulator rather than actual PITP | Subunit of a Golgi mannosyltransferase complex that also contains Anp1p, Mnn9p, Mnn10p, and Hoc1p, and mediates elongation of the polysaccharide mannan backbone; has homology to Mnn10p | Putative protein of unknown function; expression induced under calcium shortage | Signal recognition particle (SRP) receptor beta subunit; involved in SRP-dependent protein targeting; anchors Srp101p to the ER membrane | Alpha subunit of the translation initiation factor eIF2B, the guanine-nucleotide exchange factor for eIF2; activity subsequently regulated by phosphorylated eIF2; first identified as a positive regulator of GCN4 expression | Component of both the SWI/SNF and RSC chromatin remodeling complexes; actin-related protein involved in transcriptional regulation | Transcription factor of the Zn[II]2Cys6 family involved in sterol uptake; involved in induction of hypoxic gene expression | Non-essential protein of unknown function; exhibits synthetic lethal genetic interactions with PHO85; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies and is phosphorylated | GTPase activating protein (GAP) for Gsp1p, involved in nuclear transport | Translation initiation factor eIF1A, essential protein that forms a complex with Sui1p (eIF1) and the 40S ribosomal subunit and scans for the start codon; C-terminus associates with Fun12p (eIF5B); N terminus interacts with eIF2 and eIF3 | Endoribonuclease; functions in no-go mRNA decay, protein translation to promote G1 progression and differentiation, required for meiotic cell division; similar to the eukaryotic Pelota | Protein with seven cysteine-rich CCHC zinc-finger motifs, similar to human CNBP, proposed to be involved in the RAS/cAMP signaling pathway | RNA polymerase II second largest subunit B150, part of central core; similar to bacterial beta subunit | Glutamine tRNA synthetase, monomeric class I tRNA synthetase that catalyzes the specific glutaminylation of tRNA(Glu); N-terminal domain proposed to be involved in enzyme-tRNA interactions | G protein beta subunit, forms a dimer with Ste18p to activate the mating signaling pathway, forms a heterotrimer with Gpa1p and Ste18p to dampen signaling; may recruit Rho1p to the polarized growth site during mating; contains WD40 repeats | Subunit of heteropentameric Replication factor C (RF-C), which is a DNA binding protein and ATPase that acts as a clamp loader of the proliferating cell nuclear antigen (PCNA) processivity factor for DNA polymerases delta and epsilon | Subunit of Golgi mannosyltransferase complex also containing Anp1p, Mnn10p, Mnn11p, and Hoc1p that mediates elongation of the polysaccharide mannan backbone; forms a separate complex with Van1p that is also involved in backbone elongation | UDP-glucose:dolichyl-phosphate glucosyltransferase, involved in asparagine-linked glycosylation in the endoplasmic reticulum | Essential protein involved in transcription regulation; component of chromatin remodeling complexes; required for assembly and function of the INO80 complex; member of the RUVB-like protein family | Beta subunit of the translation initiation factor eIF2, involved in the identification of the start codon; proposed to be involved in mRNA binding | Methylthioribose-1-phosphate isomerase, catalyzes the isomerization of 5-methylthioribose-1-phosphate to 5-methylthioribulose-1-phosphate in the methionine salvage pathway |
| 65 | 14 | 448 | -0.809 | 0.456 |  |  |  |  |  | YDL150W YDL167C YDR060W YDR324C YDR361C YER025W YGR187C YHL039W YJL125C YLR397C YNL060C YOR091W YOR207C YER082C | RPC53 NRP1 MAK21 UTP4 BCP1 GCD11 HGH1 GCD14 AFG2 TMA46 COP1 UTP7 | RNA polymerase III subunit C53 | Protein of unknown function, rich in asparagine residues | Constituent of 66S pre-ribosomal particles, required for large (60S) ribosomal subunit biogenesis; involved in nuclear export of pre-ribosomes; required for maintenance of dsRNA virus; homolog of human CAATT-binding protein | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Essential protein involved in nuclear export of Mss4p, which is a lipid kinase that generates phosphatidylinositol 4,5-biphosphate and plays a role in actin cytoskeleton organization and vesicular transport | Gamma subunit of the translation initiation factor eIF2, involved in the identification of the start codon; binds GTP when forming the ternary complex with GTP and tRNAi-Met | Protein of unknown function with similarity to human HMG1 and HMG2; localizes to the cytoplasm | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Subunit of tRNA (1-methyladenosine) methyltransferase, with Gcd10p, required for the modification of the adenine at position 58 in tRNAs, especially tRNAi-Met; first identified as a negative regulator of GCN4 expression | ATPase of the CDC48/PAS1/SEC18 (AAA) family, forms a hexameric complex; may be involved in degradation of aberrant mRNAs | Protein of unknown function that associates with ribosomes; interacts with GTPase Rbg1p | Alpha subunit of COPI vesicle coatomer complex, which surrounds transport vesicles in the early secretory pathway |
| 35 | 25 | 435 | -0.794 | 0.428 |  |  |  |  |  | YBL049W YBR072W YBR147W YDR022C YDR262W YGL121C YGL192W YGR110W YGR256W YIL045W YIL087C YJL089W YJL185C YJR008W YJR155W YKL026C YLR311C YLR312C YMR181C YMR271C YOR031W YPL054W YPR150W YPR151C YKR009C | MOH1 HSP26 CIS1 GPG1 IME4 GND2 PIG2 LRC2 SIP4 AAD10 GPX1 QNQ1 URA10 CRS5 LEE1 SUE1 FOX2 | Protein of unknown function, has homology to kinase Snf7p; not required for growth on nonfermentable carbon sources; essential for viability in stationary phase | Small heat shock protein (sHSP) with chaperone activity; forms hollow, sphere-shaped oligomers that suppress unfolded proteins aggregation; oligomer activation requires a heat-induced conformational change; not expressed in unstressed cells | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays fluconazole resistance | Protein required for autophagosome formation in concert with Atg17p; may be involved in microtubule organization; high-copy suppressor of CIK1 deletion | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole and is induced in response to the DNA-damaging agent MMS; gene expression increases in response to Zymoliase treatment | Proposed gamma subunit of the heterotrimeric G protein that interacts with the receptor Grp1p; involved in regulation of pseudohyphal growth; requires Gpb1p or Gpb2p to interact with Gpa2p | Probable mRNA N6-adenosine methyltransferase that is required for IME1 transcript accumulation and for sporulation; expression is induced in starved MATa/MAT alpha diploid cells | Putative protein of unknown function; transcription is increased in response to genotoxic stress | 6-phosphogluconate dehydrogenase (decarboxylating), catalyzes an NADPH regenerating reaction in the pentose phosphate pathway; required for growth on D-glucono-delta-lactone | Putative type-1 protein phosphatase targeting subunit that tethers Glc7p type-1 protein phosphatase to Gsy2p glycogen synthase | Putative protein of unknown function; protein is detected in purified mitochondria in high-throughput studies; null mutant displays decreased mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | C6 zinc cluster transcriptional activator that binds to the carbon source-responsive element (CSRE) of gluconeogenic genes; involved in the positive regulation of gluconeogenesis; regulated by Snf1p protein kinase; localized to the nucleus | Putative protein of unknown function; mRNA is weakly cell cycle regulated, peaking in G2 phase; YJL185C is a non-essential gene | Putative protein of unknown function; expression repressed by inosine and choline in an Opi1p-dependent manner; expression induced by mild heat-stress on a non-fermentable carbon source. | Putative aryl-alcohol dehydrogenase with similarity to P. chrysosporium aryl-alcohol dehydrogenase; mutational analysis has not yet revealed a physiological role | Phospholipid hydroperoxide glutathione peroxidase induced by glucose starvation that protects cells from phospholipid hydroperoxides and nonphospholipid peroxides during oxidative stress | Protein of unknown function; null mutant quiescent and non-quiescent cells exhibit reduced reproductive capacity | Protein of unknown function; mRNA transcribed as part of a bicistronic transcript with a predicted transcriptional repressor RGM1/YMR182C; mRNA is destroyed by nonsense-mediated decay (NMD); YMR181C is not an essential gene | One of two orotate phosphoribosyltransferase isozymes (see also URA5) that catalyze the fifth enzymatic step in the de novo biosynthesis of pyrimidines, converting orotate into orotidine-5'-phosphate | Copper-binding metallothionein, required for wild-type copper resistance | Zinc-finger protein of unknown function | Mitochondrial protein required for degradation of unstable forms of cytochrome c | Multifunctional enzyme of the peroxisomal fatty acid beta-oxidation pathway; has 3-hydroxyacyl-CoA dehydrogenase and enoyl-CoA hydratase activities |
| 131 | 24 | 450 | -0.777 | 0.492 |  |  | 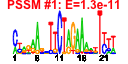 | 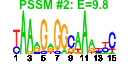 |  | YBL075C YDR287W YJL144W YBR101C YBR169C YDL020C YDR214W YDR258C YER067W YER103W YFR003C YGR111W YGR142W YHR209W YLL039C YLR216C YLR217W YLR259C YML130C YNL006W YNL007C YOL032W YOR020C YOR027W | SSA3 INM2 FES1 SSE2 RPN4 AHA1 HSP78 SSA4 YPI1 BTN2 CRG1 UBI4 CPR6 HSP60 ERO1 LST8 SIS1 OPI10 HSP10 STI1 | ATPase involved in protein folding and the response to stress; plays a role in SRP-dependent cotranslational protein-membrane targeting and translocation; member of the heat shock protein 70 (HSP70) family; localized to the cytoplasm | Inositol monophosphatase, involved in biosynthesis of inositol; enzymatic activity requires magnesium ions and is inhibited by lithium and sodium ions; inm1 inm2 double mutant lacks inositol auxotrophy | Cytoplasmic hydrophilin of unknown function, possibly involved in the dessication response; expression induced by osmotic stress, starvation and during stationary phase; GFP-fusion protein is induced by the DNA-damaging agent MMS | Hsp70 (Ssa1p) nucleotide exchange factor, cytosolic homolog of Sil1p, which is the nucleotide exchange factor for BiP (Kar2p) in the endoplasmic reticulum | Member of the heat shock protein 70 (HSP70) family; may be involved in protein folding; localized to the cytoplasm; highly homologous to the heat shock protein Sse1p | Transcription factor that stimulates expression of proteasome genes; Rpn4p levels are in turn regulated by the 26S proteasome in a negative feedback control mechanism; RPN4 is transcriptionally regulated by various stress responses | Co-chaperone that binds to Hsp82p and activates its ATPase activity; similar to Hch1p; expression is regulated by stresses such as heat shock | Oligomeric mitochondrial matrix chaperone that cooperates with Ssc1p in mitochondrial thermotolerance after heat shock; prevents the aggregation of misfolded matrix proteins; component of the mitochondrial proteolysis system | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YER067W is not an essential gene | Heat shock protein that is highly induced upon stress; plays a role in SRP-dependent cotranslational protein-membrane targeting and translocation; member of the HSP70 family; cytoplasmic protein that concentrates in nuclei upon starvation | Inhibitor of the type I protein phosphatase Glc7p, which is involved in regulation of a variety of metabolic processes; overproduction causes decreased cellular content of glycogen | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | v-SNARE binding protein that facilitates specific protein retrieval from a late endosome to the Golgi; modulates arginine uptake, possible role in mediating pH homeostasis between the vacuole and plasma membrane H(+)-ATPase | Putative S-adenosylmethionine-dependent methyltransferase; mediates cantharidin resistance | Ubiquitin, becomes conjugated to proteins, marking them for selective degradation via the ubiquitin-26S proteasome system; essential for the cellular stress response; encoded as a polyubiquitin precursor comprised of 5 head-to-tail repeats | Peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; binds to Hsp82p and contributes to chaperone activity | Tetradecameric mitochondrial chaperonin required for ATP-dependent folding of precursor polypeptides and complex assembly; prevents aggregation and mediates protein refolding after heat shock; role in mtDNA transmission; phosphorylated | Thiol oxidase required for oxidative protein folding in the endoplasmic reticulum | Protein required for the transport of amino acid permease Gap1p from the Golgi to the cell surface; component of the TOR signaling pathway; associates with both Tor1p and Tor2p; contains a WD-repeat | Type II HSP40 co-chaperone that interacts with the HSP70 protein Ssa1p; not functionally redundant with Ydj1p due to due to substrate specificity; shares similarity with bacterial DnaJ proteins | Protein with a possible role in phospholipid biosynthesis, based on inositol-excreting phenotype of the null mutant and its suppression by exogenous choline | Mitochondrial matrix co-chaperonin that inhibits the ATPase activity of Hsp60p, a mitochondrial chaperonin; involved in protein folding and sorting in the mitochondria; 10 kD heat shock protein with similarity to E. coli groES | Hsp90 cochaperone, interacts with the Ssa group of the cytosolic Hsp70 chaperones; activates the ATPase activity of Ssa1p; homolog of mammalian Hop protein |
| 300 | 29 | 442 | -0.768 | 0.465 |  |  | 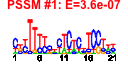 | 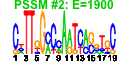 |  | YBL015W YBL045C YBR212W YDL181W YDR148C YDR178W YDR529C YER141W YGL191W YGR174C YHL032C YHR195W YIL125W YJL166W YKL016C YKL085W YKL109W YKL141W YKL148C YLL041C YML120C YMR191W YNL156C YOR065W YPL134C YPL135W YPR149W YPR155C YPR191W | ACH1 COR1 NGR1 INH1 KGD2 SDH4 QCR7 COX15 COX13 CBP4 GUT1 NVJ1 KGD1 QCR8 ATP7 MDH1 HAP4 SDH3 SOR1 SDH2 NDI1 SPG5 NSG2 CYT1 ODC1 ISU1 NCE102 NCA2 QCR2 | Acetyl-coA hydrolase, primarily localized to mitochondria; phosphorylated; required for acetate utilization and for diploid pseudohyphal growth | Core subunit of the ubiquinol-cytochrome c reductase complex (bc1 complex), which is a component of the mitochondrial inner membrane electron transport chain | RNA binding protein that negatively regulates growth rate; interacts with the 3' UTR of the mitochondrial porin (POR1) mRNA and enhances its degradation; overexpression impairs mitochondrial function; expressed in stationary phase | Protein that inhibits ATP hydrolysis by the F1F0-ATP synthase, inhibitory function is enhanced by stabilizing proteins Stf1p and Stf2p; has similarity to Stf1p and both Inh1p and Stf1p exhibit the potential to form coiled-coil structures | Dihydrolipoyl transsuccinylase, component of the mitochondrial alpha-ketoglutarate dehydrogenase complex, which catalyzes the oxidative decarboxylation of alpha-ketoglutarate to succinyl-CoA in the TCA cycle; phosphorylated | Membrane anchor subunit of succinate dehydrogenase (Sdh1p, Sdh2p, Sdh3p, Sdh4p), which couples the oxidation of succinate to the transfer of electrons to ubiquinone | Subunit 7 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; oriented facing the mitochondrial matrix; N-terminus appears to play a role in complex assembly | Protein required for the hydroxylation of heme O to form heme A, which is an essential prosthetic group for cytochrome c oxidase | Subunit VIa of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; not essential for cytochrome c oxidase activity but may modulate activity in response to ATP | Mitochondrial protein required for assembly of ubiquinol cytochrome-c reductase complex (cytochrome bc1 complex); interacts with Cbp3p and function is partially redundant with that of Cbp3p | Glycerol kinase, converts glycerol to glycerol-3-phosphate; glucose repression of expression is mediated by Adr1p and Ino2p-Ino4p; derepression of expression on non-fermentable carbon sources is mediated by Opi1p and Rsf1p | Nuclear envelope protein that interacts with the vacuolar membrane protein Vac8p to promote formation of nucleus-vacuole junctions during piecemeal microautophagy of the nucleus (PMN) | Component of the mitochondrial alpha-ketoglutarate dehydrogenase complex, which catalyzes a key step in the tricarboxylic acid (TCA) cycle, the oxidative decarboxylation of alpha-ketoglutarate to form succinyl-CoA | Subunit 8 of ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; oriented facing the intermembrane space; expression is regulated by Abf1p and Cpf1p | Subunit d of the stator stalk of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | Mitochondrial malate dehydrogenase, catalyzes interconversion of malate and oxaloacetate; involved in the tricarboxylic acid (TCA) cycle; phosphorylated | Subunit of the heme-activated, glucose-repressed Hap2p/3p/4p/5p CCAAT-binding complex, a transcriptional activator and global regulator of respiratory gene expression; provides the principal activation function of the complex | Cytochrome b subunit of succinate dehydrogenase (Sdh1p, Sdh2p, Sdh3p, Sdh4p), which couples the oxidation of succinate to the transfer of electrons to ubiquinone | Sorbitol dehydrogenase; expression is induced in the presence of sorbitol | Iron-sulfur protein subunit of succinate dehydrogenase (Sdh1p, Sdh2p, Sdh3p, Sdh4p), which couples the oxidation of succinate to the transfer of electrons to ubiquinone | NADH:ubiquinone oxidoreductase, transfers electrons from NADH to ubiquinone in the respiratory chain but does not pump protons, in contrast to the higher eukaryotic multisubunit respiratory complex I; phosphorylated; homolog of human AMID | Protein required for survival at high temperature during stationary phase; not required for growth on nonfermentable carbon sources | Protein involved in regulation of sterol biosynthesis; specifically stabilizes Hmg2p, one of two HMG-CoA isoenzymes that catalyze the rate-limiting step in sterol biosynthesis; homolog of mammalian INSIG proteins | Cytochrome c1, component of the mitochondrial respiratory chain; expression is regulated by the heme-activated, glucose-repressed Hap2p/3p/4p/5p CCAAT-binding complex | Mitochondrial inner membrane transporter, exports 2-oxoadipate and 2-oxoglutarate from the mitochondrial matrix to the cytosol for lysine and glutamate biosynthesis and lysine catabolism; suppresses, in multicopy, an fmc1 null mutation | Conserved protein of the mitochondrial matrix, performs a scaffolding function during assembly of iron-sulfur clusters, interacts physically and functionally with yeast frataxin (Yfh1p); isu1 isu2 double mutant is inviable | Protein of unknown function; contains transmembrane domains; involved in secretion of proteins that lack classical secretory signal sequences; component of the detergent-insoluble glycolipid-enriched complexes (DIGs) | Protein involved in regulation of mitochondrial expression of subunits 6 (Atp6p) and 8 (Atp8p) of the Fo-F1 ATP synthase; functions with Nca3p | Subunit 2 of the ubiquinol cytochrome-c reductase complex, which is a component of the mitochondrial inner membrane electron transport chain; phosphorylated; transcription is regulated by Hap1p, Hap2p/Hap3p, and heme |
| 211 | 27 | 446 | -0.696 | 0.499 |  |  | 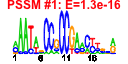 | 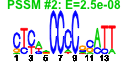 |  | YAL054C YDL085W YER121W YFL030W YGR067C YLR174W YML042W YOR032C YOR100C YPR030W YBL043W YBR051W YBR297W YFL052W YGL146C YGL250W YGL251C YGR236C YKR058W YLR377C YMR206W YNL196C YOR351C YPL017C YPL147W YPL200W YPL201C | ACS1 NDE2 AGX1 IDP2 CAT2 HMS1 CRC1 CSR2 ECM13 MAL33 RMR1 HFM1 SPG1 GLG1 FBP1 SLZ1 MEK1 IRC15 PXA1 CSM4 YIG1 | Acetyl-coA synthetase isoform which, along with Acs2p, is the nuclear source of acetyl-coA for histone acetlyation; expressed during growth on nonfermentable carbon sources and under aerobic conditions | Mitochondrial external NADH dehydrogenase, catalyzes the oxidation of cytosolic NADH; Nde1p and Nde2p are involved in providing the cytosolic NADH to the mitochondrial respiratory chain | Putative protein of unknown function; may be involved in phosphatase regulation and/or generation of precursor metabolites and energy | Alanine:glyoxylate aminotransferase (AGT), catalyzes the synthesis of glycine from glyoxylate, which is one of three pathways for glycine biosynthesis in yeast; has similarity to mammalian and plant alanine:glyoxylate aminotransferases | Putative protein of unknown function; contains a zinc finger motif similar to that of Adr1p | Cytosolic NADP-specific isocitrate dehydrogenase, catalyzes oxidation of isocitrate to alpha-ketoglutarate; levels are elevated during growth on non-fermentable carbon sources and reduced during growth on glucose | Carnitine acetyl-CoA transferase present in both mitochondria and peroxisomes, transfers activated acetyl groups to carnitine to form acetylcarnitine which can be shuttled across membranes | Basic helix-loop-helix (bHLH) protein with similarity to myc-family transcription factors; overexpression confers hyperfilamentous growth and suppresses the pseudohyphal filamentation defect of a diploid mep1 mep2 homozygous null mutant | Mitochondrial inner membrane carnitine transporter, required for carnitine-dependent transport of acetyl-CoA from peroxisomes to mitochondria during fatty acid beta-oxidation | Nuclear protein with a potential regulatory role in utilization of galactose and nonfermentable carbon sources; overproduction suppresses the lethality at high temperature of a chs5 spa2 double null mutation; potential Cdc28p substrate | Non-essential protein of unknown function | MAL-activator protein, part of complex locus MAL3; nonfunctional in genomic reference strain S288C | Putative zinc cluster protein that contains a DNA binding domain; null mutant sensitive to calcofluor white, low osmolarity and heat, suggesting a role for YFL052Wp in cell wall integrity | Putative protein of unknown function, contains two putative transmembrane spans, shows no significant homology to any other known protein sequence, YGL146C is not an essential gene | Putative protein of unknown function; deletion mutant results in reduced PIS1 expression and has growth defects on non-fermentable carbon sources and on minimal media; GFP-fusion protein localizes to both the cytoplasm and the nucleus | Meiosis specific DNA helicase involved in the conversion of double-stranded breaks to later recombination intermediates and in crossover control; catalyzes the unwinding of Holliday junctions; has ssDNA and dsDNA stimulated ATPase activity | Protein required for survival at high temperature during stationary phase; not required for growth on nonfermentable carbon sources; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Self-glucosylating initiator of glycogen synthesis, also glucosylates n-dodecyl-beta-D-maltoside; similar to mammalian glycogenin | Fructose-1,6-bisphosphatase, key regulatory enzyme in the gluconeogenesis pathway, required for glucose metabolism | Putative protein of unknown function; YMR206W is not an essential gene | Sporulation-specific protein with a leucine zipper motif | Meiosis-specific serine/threonine protein kinase, functions in meiotic checkpoint, promotes recombination between homologous chromosomes by suppressing double strand break repair between sister chromatids | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; null mutant displays increased levels of spontaneous Rad52 foci | Subunit of a heterodimeric peroxisomal ATP-binding cassette transporter complex (Pxa1p-Pxa2p), required for import of long-chain fatty acids into peroxisomes; similarity to human adrenoleukodystrophy transporter and ALD-related proteins | Protein required for accurate chromosome segregation during meiosis | Protein that interacts with glycerol 3-phosphatase and plays a role in anaerobic glycerol production; localizes to the nucleus and cytosol |
| 5 | 27 | 432 | -0.695 | 0.444 |  |  | 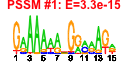 | 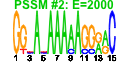 |  | YDR329C YER182W YGL098W YGL219C YGR223C YIL152W YKL034W YKL094W YKL168C YKL193C YLR201C YLR225C YLR290C YMR030W YMR139W YMR140W YNL159C YOL013C YOL065C YOR023C YOR228C YOR352W YPL003W YPL152W YPL203W YPL219W YPR172W | PEX3 FMP10 USE1 MDM34 HSV2 TUL1 YJU3 KKQ8 SDS22 COQ9 RSF1 RIM11 SIP5 ASI2 HRD1 INP54 AHC1 ULA1 YPL152W-A TPK2 PCL8 | Peroxisomal membrane protein (PMP) required required for the proper localization and stability of PMPs; interacts with Pex19p | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Essential SNARE protein localized to the ER, involved in retrograde traffic from the Golgi to the ER; forms a complex with the SNAREs Sec22p, Sec20p and Ufe1p | Mitochondrial outer membrane protein, required for normal mitochondrial morphology and inheritance; localizes to dots on the mitochondrial surface near mtDNA nucleoids; interacts genetically with MDM31 and MDM32 | Phosphatidylinositol 3,5-bisphosphate-binding protein, predicted to fold as a seven-bladed beta-propeller; displays punctate cytoplasmic localization | Putative protein of unknown function | Golgi-localized RING-finger ubiquitin ligase (E3), involved in ubiquitinating and sorting membrane proteins that contain polar transmembrane domains to multivesicular bodies for delivery to the vacuole for quality control purposes | Serine hydrolase with sequence similarity to monoglyceride lipase (MGL), localizes to lipid particles | Putative serine/threonine protein kinase with unknown cellular role | Conserved nuclear regulatory subunit of Glc7p type 1 protein serine-threonine phosphatase (PP1), functions positively with Glc7p to promote dephosphorylation of nuclear substrates required for chromosome transmission during mitosis | Protein required for ubiquinone (coenzyme Q) biosynthesis and respiratory growth; localizes to the matrix face of the mitochondrial inner membrane in a large complex with ubiquinone biosynthetic enzymes | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR225C is not an essential gene | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YLR290C is not an essential gene | Protein required for respiratory growth; localized to both the nucleus and mitochondrion; mutant displays decreased transcription of specific nuclear and mitochondrial genes whose products are involved in respiratory growth | Protein kinase required for signal transduction during entry into meiosis; promotes the formation of the Ime1p-Ume6p complex by phosphorylating Ime1p and Ume6p; shares similarity with mammalian glycogen synthase kinase 3-beta | Protein of unknown function; interacts with both the Reg1p/Glc7p phosphatase and the Snf1p kinase | Integral inner nuclear membrane protein that acts with Asi1p and Asi3p to ensure the fidelity of SPS-sensor signalling by maintaining the dormant repressed state of gene expression in the absence of inducing signals | Ubiquitin-protein ligase required for endoplasmic reticulum-associated degradation (ERAD) of misfolded proteins; genetically linked to the unfolded protein response (UPR); regulated through association with Hrd3p; contains an H2 ring finger | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase with a role in secretion, localizes to the endoplasmic reticulum via the C-terminal tail; lacks the Sac1 domain and proline-rich region found in the other 3 INP proteins | Subunit of the Ada histone acetyltransferase complex, required for structural integrity of the complex | Protein of unknown function, localized to the mitochondrial outer membrane | Putative protein of unknown function; expression levels regulated by Arg5,6p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Protein that acts together with Uba3p to activate Rub1p before its conjugation to proteins (neddylation), which may play a role in protein degradation | Identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching | cAMP-dependent protein kinase catalytic subunit; promotes vegetative growth in response to nutrients via the Ras-cAMP signaling pathway; inhibited by regulatory subunit Bcy1p in the absence of cAMP; partially redundant with Tpk1p and Tpk3p | Cyclin, interacts with Pho85p cyclin-dependent kinase (Cdk) to phosphorylate and regulate glycogen synthase, also activates Pho85p for Glc8p phosphorylation | Protein of unknown function, transcriptionally activated by Yrm1p along with genes involved in multidrug resistance |
| 55 | 17 | 444 | -0.691 | 0.484 |  |  |  |  |  | YDR449C YHR062C YIL079C YIL103W YKL144C YKL191W YLR063W YLR336C YML082W YNL124W YNL163C YNL282W YOL125W YOR001W YPR048W YPR190C YDR021W | UTP6 RPP1 AIR1 DPH1 RPC25 DPH2 SGD1 NAF1 RIA1 POP3 TRM13 RRP6 TAH18 RPC82 FAL1 | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends | RING finger protein that interacts with the arginine methyltransferase Hmt1p to regulate methylation of Npl3p, which modulates Npl3p function in mRNA processing and export; has similarity to Air2p | Protein required, along with Dph2p, Kti11p, Jjj3p, and Dph5p, for synthesis of diphthamide, which is a modified histidine residue of translation elongation factor 2 (Eft1p or Eft2p); may act in a complex with Dph2p and Kti11p | RNA polymerase III subunit C25 | Protein required, along with Dph1p, Kti11p, Jjj3p, and Dph5p, for synthesis of diphthamide, which is a modified histidine residue of translation elongation factor 2 (Eft1p or Eft2p); may act in a complex with Dph1p and Kti11p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR063W is not an essential gene | Essential nuclear protein with a possible role in the osmoregulatory glycerol response; interacts with phospholipase C (Plc1p); putative homolog of human NOM1 which is implicated in acute myeloid leukemia | Putative protein predicted to have carbon-sulfur lyase activity; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and the cytoplasm; YNML082W is not an essential gene | Protein required for the assembly of box H/ACA snoRNPs and for pre-rRNA processing, forms a complex with Shq1p and interacts with H/ACA snoRNP components Nhp2p and Cbf5p; has similarity to Gar1p and other RNA-binding proteins | Cytoplasmic GTPase involved in biogenesis of the 60S ribosome; has similarity to translation elongation factor 2 (Eft1p and Eft2p) | 2'-O-methyltransferase responsible for modification of tRNA at position 4; exhibits no obvious similarity to other known methyltransferases | Exonuclease component of the nuclear exosome; contributes to the quality-control system that retains and degrades aberrant mRNAs in the nucleus | Protein with a potential role in DNA replication; displays synthetic lethal genetic interaction with the pol3-13 allele of POL3, which encodes DNA polymerase delta | RNA polymerase III subunit C82 | Nucleolar protein required for maturation of 18S rRNA, member of the eIF4A subfamily of DEAD-box ATP-dependent RNA helicases |
| 203 | 21 | 445 | -0.628 | 0.444 |  |  |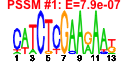  |  |  | YDR091C YJR124C YKL004W YKL216W YAR015W YBR238C YCRX16C YDL014W YER156C YGL234W YGR160W YHL011C YHR089C YHR128W YJL198W YKL181W YLR146C YML018C YML022W YOL092W YPL263C | RLI1 AUR1 URA1 ADE1 NOP1 ADE5,7 PRE7 GAR1 FUR1 PHO90 PRS1 SPE4 YLR118C KEL3 | Essential iron-sulfur protein required for ribosome biogenesis and translation initiation; facilitates binding of a multifactor complex (MFC) of translation initiation factors to the small ribosomal subunit; predicted ABC family ATPase | Putative protein of unknown function; expression induced under calcium shortage | Phosphatidylinositol:ceramide phosphoinositol transferase (IPC synthase), required for sphingolipid synthesis; can mutate to confer aureobasidin A resistance | Dihydroorotate dehydrogenase, catalyzes the fourth enzymatic step in the de novo biosynthesis of pyrimidines, converting dihydroorotic acid into orotic acid | N-succinyl-5-aminoimidazole-4-carboxamide ribotide (SAICAR) synthetase, required for 'de novo' purine nucleotide biosynthesis; red pigment accumulates in mutant cells deprived of adenine | Mitochondrial membrane protein with similarity to Rmd9p; not required for respiratory growth but causes a synthetic respiratory defect in combination with rmd9 mutations; transcriptionally up-regulated by TOR; deletion increases life span | Nucleolar protein, component of the small subunit processome complex, which is required for processing of pre-18S rRNA; has similarity to mammalian fibrillarin | Putative protein of unknown function; identified as interacting with Hsp82p in a high-throughput two-hybrid screen; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Bifunctional enzyme of the 'de novo' purine nucleotide biosynthetic pathway, contains aminoimidazole ribotide synthetase and glycinamide ribotide synthetase activities | 20S proteasome beta-type subunit | Protein component of the H/ACA snoRNP pseudouridylase complex, involved in the modification and cleavage of the 18S pre-rRNA | Uracil phosphoribosyltransferase, synthesizes UMP from uracil; involved in the pyrimidine salvage pathway | Low-affinity phosphate transporter; deletion of pho84, pho87, pho89, pho90, and pho91 causes synthetic lethality; transcription independent of Pi and Pho4p activity; overexpression results in vigorous growth | 5-phospho-ribosyl-1(alpha)-pyrophosphate synthetase, involved in nucleotide, histidine, and tryptophan biosynthesis; one of five related enzymes, which are active as heteromultimeric complexes | Spermine synthase, required for the biosynthesis of spermine and also involved in biosynthesis of pantothenic acid | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the membrane of the vacuole; YML018C is not an essential gene | Acyl-protein thioesterase responsible for depalmitoylation of Gpa1p; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and nucleus and is induced in response to the DNA-damaging agent MMS | Putative protein of unknown function; predicted to contain six transmembrane domains and is 58% similar to the uncharacterized ORF YBR147W; deletion mutant has no detectable phenotype | Cytoplasmic protein of unknown function |
| 142 | 24 | 456 | -0.625 | 0.534 |  |  | 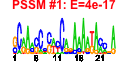 | 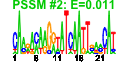 |  | YAR031W YAR033W YDL023C YDR031W YDR276C YER144C YGR209C YHR016C YKL067W YAR027W YAR028W YBR177C YDL027C YDL072C YER119C YGL053W YGL196W YGR149W YHR017W YIL062C YJR019C YJR073C YLR370C YOL129W | PRM9 MST28 MIC14 PMP3 UBP5 TRX2 YSC84 YNK1 UIP3 EHT1 YET3 AVT6 PRM8 DSD1 YSC83 ARC15 TES1 OPI3 ARC18 VPS68 | Pheromone-regulated protein with 3 predicted transmembrane segments and an FF sequence, a motif involved in COPII binding; member of DUP240 gene family | Putative integral membrane protein, involved in vesicle formation; forms complex with Mst27p; member of DUP240 gene family; binds COPI and COPII vesicles | Mitochondrial intermembrane space cysteine motif protein of 14 kDa | Small plasma membrane protein related to a family of plant polypeptides that are overexpressed under high salt concentration or low temperature, not essential for viability, deletion causes hyperpolarization of the plasma membrane potential | Putative ubiquitin-specific protease, closest paralog of Doa4p but has no functional overlap; concentrates at the bud neck | Cytoplasmic thioredoxin isoenzyme of the thioredoxin system which protects cells against oxidative and reductive stress, forms LMA1 complex with Pbi2p, acts as a cofactor for Tsa1p, required for ER-Golgi transport and vacuole inheritance | Protein involved in the organization of the actin cytoskeleton; contains SH3 domain similar to Rvs167p | Nucleoside diphosphate kinase, catalyzes the transfer of gamma phosphates from nucleoside triphosphates, usually ATP, to nucleoside diphosphates by a mechanism that involves formation of an autophosphorylated enzyme intermediate | Putative integral membrane protein of unknown function; interacts with Ulp1p at the nuclear periphery; member of DUP240 gene family | Putative integral membrane protein, member of DUP240 gene family; GFP-fusion protein is induced in response to the DNA-damaging agent MMS | Acyl-coenzymeA:ethanol O-acyltransferase that plays a minor role in medium-chain fatty acid ethyl ester biosynthesis; contains esterase activity; localizes to lipid particles and the mitochondrial outer membrane | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YDL027C is not an essential gene | Protein of unknown function; YET3 null mutant decreases the level of secreted invertase; homolog of human BAP31 protein | Vacuolar amino acid transporter, exports aspartate and glutamate from the vacuole; member of a family of seven S. cerevisiae genes (AVT1-7) related to vesicular GABA-glycine transporters | Pheromone-regulated protein with 2 predicted transmembrane segments and an FF sequence, a motif involved in COPII binding; forms a complex with Prp9p in the ER; member of DUP240 gene family | D-serine dehydratase (aka D-serine ammonia-lyase); converts D-serine to pyruvate and ammonia; specifc for D-serine, unlike the bacterial enzyme which recognizes both D-serine and L-serine as substrates | Putative protein of unknown function; predicted to be an integal membrane protein | Non-essential mitochondrial protein of unknown function; mRNA induced during meiosis, peaking between mid to late prophase of meiosis I; similar to S. douglasii YSD83 | Subunit of the ARP2/3 complex, which is required for the motility and integrity of cortical actin patches | Peroxisomal acyl-CoA thioesterase likely to be involved in fatty acid oxidation rather than fatty acid synthesis; conserved protein also found in human peroxisomes; TES1 mRNA levels increase during growth on fatty acids | Phospholipid methyltransferase (methylene-fatty-acyl-phospholipid synthase), catalyzes the last two steps in phosphatidylcholine biosynthesis | Vacuolar membrane protein of unknown function involved in vacuolar protein sorting; also detected in the mitochondria |
| 207 | 27 | 443 | -0.604 | 0.441 |  |  | 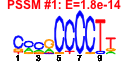 | 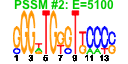 |  | YBR052C YBR214W YDR391C YDR512C YDR513W YKL103C YKR011C YLR109W YMR173W YNL134C YOL150C YOL151W YBL064C YDL124W YDR032C YDR453C YDR533C YHR008C YJR104C YKR066C YKR076W YMR090W YMR315W YNL160W YNL208W YNL274C YOR185C | RFS1 SDS24 EMI1 GRX2 LAP4 AHP1 DDR48 GRE2 PRX1 PST2 TSA2 HSP31 SOD2 SOD1 CCP1 ECM4 YGP1 GOR1 GSP2 | Protein of unknown function; member of a flavodoxin-like fold protein family that includes Pst2p and Ycp4p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | One of two S. cerevisiae homologs (Sds23p and Sds24p) of the Schizosaccharomyces pombe Sds23 protein, which genetic studies have implicated in APC/cyclosome regulation; may play an indirect role in fluid-phase endocytosis | Putative protein of unknown function, possibly involved in zinc homeostasis; Bdf1p-dependent transcription induced by salt stress; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Non-essential protein of unknown function required for transcriptional induction of the early meiotic-specific transcription factor IME1, also required for sporulation | Cytoplasmic glutaredoxin, thioltransferase, glutathione-dependent disulfide oxidoreductase involved in maintaining redox state of target proteins, also exhibits glutathione peroxidase activity, expression induced in response to stress | Vacuolar aminopeptidase, often used as a marker protein in studies of autophagy and cytosol to vacuole targeting (CVT) pathway | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus | Thiol-specific peroxiredoxin, reduces hydroperoxides to protect against oxidative damage; function in vivo requires covalent conjugation to Urm1p | DNA damage-responsive protein, expression is increased in response to heat-shock stress or treatments that produce DNA lesions; contains multiple repeats of the amino acid sequence NNNDSYGS | Putative protein of unknown function with similarity to dehydrogenases from other model organisms; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and nucleus and is induced by the DNA-damaging agent MMS | NADPH-dependent methylglyoxal reductase (D-lactaldehyde dehydrogenase); stress induced (osmotic, ionic, oxidative, heat shock and heavy metals); regulated by the HOG pathway | Mitochondrial peroxiredoxin (1-Cys Prx) with thioredoxin peroxidase activity, has a role in reduction of hydroperoxides; induced during respiratory growth and under conditions of oxidative stress; phosphorylated | NADPH-dependent alpha-keto amide reductase; reduces aromatic alpha-keto amides, aliphatic alpha-keto esters, and aromatic alpha-keto esters; member of the aldo-keto reductase (AKR) family | Protein with similarity to members of a family of flavodoxin-like proteins; induced by oxidative stress in a Yap1p dependent manner; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Stress inducible cytoplasmic thioredoxin peroxidase; cooperates with Tsa1p in the removal of reactive oxygen, nitrogen and sulfur species using thioredoxin as hydrogen donor; deletion enhances the mutator phenotype of tsa1 mutants | Possible chaperone and cysteine protease with similarity to E. coli Hsp31; member of the DJ-1/ThiJ/PfpI superfamily, which includes human DJ-1 involved in Parkinson's disease; exists as a dimer and contains a putative metal-binding site | Mitochondrial superoxide dismutase, protects cells against oxygen toxicity; phosphorylated | Cytosolic superoxide dismutase; some mutations are analogous to those that cause ALS (amyotrophic lateral sclerosis) in humans | Mitochondrial cytochrome-c peroxidase; degrades reactive oxygen species in mitochondria, involved in the response to oxidative stress | Omega class glutathione transferase; not essential; similar to Ygr154cp; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative protein of unknown function with similarity to DTDP-glucose 4,6-dehydratases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YMR090W is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YMR315W is not an essential gene | Cell wall-related secretory glycoprotein; induced by nutrient deprivation-associated growth arrest and upon entry into stationary phase; may be involved in adaptation prior to stationary phase entry; has similarity to Sps100p | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; authentic, non-tagged protein is detected in purified mitochondria in high-throughput studies; potential orthologs found in other fungi | Glyoxylate reductase; null mutation results in increased biomass after diauxic shift; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | GTP binding protein (mammalian Ranp homolog) involved in the maintenance of nuclear organization, RNA processing and transport; interacts with Kap121p, Kap123p and Pdr6p (karyophilin betas); Gsp1p homolog that is not required for viability |
| 112 | 24 | 443 | -0.576 | 0.486 |  |  | 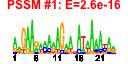 |  |  | YBR132C YBR203W YBR287W YDL233W YDL234C YDR003W YDR530C YEL039C YFR015C YGL053W YGL250W YGR268C YHR097C YJL082W YOR347C YEL060C YGL006W YGR052W YGR161C YIR016W YKL064W YKL065C YOL016C YPL247C | AGP2 COS111 GYP7 RCR2 APA2 CYC7 GSY1 PRM8 RMR1 HUA1 IML2 PYK2 PRB1 YGL006W-A FMP48 RTS3 MNR2 YET1 CMK2 | High affinity polyamine permease, preferentially uses spermidine over putrescine; expression is down-regulated by osmotic stress; plasma membrane carnitine transporter, also functions as a low-affinity amino acid permease | Protein required for resistance to the antifungal drug ciclopirox olamine; not related to the subtelomerically-encoded COS family; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the ER; YBR287W is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YDL233W is not an essential gene | GTPase-activating protein for yeast Rab family members including: Ypt7p (most effective), Ypt1p, Ypt31p, and Ypt32p (in vitro); involved in vesicle mediated protein trafficking | Vacuolar protein that presumably functions within the endosomal-vacuolar trafficking pathway, affecting events that determine whether plasma membrane proteins are degraded or routed to the plasma membrane; similar to Rcr1p | Diadenosine 5',5''-P1,P4-tetraphosphate phosphorylase II (AP4A phosphorylase), involved in catabolism of bis(5'-nucleosidyl) tetraphosphates; has similarity to Apa1p | Cytochrome c isoform 2, expressed under hypoxic conditions; electron carrier of the mitochondrial intermembrane space that transfers electrons from ubiquinone-cytochrome c oxidoreductase to cytochrome c oxidase during cellular respiration | Glycogen synthase with similarity to Gsy2p, the more highly expressed yeast homolog; expression induced by glucose limitation, nitrogen starvation, environmental stress, and entry into stationary phase | Pheromone-regulated protein with 2 predicted transmembrane segments and an FF sequence, a motif involved in COPII binding; forms a complex with Prp9p in the ER; member of DUP240 gene family | Putative protein of unknown function; deletion mutant results in reduced PIS1 expression and has growth defects on non-fermentable carbon sources and on minimal media; GFP-fusion protein localizes to both the cytoplasm and the nucleus | Cytoplasmic protein containing a zinc finger domain with sequence similarity to that of Type I J-proteins; computational analysis of large-scale protein-protein interaction data suggests a possible role in actin patch assembly | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and the nucleus | Protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Pyruvate kinase that appears to be modulated by phosphorylation; PYK2 transcription is repressed by glucose, and Pyk2p may be active under low glycolytic flux | Vacuolar proteinase B (yscB), a serine protease of the subtilisin family; involved in protein degradation in the vacuole and required for full protein degradation during sporulation | Putative protein of unknown function; identified by SAGE | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative component of the protein phosphatase type 2A complex | Putative protein of unknown function; expression directly regulated by the metabolic and meiotic transcriptional regulator Ume6p; YIR016W is a non-essential gene | Putative magnesium transporter; has similarity to Alr1p and Alr2p, which mediate influx of Mg2+ and other divalent cations | Endoplasmic reticulum transmembrane protein; may interact with ribosomes, based on co-purification experiments; homolog of human BAP31 protein | Calmodulin-dependent protein kinase; may play a role in stress response, many CA++/calmodulan dependent phosphorylation substrates demonstrated in vitro, amino acid sequence similar to Cmk1p and mammalian Cam Kinase II | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; similar to the petunia WD repeat protein an11; YPL247C is not an essential gene |
| 266 | 22 | 450 | -0.569 | 0.480 |  |  |  |  |  | YDR436W YGR250C YBR132C YBR280C YCR091W YDR085C YDR096W YDR169C YDR358W YFL016C YFL042C YFL043C YFR014C YGL059W YHR097C YHR113W YHR140W YHR171W YIL033C YLR152C YMR251W YOR228C | PPZ2 AGP2 SAF1 KIN82 AFR1 GIS1 STB3 GGA1 MDJ1 CMK1 ATG7 BCY1 GTO3 | Serine/threonine protein phosphatase Z, isoform of Ppz1p; involved in regulation of potassium transport, which affects osmotic stability, cell cycle progression, and halotolerance | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | High affinity polyamine permease, preferentially uses spermidine over putrescine; expression is down-regulated by osmotic stress; plasma membrane carnitine transporter, also functions as a low-affinity amino acid permease | F-Box protein involved in proteasome-dependent degradation of Aah1p during entry of cells into quiescence; interacts with Skp1 | Putative serine/threonine protein kinase, most similar to cyclic nucleotide-dependent protein kinase subfamily and the protein kinase C subfamily | Alpha-factor pheromone receptor regulator, negatively regulates pheromone receptor signaling; required for normal mating projection (shmoo) formation; required for Spa2p to recruit Mpk1p to shmoo tip during mating; interacts with Cdc12p | JmjC domain-containing histone demethylase; transcription factor involved in the expression of genes during nutrient limitation; also involved in the negative regulation of DPP1 and PHR1 | Protein that binds Sin3p in a two-hybrid assay | Golgi-localized protein with homology to gamma-adaptin, interacts with and regulates Arf1p and Arf2p in a GTP-dependent manner in order to facilitate traffic through the late Golgi | Protein involved in folding of mitochondrially synthesized proteins in the mitochondrial matrix; localizes to the mitochondrial inner membrane; member of the DnaJ family of molecular chaperones | Putative protein of unknown function; YFL042C is not an essential gene | Calmodulin-dependent protein kinase; may play a role in stress response, many CA++/calmodulan dependent phosphorylation substrates demonstrated in vitro, amino acid sequence similar to Cmk2p and mammalian Cam Kinase II | Protein kinase; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and the nucleus | Cytoplasmic aspartyl aminopeptidase; cleaves unblocked N-terminal acidic amino acid residues from peptide substrates; forms a 12 subunit homo-oligomeric complex; M18 metalloprotease family member; may interact with ribosomes | Putative integral membrane protein of unknown function | Autophagy-related protein and dual specificity member of the E1 family of ubiquitin-activating enzymes; mediates the conjugation of Atg12p with Atg5p and Atg8p with phosphatidylethanolamine, required steps in autophagosome formation | Regulatory subunit of the cyclic AMP-dependent protein kinase (PKA), a component of a signaling pathway that controls a variety of cellular processes, including metabolism, cell cycle, stress response, stationary phase, and sporulation | Putative protein of unknown function; YLR152C is not an essential gene | Omega class glutathione transferase; putative cytosolic localization | Protein of unknown function, localized to the mitochondrial outer membrane |
| 43 | 24 | 437 | -0.558 | 0.512 | 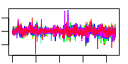 |  | 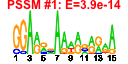 | 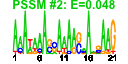 |  | YBR114W YCR011C YDR343C YDR505C YGR023W YGR288W YHL024W YKL062W YLL029W YML118W YMR008C YMR135C YMR136W YMR297W YNL037C YNL125C YNL237W YOL060C YOR113W YOR230W YPL154C YPR154W YKR067W YLL019C | RAD16 ADP1 HXT6 PSP1 MTL1 MAL13 RIM4 MSN4 RUP2 NGL3 PLB1 GID8 SCT1 PRC1 IDH1 ESBP6 YTP1 MAM3 AZF1 WTM1 PEP4 PIN3 GAT1 KNS1 | Protein that recognizes and binds damaged DNA in an ATP-dependent manner (with Rad7p) during nucleotide excision repair; subunit of Nucleotide Excision Repair Factor 4 (NEF4) and the Elongin-Cullin-Socs (ECS) ligase complex | Putative ATP-dependent permease of the ABC transporter family of proteins | High-affinity glucose transporter of the major facilitator superfamily, nearly identical to Hxt7p, expressed at high basal levels relative to other HXTs, repression of expression by high glucose requires SNF3 | Asn and gln rich protein of unknown function; high-copy suppressor of POL1 (DNA polymerase alpha) and partial suppressor of CDC2 (polymerase delta) and CDC6 (pre-RC loading factor) mutations; overexpression results in growth inhibition | Protein with both structural and functional similarity to Mid2p, which is a plasma membrane sensor required for cell integrity signaling during pheromone-induced morphogenesis; suppresses rgd1 null mutations | MAL-activator protein, part of complex locus MAL1; nonfunctional in genomic reference strain S288C | Putative RNA-binding protein required for the expression of early and middle sporulation genes | Transcriptional activator related to Msn2p; activated in stress conditions, which results in translocation from the cytoplasm to the nucleus; binds DNA at stress response elements of responsive genes, inducing gene expression | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLL029W is not an essential gene but mutant is defective in spore formation | Putative endonuclease, has a domain similar to a magnesium-dependent endonuclease motif in mRNA deadenylase Ccr4p; similar to Ngl1p and Ngl2p | Phospholipase B (lysophospholipase) involved in lipid metabolism, required for deacylation of phosphatidylcholine and phosphatidylethanolamine but not phosphatidylinositol | Protein of unknown function, involved in proteasome-dependent catabolite inactivation of fructose-1,6-bisphosphatase; contains LisH and CTLH domains, like Vid30p; dosage-dependent regulator of START | Glycerol 3-phosphate/dihydroxyacetone phosphate dual substrate-specific sn-1 acyltransferase of the glycerolipid biosynthesis pathway, prefers 16-carbon fatty acids, similar to Gpt2p, gene is constitutively transcribed | Vacuolar carboxypeptidase Y (proteinase C), broad-specificity C-terminal exopeptidase involved in non-specific protein degradation in the vacuole; member of the serine carboxypeptidase family | Subunit of mitochondrial NAD(+)-dependent isocitrate dehydrogenase, which catalyzes the oxidation of isocitrate to alpha-ketoglutarate in the TCA cycle | Protein with similarity to monocarboxylate permeases, appears not to be involved in transport of monocarboxylates such as lactate, pyruvate or acetate across the plasma membrane | Probable type-III integral membrane protein of unknown function, has regions of similarity to mitochondrial electron transport proteins | Protein required for normal mitochondrial morphology, has similarity to hemolysins | Zinc-finger transcription factor, involved in induction of CLN3 transcription in response to glucose; genetic and physical interactions indicate a possible role in mitochondrial transcription or genome maintenance | Transcriptional repressor involved in regulation of meiosis and silencing, required for nuclear localization of the ribonucleotide reductase small subunit Rnr2p and Rnr4p; contains WD repeats | Vacuolar aspartyl protease (proteinase A), required for the posttranslational precursor maturation of vacuolar proteinases; important for protein turnover after oxidative damage; synthesized as a zymogen, self-activates | Protein that induces appearance of [PIN+] prion when overproduced | Transcriptional activator of genes involved in nitrogen catabolite repression; contains a GATA-1-type zinc finger DNA-binding motif; activity and localization regulated by nitrogen limitation and Ure2p | Nonessential putative protein kinase of unknown cellular role; member of the LAMMER family of protein kinases, which are serine/threonine kinases also capable of phosphorylating tyrosine residues |
| 219 | 32 | 446 | -0.547 | 0.555 | 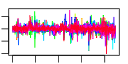 |  |  |  |  | YBR255W YDL113C YDL245C YDR124W YDR125C YFR046C YGL081W YJR159W YKL208W YNR072W YBR254C YCR011C YDL238C YDL246C YDR242W YER033C YER076C YER116C YFL059W YFL060C YGL258W YGR258C YHL016C YHR156C YIR027C YIR029W YJR127C YKL161C YLR030W YLR282C YNL333W YNL334C | ATG20 HXT15 ECM18 CNN1 SOR1 CBT1 HXT17 TRS20 ADP1 GUD1 SOR2 AMD2 ZRG8 SLX8 SNZ3 SNO3 YGL258W-A RAD2 DUR3 LIN1 DAL1 DAL2 RSF2 SNZ2 SNO2 | Protein of unknown function, required for normal growth rate at 15 degrees C; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Protein required for transport of aminopeptidase I (Lap4p) through the cytoplasm-to-vacuole targeting pathway; binds phosphatidylinositol-3-phosphate, involved in localization of membranes to the preautophagosome, potential Cdc28p substrate | Protein of unknown function with similarity to hexose transporter family members, expression is induced by low levels of glucose and repressed by high levels of glucose | Putative protein of unknown function; non-essential gene; expression is strongly induced by alpha factor | Protein of unknown function, similar to Rlp24p | Kinetochore protein of unknown function; associated with the essential kinetochore proteins Nnf1p and Spc24p; phosphorylated by both Clb5-Cdk1 and, to a lesser extent, Clb2-Cdk1. | Putative protein of unknown function; non-essential gene; interacts genetically with CHS5, a gene involved in chitin biosynthesis | Sorbitol dehydrogenase; expression is induced in the presence of sorbitol | Protein involved in 5' end processing of mitochondrial COB, 15S_rRNA, and RPM1 transcripts; may also have a role in 3' end processing of the COB pre-mRNA; displays genetic interaction with cell cycle-regulated kinase Dbf2p | Hexose transporter, up-regulated in media containing raffinose and galactose at pH 7.7 versus pH 4.7, repressed by high levels of glucose | One of 10 subunits of the transport protein particle (TRAPP) complex of the cis-Golgi which mediates vesicle docking and fusion; mutations in the human homolog cause the spondyloepiphyseal dysplasia tarda (SEDL) disorder | Putative ATP-dependent permease of the ABC transporter family of proteins | Guanine deaminase, a catabolic enzyme of the guanine salvage pathway producing xanthine and ammonia from guanine; activity is low in exponentially-growing cultures but expression is increased in post-diauxic and stationary-phase cultures | Protein of unknown function, computational analysis of large-scale protein-protein interaction data suggests a possible role in fructose or mannose metabolism | Putative amidase | Protein of unknown function; authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; GFP-fusion protein is localized to the cytoplasm; transcription induced under conditions of zinc deficiency | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; analysis of HA-tagged protein suggests a membrane localization | Subunit of the Slx5-Slx8 substrate-specific ubiquitin ligase complex; stimulated by prior attachment of SUMO to the substrate | Member of a stationary phase-induced gene family; transcription of SNZ2 is induced prior to diauxic shift, and also in the absence of thiamin in a Thi2p-dependent manner; forms a coregulated gene pair with SNO3 | Protein of unknown function, nearly identical to Sno2p; expression is induced before the diauxic shift and also in the absence of thiamin | Putative protein of unknown function | Single-stranded DNA endonuclease, cleaves single-stranded DNA during nucleotide excision repair to excise damaged DNA; subunit of Nucleotide Excision Repair Factor 3 (NEF3); homolog of human XPG protein | Plasma membrane transporter for both urea and polyamines, expression is highly sensitive to nitrogen catabolite repression and induced by allophanate, the last intermediate of the allantoin degradative pathway | Non-essential component of U5 snRNP; nuclear protein; physically interacts with Irr1p of cohesin complex; may link together proteins involved in chromosome segregation, mRNA splicing and DNA replication | Allantoinase, converts allantoin to allantoate in the first step of allantoin degradation; expression sensitive to nitrogen catabolite repression | Allantoicase, converts allantoate to urea and ureidoglycolate in the second step of allantoin degradation; expression sensitive to nitrogen catabolite repression and induced by allophanate, an intermediate in allantoin degradation | Zinc-finger protein involved in transcriptional control of both nuclear and mitochondrial genes, many of which specify products required for glycerol-based growth, respiration, and other functions | Protein kinase implicated in the Slt2p mitogen-activated (MAP) kinase signaling pathway; associates with Rlm1p | Member of a stationary phase-induced gene family; transcription of SNZ2 is induced prior to diauxic shift, and also in the absence of thiamin in a Thi2p-dependent manner; forms a coregulated gene pair with SNO2; interacts with Thi11p | Protein of unknown function, nearly identical to Sno3p; expression is induced before the diauxic shift and also in the absence of thiamin |
| 239 | 22 | 447 | -0.542 | 0.464 |  |  |  |  |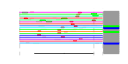  | YBL054W YBR030W YBR155W YBR266C YCL053C YCLX02C YDL063C YDR161W YDR312W YGR079W YGR280C YGR283C YKR056W YLR074C YML043C YMR014W YNL060C YNL186W YNR038W YOL022C YOR146W YPL183C | CNS1 SSF2 PXR1 TRM2 BUD20 RRN11 BUD22 UBP10 DBP6 TSR4 RTT10 | Protein of unknown function involved in rRNA and ribosome biosynthesis | Putative protein of unknown function; predicted protein contains a SET domain (S-adenosyl-L-methionine-binding fold) | TPR-containing co-chaperone; binds both Hsp82p (Hsp90) and Ssa1p (Hsp70) and stimulates the ATPase activity of SSA1, ts mutants reduce Hsp82p function while over expression suppresses the phenotypes of an HSP82 ts allele and a cpr7 deletion | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YDL063C is not an essential gene | Putative protein of unknown function; non-essential gene; proposed function in rRNA and ribosome biosynthesis based on transcriptional co-regulation | Protein required for ribosomal large subunit maturation, functionally redundant with Ssf1p; member of the Brix family | Putative protein of unknown function; YGR079W is not an essential gene | Essential protein involved in rRNA and snoRNA maturation; competes with TLC1 RNA for binding to Est2p, suggesting a role in negative regulation of telomerase; human homolog inhibits telomerase; contains a G-patch RNA interacting domain | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; deletion mutant is resistant to fluconazole; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus | tRNA methyltransferase, 5-methylates the uridine residue at position 54 of tRNAs and may also have a role in tRNA stabilization or maturation; endo-exonuclease with a role in DNA repair | Protein involved in bud-site selection; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Protein required for rDNA transcription by RNA polymerase I, component of the core factor (CF) of rDNA transcription factor, which also contains Rrn6p and Rrn7p | Ubiquitin-specific protease that deubiquitinates ubiquitin-protein moieties; may regulate silencing by acting on Sir4p; involved in posttranscriptionally regulating Gap1p and possibly other transporters; primarily located in the nucleus | Essential protein involved in ribosome biogenesis; putative ATP-dependent RNA helicase of the DEAD-box protein family | Cytoplasmic protein of unknown function; essential gene in S288C, and non-essential with reduced growth rate in CEN.PK2; Null mutant accumulates 20S pre-rRNA | Cytoplasmic protein of unknown function, plays a role in restricting Ty1 transposition |
| 285 | 27 | 444 | -0.510 | 0.498 | 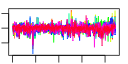 |  | 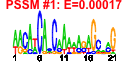 | 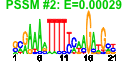 |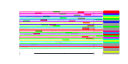  | YPL231W YAR071W YAR074C YAR075W YBL068W YBR093C YEL042W YER025W YER049W YER086W YER110C YER165W YGR094W YHR064C YHR215W YJR071W YKL004W YKL216W YLR413W YMR202W YMR246W YNL087W YNR015W YNR041C YNR043W YOL130W YPL207W | FAS2 PHO11 PRS4 PHO5 GDA1 GCD11 TPA1 ILV1 KAP123 PAB1 VAS1 SSZ1 PHO12 AUR1 URA1 ERG2 FAA4 TCB2 RSP5 COQ2 MVD1 SWC3 TYW1 | Alpha subunit of fatty acid synthetase, which catalyzes the synthesis of long-chain saturated fatty acids; contains beta-ketoacyl reductase and beta-ketoacyl synthase activities; phosphorylated | One of three repressible acid phosphatases, a glycoprotein that is transported to the cell surface by the secretory pathway; induced by phosphate starvation and coordinately regulated by PHO4 and PHO2 | 5-phospho-ribosyl-1(alpha)-pyrophosphate synthetase, involved in nucleotide, histidine, and tryptophan biosynthesis; one of five related enzymes, which are active as heteromultimeric complexes | Repressible acid phosphatase (1 of 3) that also mediates extracellular nucleotide-derived phosphate hydrolysis; secretory pathway derived cell surface glycoprotein; induced by phosphate starvation and coordinately regulated by PHO4 and PHO2 | Guanosine diphosphatase located in the Golgi, involved in the transport of GDP-mannose into the Golgi lumen by converting GDP to GMP after mannose is transferred its substrate | Gamma subunit of the translation initiation factor eIF2, involved in the identification of the start codon; binds GTP when forming the ternary complex with GTP and tRNAi-Met | Protein of unknown function; interacts with Sup45p (eRF1), Sup35p (eRF3) and Pab1p; has a role in translation termination efficiency, mRNA poly(A) tail length and mRNA stability | Threonine deaminase, catalyzes the first step in isoleucine biosynthesis; expression is under general amino acid control; ILV1 locus exhibits highly positioned nucleosomes whose organization is independent of known ILV1 regulation | Karyopherin beta, mediates nuclear import of ribosomal proteins prior to assembly into ribosomes and import of histones H3 and H4; localizes to the nuclear pore, nucleus, and cytoplasm; exhibits genetic interactions with RAI1 | Poly(A) binding protein, part of the 3'-end RNA-processing complex, mediates interactions between the 5' cap structure and the 3' mRNA poly(A) tail, involved in control of poly(A) tail length, interacts with translation factor eIF-4G | Mitochondrial and cytoplasmic valyl-tRNA synthetase | Hsp70 protein that interacts with Zuo1p (a DnaJ homolog) to form a ribosome-associated complex that binds the ribosome via the Zuo1p subunit; also involved in pleiotropic drug resistance via sequential activation of PDR1 and PDR5; binds ATP | One of three repressible acid phosphatases, a glycoprotein that is transported to the cell surface by the secretory pathway; nearly identical to Pho11p; upregulated by phosphate starvation | Phosphatidylinositol:ceramide phosphoinositol transferase (IPC synthase), required for sphingolipid synthesis; can mutate to confer aureobasidin A resistance | Dihydroorotate dehydrogenase, catalyzes the fourth enzymatic step in the de novo biosynthesis of pyrimidines, converting dihydroorotic acid into orotic acid | Putative protein of unknown function; YLR413W is not an essential gene | C-8 sterol isomerase, catalyzes the isomerization of the delta-8 double bond to the delta-7 position at an intermediate step in ergosterol biosynthesis | Long chain fatty acyl-CoA synthetase, regulates protein modification during growth in the presence of ethanol, functions to incorporate palmitic acid into phospholipids and neutral lipids | Bud-specific protein with a potential role in membrane trafficking; GFP-fusion protein migrates from the cell surface to intracellular vesicles near vacuole; contains 3 calcium and lipid binding domains; mRNA is targeted to the bud | Ubiquitin-protein ligase involved in ubiquitin-mediated protein degradation; functions in multivesicular body sorting, heat shock response and ubiquitylation of arrested RNAPII; contains a hect (homologous to E6-AP carboxyl terminus) domain | Para hydroxybenzoate: polyprenyl transferase, catalyzes the second step in ubiquinone (coenzyme Q) biosynthesis | Mevalonate pyrophosphate decarboxylase, essential enzyme involved in the biosynthesis of isoprenoids and sterols, including ergosterol; acts as a homodimer | Protein of unknown function, component of the SWR1 complex, which exchanges histone variant H2AZ (Htz1p) for chromatin-bound histone H2A; required for formation of nuclear-associated array of smooth endoplasmic reticulum known as karmellae | Protein required for the synthesis of wybutosine, a modified guanosine found at the 3'-position adjacent to the anticodon of phenylalanine tRNA which supports reading frame maintenance by stabilizing codon-anticodon interactions |
| 3 | 38 | 448 | -0.495 | 0.554 | 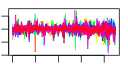 |  | 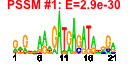 | 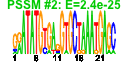 |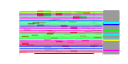  | YBR065C YBR270C YCR104W YDR030C YDR138W YDR283C YDR402C YDR458C YGL260W YGL261C YHR006W YIL066C YIL120W YIL144W YIR027C YIR028W YIR029W YJL105W YKL224C YKR015C YKR045C YLL003W YLL033W YLR082C YLR128W YNL133C YNL187W YNL285W YNR076W YOL075C YOR308C YOR394W YPL052W YPL162C YPL272C YPL282C YPR007C YPR083W | ECM2 BIT2 PAU3 RAD28 HPR1 GCN2 DIT2 HEH2 PAU11 STP2 RNR3 QDR1 TID3 DAL1 DAL4 DAL2 SET4 PAU16 SFI1 IRC19 SRL2 DCN1 FYV6 PAU6 SNU66 PAU21 OAZ1 PAU22 REC8 MDM36 | Pre-mRNA splicing factor, facilitates the cooperative formation of U2/U6 helix II in association with stem II in the spliceosome, function may be regulated by Slu7p | hypothetical protein | Part of 23-member seripauperin multigene family encoded mainly in subtelomeric regions, active during alcoholic fermentation, regulated by anaerobiosis, negatively regulated by oxygen, repressed by heme | Protein involved in transcription-coupled repair nucleotide excision repair of UV-induced DNA lesions; homolog of human CSA protein | Subunit of THO/TREX complexes that couple transcription elongation with mitotic recombination and with mRNA metabolism and export, subunit of an RNA Pol II complex; regulates lifespan; involved in telomere maintenance; similar to Top1p | Protein kinase, phosphorylates the alpha-subunit of translation initiation factor eIF2 (Sui2p) in response to starvation; activated by uncharged tRNAs and the Gcn1p-Gcn20p complex | N-formyltyrosine oxidase, sporulation-specific microsomal enzyme involved in the production of N,N-bisformyl dityrosine required for spore wall maturation, homologous to cytochrome P-450s | Inner nuclear membrane (INM) protein; contains helix-extension-helix (HEH) motif, nuclear localization signal sequence; targeting to the INM requires the Srp1p-Kap95p karyopherins and the Ran cycle | Putative protein of unknown function; transcription is significantly increased in a NAP1 deletion background; deletion mutant has increased accumulation of nickel and selenium | Putative protein of unknown function; mRNA expression appears to be regulated by SUT1 and UPC2 | Transcription factor, activated by proteolytic processing in response to signals from the SPS sensor system for external amino acids; activates transcription of amino acid permease genes | Ribonucleotide-diphosphate reductase (RNR), large subunit; the RNR complex catalyzes the rate-limiting step in dNTP synthesis and is regulated by DNA replication and DNA damage checkpoint pathways via localization of the small subunits | Multidrug transporter of the major facilitator superfamily, required for resistance to quinidine, ketoconazole, fluconazole, and barban | Component of the evolutionarily conserved kinetochore-associated Ndc80 complex (Ndc80p-Nuf2p-Spc24p-Spc25p); conserved coiled-coil protein involved in chromosome segregation, spindle checkpoint activity, kinetochore assembly and clustering | Allantoinase, converts allantoin to allantoate in the first step of allantoin degradation; expression sensitive to nitrogen catabolite repression | Allantoin permease; expression sensitive to nitrogen catabolite repression and induced by allophanate, an intermediate in allantoin degradation | Allantoicase, converts allantoate to urea and ureidoglycolate in the second step of allantoin degradation; expression sensitive to nitrogen catabolite repression and induced by allophanate, an intermediate in allantoin degradation | Protein of unknown function, contains a SET domain | Putative protein of unknown function | Putative protein of unknown function; epitope-tagged protein localizes to the cytoplasm | Centrin (Cdc31p)-binding protein required for spindle pole body (SPB) duplication, localizes to the half-bridge of the SPB, required for progression through G(2)-M transition, has similarity to Xenopus laevis XCAP-C | Putative protein of unknown function; YLL033W is not an essential gene but mutant is defective in spore formation; null mutant displays increased levels of spontaneous Rad52 foci | Protein of unknown function; overexpression suppresses the lethality caused by a rad53 null mutation | Putative Nedd8 ligase; binds Nedd8; involved in cullin neddylation; not essential; similar to C.elegans DCN-1; contains UBA-like ubiquitin-binding domain and a DUF298 domain | Protein of unknown function, required for survival upon exposure to K1 killer toxin; proposed to regulate double-strand break repair via non-homologous end-joining | Putative protein of unknown function; contains a consensus nuclear export signal (NES) sequence similar to the consensus sequence recognized by Crm1p; may interact with or be fuctionally redundant with Prp40p | Putative ABC transporter | Component of the U4/U6.U5 snRNP complex involved in pre-mRNA splicing via spliceosome; has homology to human SART-1 and to an S. pombe protein; snu66 null mutation confers cold-sensitivity but is not lethal at normal growth temperatures | Regulator of ornithine decarboxylase (Spe1p), antizyme that binds to Spe1p to regulate ubiquitin-independent degradation; ribosomal frameshifting during synthesis of Oaz1p and its ubiquitin-mediated degradation are both polyamine-regulated | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the membrane of vacuole with cell cycle-correlated morphology | Putative protein of unknown function; gene expression induced in response to ketoconazole; YPL272C is not an essential gene | Meiosis-specific component of sister chromatid cohesion complex; maintains cohesion between sister chromatids during meiosis I; maintains cohesion between centromeres of sister chromatids until meiosis II; homolog of S. pombe Rec8p | Protein required for normal mitochondrial morphology and inheritance |
| 225 | 31 | 434 | -0.478 | 0.490 |  |  | 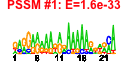 |  |  | YBL101C YDR096W YIL033C YIR007W YKL124W YMR041C YNR034W YOR022C YPR155C YBR203W YCR038C YDR505C YGL160W YGL248W YGR237C YGR255C YJL082W YJR039W YKL140W YKL168C YLR392C YML129C YMR030W YMR173W YMR278W YNL115C YNL293W YOL100W YOL117W YOR208W YPR006C | ECM21 GIS1 BCY1 SSH4 ARA2 SOL1 NCA2 COS111 BUD5 PSP1 AIM14 PDE1 COQ6 IML2 TGL1 KKQ8 COX14 RSF1 DDR48 PGM3 MSB3 PKH2 RRI2 PTP2 ICL2 | Non-essential protein of unknown function; promoter contains several Gcn4p binding elements | JmjC domain-containing histone demethylase; transcription factor involved in the expression of genes during nutrient limitation; also involved in the negative regulation of DPP1 and PHR1 | Regulatory subunit of the cyclic AMP-dependent protein kinase (PKA), a component of a signaling pathway that controls a variety of cellular processes, including metabolism, cell cycle, stress response, stationary phase, and sporulation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YIR007W is a non-essential gene | Vacuolar protein that presumably functions within the endosomal-vacuolar trafficking pathway, affecting events that determine whether plasma membrane proteins are degraded or routed to the plasma membrane | NAD-dependent arabinose dehydrogenase, involved in biosynthesis of erythroascorbic acid; similar to plant L-galactose dehydrogenase | Protein with a possible role in tRNA export; shows similarity to 6-phosphogluconolactonase non-catalytic domains but does not exhibit this enzymatic activity; homologous to Sol2p, Sol3p, and Sol4p | Protein with similarity to bovine phospholipase A1; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein involved in regulation of mitochondrial expression of subunits 6 (Atp6p) and 8 (Atp8p) of the Fo-F1 ATP synthase; functions with Nca3p | Protein required for resistance to the antifungal drug ciclopirox olamine; not related to the subtelomerically-encoded COS family; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | GTP/GDP exchange factor for Rsr1p (Bud1p) required for both axial and bipolar budding patterns; mutants exhibit random budding in all cell types | Asn and gln rich protein of unknown function; high-copy suppressor of POL1 (DNA polymerase alpha) and partial suppressor of CDC2 (polymerase delta) and CDC6 (pre-RC loading factor) mutations; overexpression results in growth inhibition | Putative protein of unknown function; similar to iron/copper reductases (FRE1-8), possibly involved in iron homeostasis; may interact with ribosomes, null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Low-affinity cyclic AMP phosphodiesterase, controls glucose and intracellular acidification-induced cAMP signaling, target of the cAMP-protein kinase A (PKA) pathway; glucose induces transcription and inhibits translation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative flavin-dependent monooxygenase, involved in ubiquinone (Coenzyme Q) biosynthesis; localizes to the matrix face of the mitochondrial inner membrane in a large complex with other ubiquinone biosynthetic enzymes | Protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Steryl ester hydrolase, one of three gene products (Yeh1p, Yeh2p, Tgl1p) responsible for steryl ester hydrolase activity and involved in sterol homeostasis; localized to lipid particle membranes | Putative serine/threonine protein kinase with unknown cellular role | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR392C is not an essential gene | Mitochondrial membrane protein, involved in translational regulation of Cox1p and assembly of cytochrome c oxidase (complex IV); associates with complex IV assembly intermediates and complex III/complex IV supercomplexes | Protein required for respiratory growth; localized to both the nucleus and mitochondrion; mutant displays decreased transcription of specific nuclear and mitochondrial genes whose products are involved in respiratory growth | DNA damage-responsive protein, expression is increased in response to heat-shock stress or treatments that produce DNA lesions; contains multiple repeats of the amino acid sequence NNNDSYGS | Phosphoglucomutase, catalyzes interconversion of glucose-1-phosphate and glucose-6-phospate; transcription induced in response to stress; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; non-essential | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YNL115C is not an essential gene | GTPase-activating protein for Sec4p and several other Rab GTPases, regulates exocytosis via its action on Sec4p, also required for proper actin organization; similar to Msb4p; both Msb3p and Msb4p localize to sites of polarized growth | Serine/threonine protein kinase involved in sphingolipid-mediated signaling pathway that controls endocytosis; activates Ypk1p and Ykr2p, components of signaling cascade required for maintenance of cell wall integrity; redundant with Pkh1p | Subunit of the COP9 signalosome (CSN) complex that cleaves the ubiquitin-like protein Nedd8 from SCF ubiquitin ligases; plays a role in the mating pheromone response | Phosphotyrosine-specific protein phosphatase involved in the inactivation of mitogen-activated protein kinase (MAPK) during osmolarity sensing; dephosporylates Hog1p MAPK and regulates its localization; localized to the nucleus | 2-methylisocitrate lyase of the mitochondrial matrix, functions in the methylcitrate cycle to catalyze the conversion of 2-methylisocitrate to succinate and pyruvate; ICL2 transcription is repressed by glucose and induced by ethanol |
| 172 | 37 | 436 | -0.447 | 0.514 | 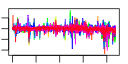 |  | 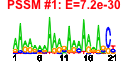 | 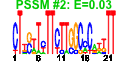 |  | YDL161W YJL203W YJL204C YJR005W YLR039C YLR086W YLR115W YLR182W YLR187W YLR207W YLR319C YLR320W YML062C YMR066W YMR172W YNL076W YNL118C YNL267W YNL316C YOL107W YOL113W YOR109W YOR141C YKL195W YKR023W YLR085C YLR277C YLR427W YMR270C YNL027W YNL148C YNL236W YNL304W YNL310C YOR112W YOR299W YPL155C | ENT1 PRP21 RCY1 APL1 RIC1 SMC4 CFT2 SWI6 SKG3 HRD3 BUD6 MMS22 MFT1 SOV1 HOT1 MKS1 DCP2 PIK1 PHA2 SKM1 SRO77 ARP8 MIA40 ARP6 YSH1 MAG2 RRN9 CRZ1 ALF1 SIN4 YPT11 ZIM17 CEX1 BUD7 KIP2 | Epsin-like protein involved in endocytosis and actin patch assembly and functionally redundant with Ent2p; binds clathrin via a clathrin-binding domain motif at C-terminus | Subunit of the SF3a splicing factor complex, required for spliceosome assembly | F-box protein involved in recycling plasma membrane proteins internalized by endocytosis; localized to sites of polarized growth | Beta-adaptin, large subunit of the clathrin associated protein complex (AP-2); involved in vesicle mediated transport; similar to mammalian beta-chain of the clathrin associated protein complex | Protein involved in retrograde transport to the cis-Golgi network; forms heterodimer with Rgp1p that acts as a GTP exchange factor for Ypt6p; involved in transcription of rRNA and ribosomal protein genes | Subunit of the condensin complex, which reorganizes chromosomes during cell division, forms a stable complex with Smc2p that has ATP-hydrolyzing and DNA-binding activity and promotes knotting of circular DNA; potential Cdc28p substrate | Subunit of the mRNA cleavage and polyadenlylation factor (CPF); required for pre-mRNA cleavage, polyadenylation and poly(A) site recognition, 43% similarity with the mammalian CPSF-100 protein. | Transcription cofactor, forms complexes with DNA-binding proteins Swi4p and Mbp1p to regulate transcription at the G1/S transition; involved in meiotic gene expression; localization regulated by phosphorylation; potential Cdc28p substrate | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery, cytoplasm, bud, and bud neck; potential Cdc28p substrate; similar to Caf120p and Skg4p | Resident protein of the ER membrane that plays a central role in ER-associated protein degradation (ERAD), forms HRD complex with Hrd1p and ERAD determinants that engages in lumen to cytosol communication and coordination of ERAD events | Actin- and formin-interacting protein, involved in actin cable nucleation and polarized cell growth; isolated as bipolar budding mutant; potential Cdc28p substrate | Protein involved in resistance to ionizing radiation; acts with Mms1p in a repair pathway that may be involved in resolving replication intermediates or preventing the damage caused by blocked replication forks | Subunit of the THO complex, which is a nuclear complex comprised of Hpr1p, Mft1p, Rlr1p, and Thp2p, that is involved in transcription elongation and mitotic recombination; involved in telomere maintenance | Mitochondrial protein of unknown function | Transcription factor required for the transient induction of glycerol biosynthetic genes GPD1 and GPP2 in response to high osmolarity; targets Hog1p to osmostress responsive promoters; has similarity to Msn1p and Gcr1p | Pleiotropic negative transcriptional regulator involved in Ras-CAMP and lysine biosynthetic pathways and nitrogen regulation; involved in retrograde (RTG) mitochondria-to-nucleus signaling | Catalytic subunit of the Dcp1p-Dcp2p decapping enzyme complex, which removes the 5' cap structure from mRNAs prior to their degradation; member of the Nudix hydrolase family | Phosphatidylinositol 4-kinase; catalyzes first step in the biosynthesis of phosphatidylinositol-4,5-biphosphate; may control cytokineses through the actin cytoskeleton | Prephenate dehydratase, catalyzes the conversion of prephanate to phenylpyruvate, which is a step in the phenylalanine biosynthesis pathway | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and colocalizes in a punctate pattern with the early golgi/COPI vesicles; YOL107W is not an essential protein | Member of the PAK family of serine/threonine protein kinases with similarity to Ste20p and Cla4p; proposed to be a downstream effector of Cdc42p during polarized growth | Protein with roles in exocytosis and cation homeostasis; functions in docking and fusion of post-Golgi vesicles with plasma membrane; homolog of Sro7p and Drosophila lethal giant larvae tumor suppressor; interacts with SNARE protein Sec9p | Nuclear actin-related protein involved in chromatin remodeling, component of chromatin-remodeling enzyme complexes | Essential protein of the mitochondrial intermembrane space (IMS); promotes retention of newly imported proteins; may do so by stabilizing client protein folding as part of a disulfide relay system or transferring metal to client proteins | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Actin-related protein that binds nucleosomes; a component of the SWR1 complex, which exchanges histone variant H2AZ (Htz1p) for chromatin-bound histone H2A | Putative endoribonuclease, subunit of the mRNA cleavage and polyadenylation specificity complex required for 3' processing of mRNAs | Cytoplasmic protein of unknown function predicted to encode a DNA-3-methyladenine glycosidase II that catalyzes the hydrolysis of alkylated DNA | Protein involved in promoting high level transcription of rDNA, subunit of UAF (upstream activation factor) for RNA polymerase I | Transcription factor that activates transcription of genes involved in stress response; nuclear localization is positively regulated by calcineurin-mediated dephosphorylation | Alpha-tubulin folding protein, similar to mammalian cofactor B; Alf1p-GFP localizes to cytoplasmic microtubules; required for the folding of alpha-tubulin and may play an additional role in microtubule maintenance | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; contributes to both postive and negative transcriptional regulation; dispensible for basal transcription | Rab-type small GTPase that interacts with the C-terminal tail domain of Myo2p to mediate distribution of mitochondria to daughter cells | Heat shock protein with a zinc finger motif; essential for protein import into mitochondria; may act with Pam18p to facilitate recognition and folding of imported proteins by Ssc1p (mtHSP70) in the mitochondrial matrix | Cytoplasmic component of the nuclear aminoacylation-dependent tRNA export pathway; interacts with nuclear pore component Nup116p; copurifies with tRNA export receptors Los1p and Msn5p, as well as eIF-1a and the RAN GTPase Gsp1p | Member of the ChAPs family of proteins (Chs5p-Arf1p-binding proteins: Bch1p, Bch2p, Bud7p, Chs6p), that forms the exomer complex with Chs5p to mediate export of specific cargo proteins, including Chs3p, from the Golgi to the plasma membrane | Kinesin-related motor protein involved in mitotic spindle positioning, stabilizes microtubules by targeting Bik1p to the plus end; Kip2p levels are controlled during the cell cycle |
| 57 | 36 | 440 | -0.437 | 0.499 | 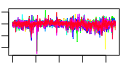 |  | 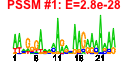 | 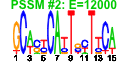 |  | YDR172W YER125W YGR204W YJL130C YJR016C YJR137C YKL057C YKL068W YKL127W YKL212W YLR347C YMR080C YMR125W YMR205C YMR300C YMR308C YNL085W YNL197C YNL287W YOL098C YOL149W YOR151C YOR204W YOR217W YOR270C YOR326W YOR335C YPL115C YPL128C YPL210C YPL244C YPL254W YPR074C YPR181C YJL168C YNL216W | SUP35 RSP5 ADE3 URA2 ILV3 ECM17 NUP120 YKL068W-A PGM1 SAC1 KAP95 NAM7 SUT1 PFK2 ADE4 PSE1 MKT1 WHI3 SEC21 DCP1 RPB2 DED1 RFC1 VPH1 MYO2 ALA1 BEM3 TBF1 SRP72 HUT1 HFI1 TKL1 SEC23 SET2 RAP1 | Translation termination factor eRF3; altered protein conformation creates the [PSI(+)] prion, a dominant cytoplasmically inherited protein aggregate that alters translational fidelity and creates a nonsense suppressor phenotype | Ubiquitin-protein ligase involved in ubiquitin-mediated protein degradation; functions in multivesicular body sorting, heat shock response and ubiquitylation of arrested RNAPII; contains a hect (homologous to E6-AP carboxyl terminus) domain | Cytoplasmic trifunctional enzyme C1-tetrahydrofolate synthase, involved in single carbon metabolism and required for biosynthesis of purines, thymidylate, methionine, and histidine | Bifunctional carbamoylphosphate synthetase (CPSase)-aspartate transcarbamylase (ATCase), catalyzes the first two enzymatic steps in the de novo biosynthesis of pyrimidines; both activities are subject to feedback inhibition by UTP | Dihydroxyacid dehydratase, catalyzes third step in the common pathway leading to biosynthesis of branched-chain amino acids | Sulfite reductase beta subunit, involved in amino acid biosynthesis, transcription repressed by methionine | Subunit of the Nup84p subcomplex of the nuclear pore complex (NPC), required for even distribution of NPCs around the nuclear envelope, involved in establishment of a normal nucleocytoplasmic concentration gradient of the GTPase Gsp1p | Putative protein of unknown function; identified by homology to Ashbya gossypii | Phosphoglucomutase, minor isoform; catalyzes the conversion from glucose-1-phosphate to glucose-6-phosphate, which is a key step in hexose metabolism | Phosphatidylinositol (PI) phosphatase, involved in hydrolysis of PI 3-phosphate, PI 4-phosphate and PI 3,5-bisphosphate to PI; membrane protein localizes to ER and Golgi; involved in protein trafficking, secretion and inositol metabolism | Karyopherin beta, forms a dimeric complex with Srp1p (Kap60p) that mediates nuclear import of cargo proteins via a nuclear localization signal (NLS), interacts with nucleoporins to guide transport across the nuclear pore complex | ATP-dependent RNA helicase of the SFI superfamily, required for nonsense mediated mRNA decay and for efficient translation termination at nonsense codons; involved in telomere maintenance | Transcription factor of the Zn[II]2Cys6 family involved in sterol uptake; involved in induction of hypoxic gene expression | Beta subunit of heterooctameric phosphofructokinase involved in glycolysis, indispensable for anaerobic growth, activated by fructose-2,6-bisphosphate and AMP, mutation inhibits glucose induction of cell cycle-related genes | Phosphoribosylpyrophosphate amidotransferase (PRPPAT; amidophosphoribosyltransferase), catalyzes first step of the 'de novo' purine nucleotide biosynthetic pathway | Karyopherin/importin that interacts with the nuclear pore complex; acts as the nuclear import receptor for specific proteins, including Pdr1p, Yap1p, Ste12p, and Aft1p | Protein that forms a complex with Pbp1p that may mediate posttranscriptional regulation of HO endonuclease; involved in propagation of M2 dsRNA satellite of L-A virus; contains a DTG signature typical of retroviral proteases | RNA binding protein that sequesters CLN3 mRNA in cytoplasmic foci; cytoplasmic retention factor for Cdc28p and associated cyclins; regulates cell fate and dose-dependently regulates the critical cell size required for passage through Start | Gamma subunit of coatomer, a heptameric protein complex that together with Arf1p forms the COPI coat; involved in ER to Golgi transport of selective cargo | Putative metalloprotease | Subunit of the Dcp1p-Dcp2p decapping enzyme complex, which removes the 5' cap structure from mRNAs prior to their degradation; enhances the activity of catalytic subunit Dcp2p; regulated by DEAD box protein Dhh1p | RNA polymerase II second largest subunit B150, part of central core; similar to bacterial beta subunit | ATP-dependent DEAD (Asp-Glu-Ala-Asp)-box RNA helicase, required for translation initiation of all yeast mRNAs; mutations in human DEAD-box DBY are a frequent cause of male infertility | Subunit of heteropentameric Replication factor C (RF-C), which is a DNA binding protein and ATPase that acts as a clamp loader of the proliferating cell nuclear antigen (PCNA) processivity factor for DNA polymerases delta and epsilon | Subunit a of vacuolar-ATPase V0 domain, one of two isoforms (Vph1p and Stv1p); Vph1p is located in V-ATPase complexes of the vacuole while Stv1p is located in V-ATPase complexes of the Golgi and endosomes | One of two type V myosin motors (along with MYO4) involved in actin-based transport of cargos; required for the polarized delivery of secretory vesicles, the vacuole, late Golgi elements, peroxisomes, and the mitotic spindle | Cytoplasmic alanyl-tRNA synthetase, required for protein synthesis; point mutation (cdc64-1 allele) causes cell cycle arrest at G1; lethality of null mutation is functionally complemented by human homolog | Rho GTPase activating protein (RhoGAP) involved in control of the cytoskeleton organization; targets the essential Rho-GTPase Cdc42p, which controls establishment and maintenance of cell polarity, including bud-site assembly | Telobox-containing general regulatory factor; binds to TTAGGG repeats within subtelomeric anti-silencing regions (STARs) and possibly throughout the genome and mediates their insulating capacity by blocking silent chromatin propagation | Core component of the signal recognition particle (SRP) ribonucleoprotein (RNP) complex that functions in targeting nascent secretory proteins to the endoplasmic reticulum (ER) membrane | Protein with a role in UDP-galactose transport to the Golgi lumen, has similarity to human UDP-galactose transporter UGTrel1, exhibits a genetic interaction with S. cerevisiae ERO1 | Adaptor protein required for structural integrity of the SAGA complex, a histone acetyltransferase-coactivator complex that is involved in global regulation of gene expression through acetylation and transcription functions | Transketolase, similar to Tkl2p; catalyzes conversion of xylulose-5-phosphate and ribose-5-phosphate to sedoheptulose-7-phosphate and glyceraldehyde-3-phosphate in the pentose phosphate pathway; needed for synthesis of aromatic amino acids | GTPase-activating protein; component of the Sec23p-Sec24p heterodimeric complex of the COPII vesicle coat, involved in ER to Golgi transport and autophagy; stimulates the GDP-bound form of Sar1p | Histone methyltransferase with a role in transcriptional elongation, methylates a lysine residue of histone H3; associates with the C-terminal domain of Rpo21p; histone methylation activity is regulated by phosphorylation status of Rpo21p | DNA-binding protein involved in either activation or repression of transcription, depending on binding site context; also binds telomere sequences and plays a role in telomeric position effect (silencing) and telomere structure |
| 81 | 48 | 435 | -0.433 | 0.523 |  |  | 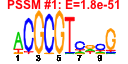 | 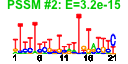 |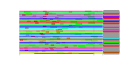  | YBL035C YBR088C YDL101C YDR501W YGR188C YJL115W YJR030C YLL002W YLR313C YNL206C YOL017W YOL102C YOR074C YOR242C YPL208W YPR135W YPR141C YBR070C YCL024W YCL060C YCL061C YDL003W YDR097C YJL074C YKL042W YKL045W YKL113C YKR077W YLL021W YLR103C YLR383W YMR076C YMR078C YMR144W YMR179W YNL082W YNL088W YNL102W YNL126W YNL166C YNR009W YOR144C YOR368W YPL209C YPL241C YPL255W YPL267W YPR120C | POL12 POL30 DUN1 PLM2 BUB1 CIA1 RTT109 SPH1 RTT106 ESC8 TPT1 CDC21 SSP2 RKM1 CTF4 KAR3 ALG14 KCC4 MRC1 MCD1 MSH6 SMC3 SPC42 PRI2 FEN1 MSA2 SPA2 CDC45 SMC6 PDS5 CTF18 SPT21 PMS1 TOP2 POL1 SPC98 BNI5 NRM1 ELG1 RAD17 IPL1 CIN2 BBP1 ACM1 CLB5 | B subunit of DNA polymerase alpha-primase complex, required for initiation of DNA replication during mitotic and premeiotic DNA synthesis; also functions in telomere capping and length regulation | Proliferating cell nuclear antigen (PCNA), functions as the sliding clamp for DNA polymerase delta; may function as a docking site for other proteins required for mitotic and meiotic chromosomal DNA replication and for DNA repair | Cell-cycle checkpoint serine-threonine kinase required for DNA damage-induced transcription of certain target genes, phosphorylation of Rad55p and Sml1p, and transient G2/M arrest after DNA damage; also regulates postreplicative DNA repair | Protein required for partitioning of the 2-micron plasmid | Protein kinase that forms a complex with Mad1p and Bub3p that is crucial in the checkpoint mechanism required to prevent cell cycle progression into anaphase in the presence of spindle damage, associates with centromere DNA via Skp1p | Essential protein involved in assembly of cytosolic and nuclear iron-sulfur proteins | Putative protein of unknown function; expression repressed in carbon limited vs carbon replete chemostat cultures; YJR030C is a non-essential gene | Histone acetyltransferase critical for cell survival in the presence of DNA damage during S phase, acetylates H3-K56; plays a role in regulation of Ty1 transposition | Protein involved in shmoo formation and bipolar bud site selection; homologous to Spa2p, localizes to sites of polarized growth in a cell cycle dependent- and Spa2p-dependent manner, interacts with MAPKKs Mkk1p, Mkk2p, and Ste7p | Protein with a role in regulation of Ty1 transposition | Protein involved in telomeric and mating-type locus silencing, interacts with Sir2p and also interacts with the Gal11p, which is a component of the RNA pol II mediator complex | tRNA 2'-phosphotransferase, catalyzes the final step in yeast tRNA splicing: the transfer of the 2'-PO(4) from the splice junction to NAD(+) to form ADP-ribose 1''-2''cyclic phosphate and nicotinamide | Thymidylate synthase, required for de novo biosynthesis of pyrimidine deoxyribonucleotides; expression is induced at G1/S | Sporulation specific protein that localizes to the spore wall; required for sporulation at a point after meiosis II and during spore wall formation; SSP2 expression is induced midway in meiosis | SET-domain lysine-N-methyltransferase, catalyzes the formation of dimethyllysine residues on the large ribsomal subunit protein L23a (RPL23A and RPL23B) | Chromatin-associated protein, required for sister chromatid cohesion; interacts with DNA polymerase alpha (Pol1p) and may link DNA synthesis to sister chromatid cohesion | Minus-end-directed microtubule motor that functions in mitosis and meiosis, localizes to the spindle pole body and localization is dependent on functional Cik1p, required for nuclear fusion during mating; potential Cdc28p substrate | Component of UDP-GlcNAc transferase required for the second step of dolichyl-linked oligosaccharide synthesis; anchors the catalytic subunit Alg13p to the ER membrane; similar to bacterial and human glycosyltransferases | Protein kinase of the bud neck involved in the septin checkpoint, associates with septin proteins, negatively regulates Swe1p by phosphorylation, shows structural homology to bud neck kinases Gin4p and Hsl1p | S-phase checkpoint protein found at replication forks, required for DNA replication; also required for Rad53p activation during DNA replication stress, where it forms a replication-pausing complex with Tof1p and is phosphorylated by Mec1p | Essential protein required for sister chromatid cohesion in mitosis and meiosis; subunit of the cohesin complex; expression is cell cycle regulated and peaks in S phase | Protein required for mismatch repair in mitosis and meiosis, forms a complex with Msh2p to repair both single-base & insertion-deletion mispairs; potentially phosphorylated by Cdc28p | Subunit of the multiprotein cohesin complex required for sister chromatid cohesion in mitotic cells; also required, with Rec8p, for cohesion and recombination during meiosis; phylogenetically conserved SMC chromosomal ATPase family member | Central plaque component of spindle pole body (SPB); involved in SPB duplication, may facilitate attachment of the SPB to the nuclear membrane | Subunit of DNA primase, which is required for DNA synthesis and double-strand break repair | Fatty acid elongase, involved in sphingolipid biosynthesis; acts on fatty acids of up to 24 carbons in length; mutations have regulatory effects on 1,3-beta-glucan synthase, vacuolar ATPase, and the secretory pathway | Putative transcriptional activator, that interacts with G1-specific transcription factor, MBF and G1-specific promoters; ortholog of Msa2p, an MBF and SBF activator that regulates G1-specific transcription and cell cycle initiation | Component of the polarisome, which functions in actin cytoskeletal organization during polarized growth; acts as a scaffold for Mkk1p and Mpk1p cell wall integrity signaling components; potential Cdc28p substrate | DNA replication initiation factor; recruited to MCM pre-RC complexes at replication origins; promotes release of MCM from Mcm10p, recruits elongation machinery; mutants in human homolog may cause velocardiofacial and DiGeorge syndromes | Protein involved in structural maintenance of chromosomes; essential subunit of Mms21-Smc5-Smc6 complex; required for growth, DNA repair, interchromosomal and sister chromatid recombination; homologous to S. pombe rad18 | Protein required for establishment and maintenance of sister chromatid condensation and cohesion, colocalizes with cohesin on chromosomes in an interdependent manner, may function as a protein-protein interaction scaffold | Subunit of a complex with Ctf8p that shares some subunits with Replication Factor C and is required for sister chromatid cohesion; may have overlapping functions with Rad24p in the DNA damage replication checkpoint | Putative protein of unknown function; localized to the nucleus; YMR144W is not an essential gene | Protein required for normal transcription at several loci including HTA2-HTB2 and HHF2-HHT2, but not required at the other histone loci; functionally related to Spt10p; involved in telomere maintenance | ATP-binding protein required for mismatch repair in mitosis and meiosis; functions as a heterodimer with Mlh1p, binds double- and single-stranded DNA via its N-terminal domain, similar to E. coli MutL | Essential type II topoisomerase, relieves torsional strain in DNA by cleaving and re-sealing the phosphodiester backbone of both positively and negatively supercoiled DNA; cleaves complementary strands; localizes to axial cores in meiosis | Catalytic subunit of the DNA polymerase I alpha-primase complex, required for the initiation of DNA replication during mitotic DNA synthesis and premeiotic DNA synthesis | Component of the microtubule-nucleating Tub4p (gamma-tubulin) complex; interacts with Spc110p at the spindle pole body (SPB) inner plaque and with Spc72p at the SPB outer plaque | Protein involved in organization of septins at the mother-bud neck, may interact directly with the Cdc11p septin, localizes to bud neck in a septin-dependent manner | Transcriptional co-repressor of MBF (MCB binding factor)-regulated gene expression; Nrm1p associates stably with promoters via MBF to repress transcription upon exit from G1 phase | Protein required for S phase progression and telomere homeostasis, forms an alternative replication factor C complex important for DNA replication and genome integrity; involved in homologous recombination-mediated DNA repair | Checkpoint protein, involved in the activation of the DNA damage and meiotic pachytene checkpoints; with Mec3p and Ddc1p, forms a clamp that is loaded onto partial duplex DNA; homolog of human and S. pombe Rad1 and U. maydis Rec1 proteins | Aurora kinase involved in regulating kinetochore-microtubule attachments; helps maintain condensed chromosomes during anaphase and early telophase; associates with Sli15p, which promotes Ipl1p association with mitotic spindle | Tubulin folding factor C (putative) involved in beta-tubulin (Tub2p) folding; isolated as mutant with increased chromosome loss and sensitivity to benomyl | Protein required for the spindle pole body (SPB) duplication, localized at the central plaque periphery; forms a complex with a nuclear envelope protein Mps2p and SPB components Spc29p and Kar1p; required for mitotic functions of Cdc5p | Cell cycle regulated protein of unknown function; associated with Cdh1p and may supress the APC/C[Cdh1]-mediated proteolysis of mitotic cyclins | B-type cyclin involved in DNA replication during S phase; activates Cdc28p to promote initiation of DNA synthesis; functions in formation of mitotic spindles along with Clb3p and Clb4p; most abundant during late G1 phase |
| 143 | 23 | 439 | -0.431 | 0.462 |  |  | 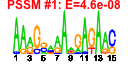 |  |  | YBR269C YDL021W YDL091C YER033C YHR140W YIL097W YJL016W YJL017W YKR049C YLR252W YBL075C YBR204C YDR003W YDR425W YEL039C YER053C YGR127W YGR256W YIL113W YJL057C YLR201C YMR133W YOR220W | FMP21 GPM2 UBX3 ZRG8 FYV10 FMP46 SSA3 RCR2 SNX41 CYC7 PIC2 GND2 SDP1 IKS1 COQ9 REC114 RCN2 | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Homolog of Gpm1p phosphoglycerate mutase which converts 3-phosphoglycerate to 2-phosphoglycerate in glycolysis; may be non-functional derivative of a gene duplication event | UBX (ubiquitin regulatory X) domain-containing protein that interacts with Cdc48p, green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Protein of unknown function; authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; GFP-fusion protein is localized to the cytoplasm; transcription induced under conditions of zinc deficiency | Putative integral membrane protein of unknown function | Protein of unknown function, required for survival upon exposure to K1 killer toxin; involved in proteasome-dependent catabolite inactivation of fructose-1,6-bisphosphatase; contains CTLH domain | Putative protein of unknown function; GFP-fusion protein localizes to the cytoplasm; conserved in closely related Saccharomyces species | Putative redox protein containing a thioredoxin fold; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | ATPase involved in protein folding and the response to stress; plays a role in SRP-dependent cotranslational protein-membrane targeting and translocation; member of the heat shock protein 70 (HSP70) family; localized to the cytoplasm | Serine hydrolase; YBR204C is not an essential gene | Vacuolar protein that presumably functions within the endosomal-vacuolar trafficking pathway, affecting events that determine whether plasma membrane proteins are degraded or routed to the plasma membrane; similar to Rcr1p | Sorting nexin, involved in the retrieval of late-Golgi SNAREs from the post-Golgi endosome to the trans-Golgi network; forms a complex with Snx4p and Atg20p | Cytochrome c isoform 2, expressed under hypoxic conditions; electron carrier of the mitochondrial intermembrane space that transfers electrons from ubiquinone-cytochrome c oxidoreductase to cytochrome c oxidase during cellular respiration | Mitochondrial phosphate carrier, imports inorganic phosphate into mitochondria; functionally redundant with Mir1p but less abundant than Mir1p under normal conditions; expression is induced at high temperature | Putative protein of unknown function; expression is regulated by Msn2p/Msn4p, indicating a possible role in stress response | 6-phosphogluconate dehydrogenase (decarboxylating), catalyzes an NADPH regenerating reaction in the pentose phosphate pathway; required for growth on D-glucono-delta-lactone | Stress-inducible dual-specificity MAP kinase phosphatase, negatively regulates Slt2p MAP kinase by direct dephosphorylation, diffuse localization under normal conditions shifts to punctate localization after heat shock | Putative serine/threonine kinase; expression is induced during mild heat stress; deletion mutants are hypersensitive to copper sulphate and resistant to sorbate; interacts with an N-terminal fragment of Sst2p | Protein required for ubiquinone (coenzyme Q) biosynthesis and respiratory growth; localizes to the matrix face of the mitochondrial inner membrane in a large complex with ubiquinone biosynthetic enzymes | Protein involved in early stages of meiotic recombination; possibly involved in the coordination of recombination and meiotic division; mutations lead to premature initiation of the first meiotic division | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and is induced in response to the DNA-damaging agent MMS |
| 8 | 32 | 439 | -0.421 | 0.482 |  |  |  |  |  | YCL052C YDR421W YER087W YGL004C YJL036W YKL013C YKL064W YKL100C YKR086W YKR091W YLL001W YLR035C YLR273C YLR369W YMR020W YMR027W YMR152W YMR160W YMR258C YNL214W YNL242W YOL159C YOR037W YOR059C YOR155C YOR386W YPR006C YPR081C YIL152W YML117W YOL065C YOR352W | PBN1 ARO80 AIM10 RPN14 SNX4 ARC19 MNR2 PRP16 SRL3 DNM1 MLH2 PIG1 SSQ1 FMS1 YIM1 PEX17 ATG2 YOL159C-A CYC2 ISN1 PHR1 ICL2 GRS2 NAB6 INP54 | Essential component of glycosylphosphatidylinositol-mannosyltransferase I, required for the autocatalytic post-translational processing of the protease B precursor Prb1p, localizes to ER in lumenal orientation; homolog of mammalian PIG-X | Zinc finger transcriptional activator of the Zn2Cys6 family; activates transcription of aromatic amino acid catabolic genes in the presence of aromatic amino acids | Protein with similarity to tRNA synthetases; non-tagged protein is detected in purified mitochondria; null mutant displays decreased frequency of mitochondrial genome loss (petite formation), severe growth defect in minimal glycerol media | Putative non-ATPase subunit of the 19S regulatory particle of the 26S proteasome; localized to the cytoplasm | Sorting nexin, involved in the retrieval of late-Golgi SNAREs from the post-Golgi endosome to the trans-Golgi network and in cytoplasm to vacuole transport; contains a PX domain; forms complex with Snx41p and Atg20p | Subunit of the ARP2/3 complex, which is required for the motility and integrity of cortical actin patches | Putative magnesium transporter; has similarity to Alr1p and Alr2p, which mediate influx of Mg2+ and other divalent cations | Putative protein of unknown function with similarity to a human minor histocompatibility antigen; YKL100C is not an essential gene | RNA helicase in the DEAH-box family involved in the second catalytic step of splicing, exhibits ATP-dependent RNA unwinding activity | Cytoplasmic protein that, when overexpressed, suppresses the lethality of a rad53 null mutation; potential Cdc28p substrate | Dynamin-related GTPase required for mitochondrial fission and the maintenance of mitochondrial morphology, assembles on the cytoplasmic face of mitochondrial tubules at sites at which division will occur; also participates in endocytosis | Protein involved in the mismatch repair of certain frameshift intermediates and involved in meiotic recombination; forms a complex with Mlh1p | Putative targeting subunit for the type-1 protein phosphatase Glc7p that tethers it to the Gsy2p glycogen synthase | Mitochondrial hsp70-type molecular chaperone, required for assembly of iron/sulfur clusters into proteins at a step after cluster synthesis, and for maturation of Yfh1p, which is a homolog of human frataxin implicated in Friedreich's ataxia | Polyamine oxidase, converts spermine to spermidine, which is required for the essential hypusination modification of translation factor eIF-5A; also involved in pantothenic acid biosynthesis | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and cytoplasm; YMR027W is not an essential gene | Protein of unknown function; null mutant displays sensitivity to DNA damaging agents; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the membrane of the vacuole; mutant has enhanced sensitivity to overexpression of mutant huntingtin; YMR160W is not an essential gene | Protein of unknown function with similarity to F-box proteins; physically interacts with Skp1p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YMR258C is not an essential gene | Peroxisomal membrane peroxin and subunit of the docking complex that facilitates the import of peroxisomal matrix proteins; required for peroxisome biogenesis | Peripheral membrane protein required for the formation of cytosolic sequestering vesicles involved in vacuolar import through both the Cvt pathway and autophagy; interacts with Atg9p and is necessary for its trafficking | Putative protein of unknown function; identified by sequence comparison with hemiascomycetous yeast species | Mitochondrial peripheral inner membrane protein, contains a FAD cofactor in a domain exposed in the intermembrane space; exhibits redox activity in vitro; likely participates in ligation of heme to acytochromes c and c1 (Cyc1p and Cyt1p) | hypothetical protein | Inosine 5'-monophosphate (IMP)-specific 5'-nucleotidase, catalyzes the breakdown of IMP to inosine, does not show similarity to known 5'-nucleotidases from other organisms | DNA photolyase involved in photoreactivation, repairs pyrimidine dimers in the presence of visible light; induced by DNA damage; regulated by transcriptional repressor Rph1p | 2-methylisocitrate lyase of the mitochondrial matrix, functions in the methylcitrate cycle to catalyze the conversion of 2-methylisocitrate to succinate and pyruvate; ICL2 transcription is repressed by glucose and induced by ethanol | Protein with sequence similarity to Grs1p, which is a glycyl-tRNA synthetase; cannot substitute for Grs1p; possible pseudogene that is expressed at very low levels | Putative protein of unknown function | Putative RNA-binding protein, based on computational analysis of large-scale protein-protein interaction data | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase with a role in secretion, localizes to the endoplasmic reticulum via the C-terminal tail; lacks the Sac1 domain and proline-rich region found in the other 3 INP proteins | Putative protein of unknown function; expression levels regulated by Arg5,6p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus |
| 262 | 45 | 425 | -0.420 | 0.499 | 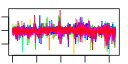 |  | 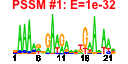 | 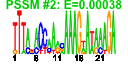 |  | YLR057W YNL032W YPL030W YBR102C YCR073C YDR257C YJL085W YJL087C YJL105W YJL197W YJR140C YKL033W YKL072W YLL018C YLR015W YLR067C YLR086W YLR263W YLR272C YLR319C YLR320W YLR321C YLR442C YML006C YML037C YML065W YML071C YML097C YML109W YMR038C YNL254C YNR011C YOL006C YOL028C YOL115W YOR337W YPL029W YPL083C YPL122C YPL175W YPL210C YPR141C YPR174C YPR175W YPR189W | SIW14 TRM44 EXO84 SSK22 SET7 EXO70 TRL1 SET4 UBP12 HIR3 YKL033W-A STB6 COX19 BRE2 PET309 SMC4 RED1 YCS4 BUD6 MMS22 YKL091C SIR3 GIS4 ORC1 COG8 VPS9 ZDS2 CCS1 PRP2 TOP1 YAP7 PAP2 TEA1 SUV3 SEN54 TFB2 SPT14 SRP72 KAR3 DPB2 SKI3 | Putative protein of unknown function; YLR050C is not an essential gene | Tyrosine phosphatase that plays a role in actin filament organization and endocytosis; localized to the cytoplasm | Putative tRNA U44 2'-O-methyltransferase; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; non-essential gene | Essential protein with dual roles in spliceosome assembly and exocytosis; the exocyst complex (Sec3p, Sec5p, Sec6p, Sec8p, Sec10p, Sec15p, Exo70p, and Exo84p) mediates polarized targeting of secretory vesicles to active sites of exocytosis | MAP kinase kinase kinase of the HOG1 mitogen-activated signaling pathway; functionally redundant with, and homologous to, Ssk2p; interacts with and is activated by Ssk1p; phosphorylates Pbs2p | Nuclear protein that contains a SET-domain, which have been shown to mediate methyltransferase activity in other proteins | Essential 70kDa subunit of the exocyst complex (Sec3p, Sec5p, Sec6p, Sec8p, Sec10p, Sec15p, Exo70p, and Exo84p), which has the essential function of mediating polarized targeting of secretory vesicles to active sites of exocytosis | tRNA ligase, required for tRNA splicing; composed of three essential domains containing the phosphodiesterase, polynucleotide kinase, and ligase activities required for ligation; localized at the inner membrane of the nuclear envelope | Protein of unknown function, contains a SET domain | Ubiquitin carboxyl-terminal hydrolase, ubiquitin-specific protease present in the nucleus and cytoplasm that cleaves ubiquitin from ubiquitinated proteins | Transcriptional corepressor involved in the cell cycle-regulated transcription of histone genes HTA1, HTB1, HHT1, and HHT2; involved in position-dependent gene silencing and nucleosome reassembly | Putative protein of unknown function; similar to uncharacterized proteins from other fungi | Protein that binds Sin3p in a two-hybrid assay | Protein required for cytochrome c oxidase assembly, located in the cytosol and mitochondrial intermembrane space; putative copper metallochaperone that delivers copper to cytochrome c oxidase | Subunit of the COMPASS (Set1C) complex, which methylates histone H3 on lysine 4 and is required in transcriptional silencing near telomeres; involved in telomere maintenance; similar to trithorax-group protein ASH2L | Specific translational activator for the COX1 mRNA, also influences stability of intron-containing COX1 primary transcripts; located in the mitochondrial inner membrane | Subunit of the condensin complex, which reorganizes chromosomes during cell division, forms a stable complex with Smc2p that has ATP-hydrolyzing and DNA-binding activity and promotes knotting of circular DNA; potential Cdc28p substrate | Protein component of the axial elements of the synaptonemal complex, involved in chromosome segregation during the first meiotic division; interacts with Hop1p; required for wild-type levels of Mek1p kinase activity | Non-SMC subunit of the condensin complex (Smc2p-Smc4p-Ycs4p-Brn1p-Ycg1p); required for establishment and maintenance of chromosome condensation, chromosome segregation, chromatin binding of condensin and silencing at the mating type locus | Actin- and formin-interacting protein, involved in actin cable nucleation and polarized cell growth; isolated as bipolar budding mutant; potential Cdc28p substrate | Protein involved in resistance to ionizing radiation; acts with Mms1p in a repair pathway that may be involved in resolving replication intermediates or preventing the damage caused by blocked replication forks | Putative homolog of Sec14p, which is a phosphatidylinositol/phosphatidylcholine transfer protein involved in lipid metabolism; localizes to the nucleus | Silencing protein that interacts with Sir2p and Sir4p, and histone H3 and H4 tails, to establish a transcriptionally silent chromatin state; required for spreading of silenced chromatin; recruited to chromatin through interaction with Rap1p | CAAX box containing protein of unknown function, proposed to be involved in the RAS/cAMP signaling pathway | Putative protein of unknown function with some characteristics of a transcriptional activator; may be a target of Dbf2p-Mob1p kinase; GFP-fusion protein co-localizes with clathrin-coated vesicles; YML037C is not an essential gene | Largest subunit of the origin recognition complex, which directs DNA replication by binding to replication origins and is also involved in transcriptional silencing; exhibits ATPase activity | Component of the conserved oligomeric Golgi complex (Cog1p through Cog8p), a cytosolic tethering complex that functions in protein trafficking to mediate fusion of transport vesicles to Golgi compartments | A guanine nucleotide exchange factor involved in vesicle-mediated vacuolar protein transport; specifically stimulates the intrinsic guanine nucleotide exchange activity of Vps21p/Rab5: similar to mammalian ras inhibitors; binds ubiquitin | Protein that interacts with silencing proteins at the telomere, involved in transcriptional silencing; paralog of Zds1p | Copper chaperone for superoxide dismutase Sod1p, involved in oxidative stress protection; Met-X-Cys-X2-Cys motif within the N-terminal portion is involved in insertion of copper into Sod1p under conditions of copper deprivation | hypothetical protein | RNA-dependent ATPase in the DEAH-box family, required for activation of the spliceosome before the first transesterification step in RNA splicing | Topoisomerase I, nuclear enzyme that relieves torsional strain in DNA by cleaving and re-sealing the phosphodiester backbone; relaxes both positively and negatively supercoiled DNA; functions in replication, transcription, and recombination | Putative basic leucine zipper (bZIP) transcription factor | Catalytic subunit of TRAMP (Trf4/Pap2p-Mtr4p-Air1p/2p), a nuclear poly (A) polymerase complex involved in RNA quality control; catalyzes polyadenylation of unmodified tRNAs, and snoRNA and rRNA precursors; disputed role as a DNA polymerase | Ty1 enhancer activator required for full levels of Ty enhancer-mediated transcription; C6 zinc cluster DNA-binding protein | ATP-dependent RNA helicase, component of the mitochondrial degradosome along with the RNase Dss1p; the degradosome associates with the ribosome and mediates turnover of aberrant or unprocessed RNAs | Subunit of the tRNA splicing endonuclease, which is composed of Sen2p, Sen15p, Sen34p, and Sen54p | Subunit of TFIIH and nucleotide excision repair factor 3 complexes, involved in transcription initiation, required for nucleotide excision repair, similar to 52 kDa subunit of human TFIIH | UDP-GlcNAc-binding and catalytic subunit of the enzyme that mediates the first step in glycosylphosphatidylinositol (GPI) biosynthesis, mutations cause defects in transcription and in biogenesis of cell wall proteins | Core component of the signal recognition particle (SRP) ribonucleoprotein (RNP) complex that functions in targeting nascent secretory proteins to the endoplasmic reticulum (ER) membrane | Minus-end-directed microtubule motor that functions in mitosis and meiosis, localizes to the spindle pole body and localization is dependent on functional Cik1p, required for nuclear fusion during mating; potential Cdc28p substrate | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nuclear periphery; potential Cdc28p substrate | Second largest subunit of DNA polymerase II (DNA polymerase epsilon), required for normal yeast chromosomal replication; expression peaks at the G1/S phase boundary; potential Cdc28p substrate | Protein involved in exosome mediated 3' to 5' mRNA degradation and translation inhibition of non-poly(A) mRNAs; forms complex with Ski2p and Ski8p; required for repressing propagation of dsRNA viruses |
| 34 | 23 | 445 | -0.414 | 0.466 | 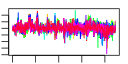 |  |  | 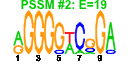 |  | YBR212W YDR148C YDR178W YDR254W YDR313C YGL010W YGR161C YGR243W YHR096C YIR016W YJL066C YJL103C YJL141C YJL142C YJL163C YKL171W YLL020C YLR149C YNL115C YNR001C YOR215C YEL012W YJL164C | NGR1 KGD2 SDH4 CHL4 PIB1 RTS3 FMP43 HXT5 MPM1 GSM1 YAK1 CIT1 AIM41 UBC8 TPK1 | RNA binding protein that negatively regulates growth rate; interacts with the 3' UTR of the mitochondrial porin (POR1) mRNA and enhances its degradation; overexpression impairs mitochondrial function; expressed in stationary phase | Dihydrolipoyl transsuccinylase, component of the mitochondrial alpha-ketoglutarate dehydrogenase complex, which catalyzes the oxidative decarboxylation of alpha-ketoglutarate to succinyl-CoA in the TCA cycle; phosphorylated | Membrane anchor subunit of succinate dehydrogenase (Sdh1p, Sdh2p, Sdh3p, Sdh4p), which couples the oxidation of succinate to the transfer of electrons to ubiquinone | Outer kinetochore protein required for chromosome stability, interacts with kinetochore proteins Ctf19p, Ctf3p, and Iml3p; exhibits a two-hybrid interaction with Mif2p; association with CEN DNA requires Ctf19p | RING-type ubiquitin ligase of the endosomal and vacuolar membranes, binds phosphatidylinositol(3)-phosphate; contains a FYVE finger domain | Putative protein of unknown function; YGL010W is not an essential gene | Putative component of the protein phosphatase type 2A complex | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Hexose transporter with moderate affinity for glucose, induced in the presence of non-fermentable carbon sources, induced by a decrease in growth rate, contains an extended N-terminal domain relative to other HXTs | Putative protein of unknown function; expression directly regulated by the metabolic and meiotic transcriptional regulator Ume6p; YIR016W is a non-essential gene | Mitochondrial membrane protein of unknown function, contains no hydrophobic stretches | Putative zinc cluster protein of unknown function; proposed to be involved in the regulation of energy metabolism, based on patterns of expression and sequence analysis | Serine-threonine protein kinase that is part of a glucose-sensing system involved in growth control in response to glucose availability; translocates from the cytoplasm to the nucleus and phosphorylates Pop2p in response to a glucose signal | Putative protein of unknown function | Putative protein of unknown function; predicted protein kinase; implicated in proteasome function; epitope-tagged protein localizes to the cytoplasm | Putative protein of unknown function; YLR149C is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YNL115C is not an essential gene | Citrate synthase, catalyzes the condensation of acetyl coenzyme A and oxaloacetate to form citrate; the rate-limiting enzyme of the TCA cycle; nuclear encoded mitochondrial protein | Putative protein of unknown function; the authentic protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays altered rate of mitochondrial loss and reduced growth rate in minimal glycerol media | Ubiquitin-conjugating enzyme that negatively regulates gluconeogenesis by mediating the glucose-induced ubiquitination of fructose-1,6-bisphosphatase (FBPase); cytoplasmic enzyme that catalyzes the ubiquitination of histones in vitro | cAMP-dependent protein kinase catalytic subunit; promotes vegetative growth in response to nutrients via the Ras-cAMP signaling pathway; inhibited by regulatory subunit Bcy1p in the absence of cAMP; partially redundant with Tpk2p and Tpk3p |
| 187 | 30 | 437 | -0.411 | 0.506 |  |  | 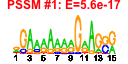 | 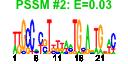 |  | YGR237C YGR255C YIL124W YJL102W YJR059W YLR299W YMR278W YOR018W YOR124C YOR132W YPR075C YGL095C YHL019C YJR091C YKL100C YKL129C YKR018C YLR248W YLR324W YML007W YMR139W YMR160W YMR304W YOL087C YOR035C YOR042W YOR113W YOR162C YOR227W YOR292C | COQ6 AYR1 MEF2 PTK2 ECM38 PGM3 ROD1 UBP2 VPS17 OPY2 VPS45 APM2 JSN1 MYO3 RCK2 PEX30 YAP1 RIM11 UBP15 SHE4 CUE5 AZF1 YRR1 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative flavin-dependent monooxygenase, involved in ubiquinone (Coenzyme Q) biosynthesis; localizes to the matrix face of the mitochondrial inner membrane in a large complex with other ubiquinone biosynthetic enzymes | NADPH-dependent 1-acyl dihydroxyacetone phosphate reductase found in lipid particles, ER, and mitochondrial outer membrane; involved in phosphatidic acid biosynthesis; required for spore germination; capable of metabolizing steroid hormones | Mitochondrial elongation factor involved in translational elongation | Putative serine/threonine protein kinase involved in regulation of ion transport across plasma membrane; enhances spermine uptake | Gamma-glutamyltranspeptidase, major glutathione-degrading enzyme; involved in detoxification of electrophilic xenobiotics; expression induced mainly by nitrogen starvation | Phosphoglucomutase, catalyzes interconversion of glucose-1-phosphate and glucose-6-phospate; transcription induced in response to stress; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; non-essential | Membrane protein; overexpression confers resistance to the GST substrate o-dinitrobenzene as well as to zinc and calcium; contains 2 PY motifs, which are required for Rod1p interaction with Rsp5p, a hect-type ubiquitin ligase | Ubiquitin-specific protease that removes ubiquitin from ubiquitinated proteins, cleaves at the C terminus of ubiquitin fusions; capable of cleaving polyubiquitin and possesses isopeptidase activity | Subunit of the membrane-associated retromer complex essential for endosome-to-Golgi retrograde protein transport; peripheral membrane protein that assembles onto the membrane with Vps5p to promote vesicle formation | Integral membrane protein that functions in the signaling branch of the high-osmolarity glycerol (HOG) pathway; interacts with Ste50p; overproduction blocks cell cycle arrest in the presence of mating pheromone | Protein of the Sec1p/Munc-18 family, essential for vacuolar protein sorting; required for the function of Pep12p and the early endosome/late Golgi SNARE Tlg2p; essential for fusion of Golgi-derived vesicles with the prevacuolar compartment | Protein of unknown function, homologous to the medium chain of mammalian clathrin-associated protein complex; involved in vesicular transport | Member of the Puf family of RNA-binding proteins, interacts with mRNAs encoding membrane-associated proteins; overexpression suppresses a tub2-150 mutation and causes increased sensitivity to benomyl in wild-type cells | Putative protein of unknown function with similarity to a human minor histocompatibility antigen; YKL100C is not an essential gene | One of two type I myosins; localizes to actin cortical patches; deletion of MYO3 has little effect on growth, but myo3 myo5 double deletion causes severe defects in growth and actin cytoskeleton organization | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Protein kinase involved in the response to oxidative and osmotic stress; identified as suppressor of S. pombe cell cycle checkpoint mutations | Peroxisomal integral membrane protein, involved in negative regulation of peroxisome number; partially functionally redundant with Pex31p; genetic interactions suggest action at a step downstream of steps mediated by Pex28p and Pex29p | Basic leucine zipper (bZIP) transcription factor required for oxidative stress tolerance; activated by H2O2 through the multistep formation of disulfide bonds and transit from the cytoplasm to the nucleus; mediates resistance to cadmium | Protein kinase required for signal transduction during entry into meiosis; promotes the formation of the Ime1p-Ume6p complex by phosphorylating Ime1p and Ume6p; shares similarity with mammalian glycogen synthase kinase 3-beta | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the membrane of the vacuole; mutant has enhanced sensitivity to overexpression of mutant huntingtin; YMR160W is not an essential gene | Ubiquitin-specific protease that may play a role in ubiquitin precursor processing | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; deletion mutant is sensitive to various chemicals including phenanthroline, sanguinarine, and nordihydroguaiaretic acid | Protein containing a UCS (UNC-45/CRO1/SHE4) domain, binds to myosin motor domains to regulate myosin function; involved in endocytosis, polarization of the actin cytoskeleton, and asymmetric mRNA localization | Protein containing a CUE domain that binds ubiquitin, which may facilitate intramolecular monoubiquitination; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Zinc-finger transcription factor, involved in induction of CLN3 transcription in response to glucose; genetic and physical interactions indicate a possible role in mitochondrial transcription or genome maintenance | Zn2-Cys6 zinc-finger transcription factor that activates genes involved in multidrug resistance; paralog of Yrm1p, acting on an overlapping set of target genes | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; YOR292C is not an essential gene |
| 93 | 27 | 439 | -0.385 | 0.481 |  |  |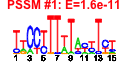  |  |  | YBR024W YBR273C YDR202C YEL005C YGL059W YGL104C YGL124C YGL185C YGR111W YIL017W YJL106W YJL155C YLR362W YNL025C YNL223W YNR007C YOR054C YOR227W YOR292C YPR149W YCL052C YDR330W YER101C YLR290C YMR302C YOR052C YPL152W | SCO2 UBX7 RAV2 VAB2 VPS73 MON1 VID28 IME2 FBP26 STE11 SSN8 ATG4 ATG3 VHS3 NCE102 PBN1 UBX5 AST2 YME2 YPL152W-A | Protein anchored to the mitochondrial inner membrane, similar to Sco1p and may have a redundant function with Sco1p in delivery of copper to cytochrome c oxidase; interacts with Cox2p | UBX (ubiquitin regulatory X) domain-containing protein that interacts with Cdc48p | Subunit of RAVE (Rav1p, Rav2p, Skp1p), a complex that associates with the V1 domain of the vacuolar membrane (H+)-ATPase (V-ATPase) and promotes assembly and reassembly of the holoenzyme | Protein with a potential role in vacuolar function, as suggested by its ability to bind Vac8p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Protein kinase; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Mitochondrial protein; mutation affects vacuolar protein sorting; putative transporter; member of the sugar porter family | Protein required for fusion of cvt-vesicles and autophagosomes with the vacuole; associates, as a complex with Ccz1p, with a perivacuolar compartment; potential Cdc28p substrate | Putative protein with sequence similarity to hydroxyacid dehydrogenases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Protein involved in proteasome-dependent catabolite degradation of fructose-1,6-bisphosphatase (FBPase); localized to the nucleus and the cytoplasm | Serine/threonine protein kinase involved in activation of meiosis, associates with Ime1p and mediates its stability, activates Ndt80p; IME2 expression is positively regulated by Ime1p | Fructose-2,6-bisphosphatase, required for glucose metabolism | Signal transducing MEK kinase involved in pheromone response and pseudohyphal/invasive growth pathways where it phosphorylates Ste7p, and the high osmolarity response pathway, via phosphorylation of Pbs2p; regulated by Ste20p and Ste50p | Cyclin-like component of the RNA polymerase II holoenzyme, involved in phosphorylation of the RNA polymerase II C-terminal domain; involved in glucose repression and telomere maintenance | Cysteine protease required for autophagy; cleaves Atg8p to a form required for autophagosome and Cvt vesicle generation; mediates attachment of autophagosomes to microtubules through interactions with Tub1p and Tub2p | E2-like enzyme involved in autophagy and the cytoplasm-to-vacuole targeting (Cvt) pathway; plays a role in formation of Atg8p-phosphatidylethanolamine conjugates, which are involved in membrane dynamics during autophagy and Cvt | Functionally redundant (see also SIS2) inhibitory subunit of Ppz1p, a PP1c-related ser/thr protein phosphatase Z isoform; synthetically lethal with sis2; putative phosphopantothenoylcysteine decarboxylase involved in coenzyme A biosynthesis | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; YOR292C is not an essential gene | Protein of unknown function; contains transmembrane domains; involved in secretion of proteins that lack classical secretory signal sequences; component of the detergent-insoluble glycolipid-enriched complexes (DIGs) | Essential component of glycosylphosphatidylinositol-mannosyltransferase I, required for the autocatalytic post-translational processing of the protease B precursor Prb1p, localizes to ER in lumenal orientation; homolog of mammalian PIG-X | Protein that may have a role in targeting of plasma membrane [H+]ATPase (Pma1p) to the plasma membrane, as suggested by analysis of genetic interactions | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YLR290C is not an essential gene | Integral inner mitochondrial membrane protein with a role in maintaining mitochondrial nucleoid structure and number; mutants exhibit an increased rate of mitochondrial DNA escape; shows some sequence similarity to exonucleases | Nuclear protein of unknown function; expression induced by nitrogen limitation in a GLN3, GAT1-independent manner and by weak acid | Identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching |
| 223 | 26 | 444 | -0.381 | 0.491 |  |  |  | 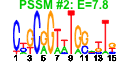 |  | YDR399W YKR056W YPL183C YBL035C YBR141C YCR058C YDR060W YDR527W YEL055C YFL023W YHR069C YHR070W YJL098W YKL014C YKR099W YLL035W YLR063W YLR336C YMR128W YMR185W YNL299W YOL125W YOR078W YOR101W YPL044C YPL068C | HPT1 TRM2 RTT10 POL12 PWP2 MAK21 RBA50 POL5 BUD27 RRP4 TRM5 SAP185 URB1 BAS1 GRC3 SGD1 ECM16 TRF5 TRM13 BUD21 RAS1 | Dimeric hypoxanthine-guanine phosphoribosyltransferase, catalyzes the formation of both inosine monophosphate and guanosine monophosphate; mutations in the human homolog HPRT1 can cause Lesch-Nyhan syndrome and Kelley-Seegmiller syndrome | tRNA methyltransferase, 5-methylates the uridine residue at position 54 of tRNAs and may also have a role in tRNA stabilization or maturation; endo-exonuclease with a role in DNA repair | Cytoplasmic protein of unknown function, plays a role in restricting Ty1 transposition | B subunit of DNA polymerase alpha-primase complex, required for initiation of DNA replication during mitotic and premeiotic DNA synthesis; also functions in telomere capping and length regulation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus; YBR141C is not an essential gene | Conserved 90S pre-ribosomal component essential for proper endonucleolytic cleavage of the 35 S rRNA precursor at A0, A1, and A2 sites; contains eight WD-repeats; PWP2 deletion leads to defects in cell cycle and bud morphogenesis | Constituent of 66S pre-ribosomal particles, required for large (60S) ribosomal subunit biogenesis; involved in nuclear export of pre-ribosomes; required for maintenance of dsRNA virus; homolog of human CAATT-binding protein | Protein involved in transcription; interacts with RNA polymerase II subunits Rpb2p, Rpb3, and Rpb11p; has similarity to human RPAP1 | DNA Polymerase phi; has sequence similarity to the human MybBP1A and weak sequence similarity to B-type DNA polymerases, not required for chromosomal DNA replication; required for the synthesis of rRNA | Protein involved in bud-site selection, nutrient signaling, and gene expression controlled by the TOR kinase; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | 3'-5' exoribonuclease involved in rRNA processing; component of the exosome 3->5 exonuclease complex with Rrp41p, Rrp42p, Rrp43p and Dis3p | tRNA(m(1)G37)methyltransferase, methylates a tRNA base adjacent to the anticodon that has a role in prevention of frameshifting; highly conserved across Archaea, Bacteria, and Eukarya | Protein that forms a complex with the Sit4p protein phosphatase and is required for its function; member of a family of similar proteins including Sap4p, Sap155p, and Sap190p | Nucleolar protein required for the normal accumulation of 25S and 5.8S rRNAs, associated with the 27SA2 pre-ribosomal particle; proposed to be involved in the biogenesis of the 60S ribosomal subunit | Myb-related transcription factor involved in regulating basal and induced expression of genes of the purine and histidine biosynthesis pathways; also involved in regulation of meiotic recombination at specific genes | Protein of unknown function, required for cell growth and possibly involved in rRNA processing; mRNA is cell cycle regulated | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR063W is not an essential gene | Essential nuclear protein with a possible role in the osmoregulatory glycerol response; interacts with phospholipase C (Plc1p); putative homolog of human NOM1 which is implicated in acute myeloid leukemia | Essential DEAH-box ATP-dependent RNA helicase specific to the U3 snoRNP, predominantly nucleolar in distribution, required for 18S rRNA synthesis | Putative protein of unknown function; essential gene required for viability | Poly (A) polymerase involved in nuclear RNA quality control based on: homology with Trf4p, genetic interactions with TRF4 mutants, physical interaction with Mtr4p (TRAMP subunit), and by direct assay; disputed role as a sigma DNA polymerase | 2'-O-methyltransferase responsible for modification of tRNA at position 4; exhibits no obvious similarity to other known methyltransferases | Component of small ribosomal subunit (SSU) processosome that contains U3 snoRNA; originally isolated as bud-site selection mutant that displays a random budding pattern | GTPase involved in G-protein signaling in the adenylate cyclase activating pathway, plays a role in cell proliferation; localized to the plasma membrane; homolog of mammalian RAS proto-oncogenes | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and is induced in response to the DNA-damaging agent MMS |
| 268 | 39 | 442 | -0.369 | 0.537 |  |  | 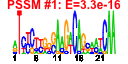 | 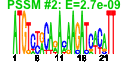 |  | YJR158W YBL029W YBR157C YBR298C YCR007C YCR098C YCR100C YDL149W YDL183C YDL244W YDL245C YEL069C YEL070W YER045C YFL053W YFL055W YFL058W YFL061W YGL081W YGR249W YHL008C YHR080C YHR105W YJL219W YJR149W YJR156C YJR159W YKL050C YKL107W YLR156W YLR159W YLR161W YNL270C YNL332W YNL335W YNR068C YNR072W YNR073C YOL154W | HXT16 ICS2 MAL31 GIT1 ATG9 THI13 HXT15 HXT13 DSF1 ACA1 DAK2 AGP3 THI5 DDI2 MGA1 YPT35 HXT9 THI11 SOR1 APL1 THI12 DDI3 HXT17 ZPS1 | Protein of unknown function with similarity to hexose transporter family members, expression is repressed by high levels of glucose | Non-essential protein of unknown function | Protein of unknown function; null mutation does not confer any obvious defects in growth, spore germination, viability, or carbohydrate utilization | Maltose permease, high-affinity maltose transporter (alpha-glucoside transporter); encoded in the MAL3 complex locus; member of the 12 transmembrane domain superfamily of sugar transporters; functional in genomic reference strain S288C | Putative integral membrane protein, member of DUP240 gene family; YCR007C is not an essential gene | Plasma membrane permease, mediates uptake of glycerophosphoinositol and glycerophosphocholine as sources of the nutrients inositol and phosphate; expression and transport rate are regulated by phosphate and inositol availability | Putative protein of unknown function | Transmembrane protein involved in formation of Cvt and autophagic vesicles; cycles between the pre-autophagosomal structure and other cytosolic punctate structures, not found in autophagosomes | Putative protein of unknown function; YDL183C is not an essential gene | Protein involved in synthesis of the thiamine precursor hydroxymethylpyrimidine (HMP); member of a subtelomeric gene family including THI5, THI11, THI12, and THI13 | Protein of unknown function with similarity to hexose transporter family members, expression is induced by low levels of glucose and repressed by high levels of glucose | Hexose transporter, induced in the presence of non-fermentable carbon sources, induced by low levels of glucose, repressed by high levels of glucose | Deletion suppressor of mpt5 mutation | Basic leucine zipper (bZIP) transcription factor of the ATF/CREB family, may regulate transcription of genes involved in utilization of non-optimal carbon sources | Dihydroxyacetone kinase, required for detoxification of dihydroxyacetone (DHA); involved in stress adaptation | Low-affinity amino acid permease, may act to supply the cell with amino acids as nitrogen source in nitrogen-poor conditions; transcription is induced under conditions of sulfur limitation | Protein whose expression is induced by DNA damage | Putative protein of unknown function; non-essential gene; interacts genetically with CHS5, a gene involved in chitin biosynthesis | Protein similar to heat shock transcription factor; multicopy suppressor of pseudohyphal growth defects of ammonium permease mutants | Putative protein of unknown function, does not appear to be involved in monocarboxylic acid transport; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Endosomal protein of unknown function that contains a phox (PX) homology domain and binds to both phosphatidylinositol-3-phosphate (PtdIns(3)P) and proteins involved in ER-Golgi or vesicular transport | Putative hexose transporter that is nearly identical to Hxt11p, has similarity to major facilitator superfamily (MFS) transporters, expression of HXT9 is regulated by transcription factors Pdr1p and Pdr3p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Sorbitol dehydrogenase; expression is induced in the presence of sorbitol | Protein of unknown function; the YKL050W protein is a target of the SCFCdc4 ubiquitin ligase complex and YKL050W transcription is regulated by Azf1p | Putative protein of unknown function; exhibits a two-hybrid interaction with Jsn1p in a large-scale analysis | Putative protein of unknown function; YLR156W, YLR159W, and YLR161W are three identical open reading frames encoded near the ribosomal DNA region of chromosome 12 | Beta-adaptin, large subunit of the clathrin associated protein complex (AP-2); involved in vesicle mediated transport; similar to mammalian beta-chain of the clathrin associated protein complex | hypothetical protein | Hexose transporter, up-regulated in media containing raffinose and galactose at pH 7.7 versus pH 4.7, repressed by high levels of glucose | Putative mannitol dehydrogenase | Putative GPI-anchored protein; transcription is induced under low-zinc conditions, as mediated by the Zap1p transcription factor, and at alkaline pH |
| 270 | 40 | 433 | -0.351 | 0.531 |  |  | 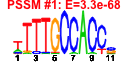 | 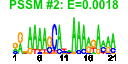 |  | YBL041W YBR173C YDL007W YDL097C YDL100C YDL126C YDL147W YDR394W YER012W YER094C YFR024C YFR050C YFR052W YGL011C YGL048C YGR135W YGR231C YGR253C YHL034C YJL001W YKL117W YKL145W YLR109W YLR387C YML028W YML092C YMR089C YMR092C YMR314W YNL079C YNL155W YOL038W YOR007C YOR157C YOR259C YOR261C YPL091W YPL240C YPR103W YPR108W | PRE7 UMP1 RPT2 RPN6 GET3 CDC48 RPN5 RPT3 PRE1 PUP3 LSB3 PRE4 RPN12 SCL1 RPT6 PRE9 PHB2 PUP2 SSB1 PRE3 SBA1 RPT1 AHP1 REH1 TSA1 PRE8 YTA12 AIP1 PRE5 TPM1 PRE6 SGT2 PUP1 RPT4 RPN8 GLR1 HSC82 PRE2 RPN7 | 20S proteasome beta-type subunit | Short-lived chaperone required for correct maturation of the 20S proteasome; may inhibit premature dimerization of proteasome half-mers; degraded by proteasome upon completion of its assembly | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; required for normal peptide hydrolysis by the core 20S particle | Essential, non-ATPase regulatory subunit of the 26S proteasome lid required for the assembly and activity of the 26S proteasome; the human homolog (S9 protein) partially rescues Rpn6p depletion | ATPase, subunit of the GET complex; required for the retrieval of HDEL proteins from the Golgi to the ER in an ERD2 dependent fashion; involved in resistance to heat and metal stress | ATPase in ER, nuclear membrane and cytosol with homology to mammalian p97; in a complex with Npl4p and Ufd1p participates in retrotranslocation of ubiquitinated proteins from the ER into the cytosol for degradation by the proteasome | Essential, non-ATPase regulatory subunit of the 26S proteasome lid, similar to mammalian p55 subunit and to another S. cerevisiae regulatory subunit, Rpn7p | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; substrate of N-acetyltransferase B | 20S proteasome beta-type subunit; localizes to the nucleus throughout the cell cycle | Beta subunit of the 20S proteasome involved in ubiquitin-dependent catabolism; human homolog is subunit C10 | Protein containing a C-terminal SH3 domain; binds Las17p, which is a homolog of human Wiskott-Aldrich Syndrome protein involved in actin patch assembly and actin polymerization | Subunit of the 19S regulatory particle of the 26S proteasome lid; synthetically lethal with RPT1, which is an ATPase component of the 19S regulatory particle; physically interacts with Nob1p and Rpn3p | Alpha subunit of the 20S core complex of the 26S proteasome involved in the degradation of ubiquitinated substrates; essential for growth; detected in the mitochondria | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; bound by ubiquitin-protein ligases Ubr1p and Ufd4p; localized mainly to the nucleus throughout the cell cycle | 20S proteasome beta-type subunit; the only nonessential 20S subunit | Subunit of the prohibitin complex (Phb1p-Phb2p), a 1.2 MDa ring-shaped inner mitochondrial membrane chaperone that stabilizes newly synthesized proteins; determinant of replicative life span; involved in mitochondrial segregation | Alpha subunit of the 20S proteasome involved in ubiquitin-dependent catabolism; human homolog is subunit zeta | Cytoplasmic ATPase that is a ribosome-associated molecular chaperone, functions with J-protein partner Zuo1p; may be involved in folding of newly-made polypeptide chains; member of the HSP70 family; interacts with phosphatase subunit Reg1p | 20S proteasome beta-type subunit, responsible for cleavage after acidic residues in peptides | Co-chaperone that binds to and regulates Hsp90 family chaperones; important for pp60v-src activity in yeast; homologous to the mammalian p23 proteins and like p23 can regulate telomerase activity | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; required for optimal CDC20 transcription; interacts with Rpn12p and the E3 ubiquitin-protein ligase Ubr1p | Thiol-specific peroxiredoxin, reduces hydroperoxides to protect against oxidative damage; function in vivo requires covalent conjugation to Urm1p | Protein of unknown function, similar to Rei1p but not involved in bud growth; contains dispersed C2H2 zinc finger domains | Ubiquitous housekeeping thioredoxin peroxidase, reduces reactive oxygen, nitrogen and sulfur species using thioredoxin as hydrogen donor; mediates redox regulation of the nuclear localization of Yap1p; deletion results in mutator phenotype | Component, with Afg3p, of the mitochondrial inner membrane m-AAA protease that mediates degradation of misfolded or unassembled proteins and is also required for correct assembly of mitochondrial enzyme complexes | Actin cortical patch component, interacts with the actin depolymerizing factor cofilin; required to restrict cofilin localization to cortical patches; contains WD repeats | 20S proteasome alpha-type subunit | Major isoform of tropomyosin; binds to and stabilizes actin cables and filaments, which direct polarized cell growth and the distribution of several organelles; acetylated by the NatB complex and acetylated form binds actin most efficiently | Putative protein of unknown function, contains DHHC domain, also predicted to have thiol-disulfide oxidoreductase active site | 20S proteasome alpha-type subunit; green fluorescent protein (GFP)-fusion protein relocates from the cytosol to the mitochondrial outer surface upon oxidative stress | Glutamine-rich cytoplasmic protein of unknown function; contains tetratricopeptide (TPR) repeats, which often mediate protein-protein interactions; has similarity to human SGT, which is a cochaperone that negatively regulates Hsp70 | Endopeptidase with trypsin-like activity that cleaves after basic residues; beta-type subunit of 20S proteasome synthesized as a proprotein before being proteolytically processed for assembly into 20S particle; human homolog is subunit Z | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; required for spindle pole body duplication; localized mainly to the nucleus throughout the cell cycle | Essential, non-ATPase regulatory subunit of the 26S proteasome; has similarity to the human p40 proteasomal subunit and to another S. cerevisiae regulatory subunit, Rpn11p | Cytosolic and mitochondrial glutathione oxidoreductase, converts oxidized glutathione to reduced glutathione | Cytoplasmic chaperone of the Hsp90 family, redundant in function and nearly identical with Hsp82p, and together they are essential; expressed constitutively at 10-fold higher basal levels than HSP82 and induced 2-3 fold by heat shock | 20S proteasome beta-type subunit, responsible for the chymotryptic activity of the proteasome | Essential, non-ATPase regulatory subunit of the 26S proteasome, similar to another S. cerevisiae regulatory subunit, Rpn5p, as well as to mammalian proteasome subunits |
| 122 | 32 | 448 | -0.343 | 0.537 |  |  | 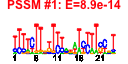 | 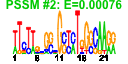 |  | YAL055W YBR033W YBR299W YCR098C YCR099C YCR100C YDR536W YEL069C YEL070W YFL055W YGR068C YGR290W YGR292W YHL008C YIL024C YJL137C YLL056C YLR281C YMR137C YMR195W YMR252C YMR317W YNR073C YPR015C YER185W YGR006W YHR213W YJL043W YJR021C YLR012C YMR017W YNL092W | PEX22 EDS1 MAL32 GIT1 STL1 HXT13 DSF1 AGP3 MAL12 GLG2 SNM1 ICY1 PUG1 PRP18 REC107 SPO20 | Putative peroxisomal membrane protein required for import of peroxisomal proteins, functionally complements a Pichia pastoris pex22 mutation | Putative zinc cluster protein; YBR033W is not an essential gene | Maltase (alpha-D-glucosidase), inducible protein involved in maltose catabolism; encoded in the MAL3 complex locus; functional in genomic reference strain S288C | Plasma membrane permease, mediates uptake of glycerophosphoinositol and glycerophosphocholine as sources of the nutrients inositol and phosphate; expression and transport rate are regulated by phosphate and inositol availability | Putative protein of unknown function | Glycerol proton symporter of the plasma membrane, subject to glucose-induced inactivation, strongly but transiently induced when cells are subjected to osmotic shock | Hexose transporter, induced in the presence of non-fermentable carbon sources, induced by low levels of glucose, repressed by high levels of glucose | Deletion suppressor of mpt5 mutation | Low-affinity amino acid permease, may act to supply the cell with amino acids as nitrogen source in nitrogen-poor conditions; transcription is induced under conditions of sulfur limitation | Putative protein of unknown function; YGR068C is not an essential gene. | Maltase (alpha-D-glucosidase), inducible protein involved in maltose catabolism; encoded in the MAL1 complex locus | Putative protein of unknown function, does not appear to be involved in monocarboxylic acid transport; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Putative protein of unknown function; non-essential gene; expression directly regulated by the metabolic and meiotic transcriptional regulator Ume6p | Self-glucosylating initiator of glycogen synthesis, also glucosylates n-dodecyl-beta-D-maltoside; similar to mammalian glycogenin | Putative protein of unknown function, transcription is activated by paralogous transcription factors Yrm1p and Yrr1p along with genes involved in pleiotropic drug resistance (PDR) phenomenon; YLL056C is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YLR281C is not an essential gene | Subunit of RNase MRP, which cleaves pre-rRNA and has a role in cell cycle-regulated degradation of daughter cell-specific mRNAs; binds to the NME1 RNA subunit of RNase MRP | Protein of unknown function, required for viability in rich media of cells lacking mitochondrial DNA; mutants have an invasive growth defect with elongated morphology; induced by amino acid starvation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YMR252C is not an essential gene | Putative protein of unknown function with some similarity to sialidase from Trypanosoma; YMR317W is not an essential gene | Putative mannitol dehydrogenase | Plasma membrane protein involved in protoporphyrin uptake; similar in sequence to RSB1, which is involved in sphingoid long-chain base release | Splicing factor involved in the positioning of the 3' splice site during the second catalytic step of splicing, part of snRNP U5, interacts with Slu7p | Putative protein of unknown function; YJL043W is a non-essential gene | Protein involved in early stages of meiotic recombination; involved in coordination between the initiation of recombination and the first division of meiosis; part of a complex (Rec107p-Mei4p-Rec114p) required for ds break formation | Putative protein of unknown function; YLR012C is not an essential gene | Meiosis-specific subunit of the t-SNARE complex, required for prospore membrane formation during sporulation; similar to but not functionally redundant with Sec9p; SNAP-25 homolog | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; YNL092W is not an essential gene |
| 298 | 30 | 434 | -0.326 | 0.505 |  |  | 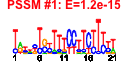 | 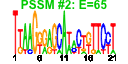 |  | YNL156C YAL049C YCL033C YCR083W YDR276C YDR512C YER035W YFR049W YGR209C YGR268C YIR036C YIR037W YJL015C YJL016W YJL017W YJL068C YJL151C YJL152W YJR085C YKL067W YKR046C YLL023C YLL027W YLR346C YOL071W YOR161C YOR215C YPL087W YPR098C YPR127W | NSG2 AIM2 TRX3 PMP3 EMI1 EDC2 YMR31 TRX2 HUA1 IRC24 HYR1 SNA3 YNK1 PET10 ISA1 EMI5 PNS1 AIM41 YDC1 | Protein involved in regulation of sterol biosynthesis; specifically stabilizes Hmg2p, one of two HMG-CoA isoenzymes that catalyze the rate-limiting step in sterol biosynthesis; homolog of mammalian INSIG proteins | Cytoplasmic protein of unknown function, potential Hsp82p interactor; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) | Putative protein-methionine-R-oxide reductase; involved in response to oxidative stress; similar to mouse Sepx1p and fly SelRp; YCL033C is not an essential gene | Mitochondrial thioredoxin, highly conserved oxidoreductase required to maintain the redox homeostasis of the cell, forms the mitochondrial thioredoxin system with Trr2p, redox state is maintained by both Trr2p and Glr1p | Small plasma membrane protein related to a family of plant polypeptides that are overexpressed under high salt concentration or low temperature, not essential for viability, deletion causes hyperpolarization of the plasma membrane potential | Non-essential protein of unknown function required for transcriptional induction of the early meiotic-specific transcription factor IME1, also required for sporulation | RNA-binding protein, activates mRNA decapping directly by binding to the mRNA substrate and enhancing the activity of the decapping proteins Dcp1p and Dcp2p | Mitochondrial ribosomal protein of the small subunit, has similarity to human mitochondrial ribosomal protein MRP-S36 | Cytoplasmic thioredoxin isoenzyme of the thioredoxin system which protects cells against oxidative and reductive stress, forms LMA1 complex with Pbi2p, acts as a cofactor for Tsa1p, required for ER-Golgi transport and vacuole inheritance | Cytoplasmic protein containing a zinc finger domain with sequence similarity to that of Type I J-proteins; computational analysis of large-scale protein-protein interaction data suggests a possible role in actin patch assembly | Putative benzil reductase;(GFP)-fusion protein localizes to the cytoplasm and is induced by the DNA-damaging agent MMS; sequence similarity with short-chain dehydrogenase/reductases; null mutant has increased spontaneous Rad52 foci | Thiol peroxidase that functions as a hydroperoxide receptor to sense intracellular hydroperoxide levels and transduce a redox signal to the Yap1p transcription factor | Putative protein of unknown function; GFP-fusion protein localizes to the cytoplasm; conserved in closely related Saccharomyces species | Non-essential intracellular esterase that can function as an S-formylglutathione hydrolase; may be involved in the detoxification of formaldehyde, which can be metabolized to S-formylglutathione; similar to human esterase D | Integral membrane protein localized to vacuolar intralumenal vesicles, computational analysis of large-scale protein-protein interaction data suggests a possible role in either cell wall synthesis or protein-vacuolar targeting | Putative protein of unknown function; GFP-fusion protein is induced in response to the DNA-damaging agent MMS; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Nucleoside diphosphate kinase, catalyzes the transfer of gamma phosphates from nucleoside triphosphates, usually ATP, to nucleoside diphosphates by a mechanism that involves formation of an autophosphorylated enzyme intermediate | Protein of unknown function that co-purifies with lipid particles; expression pattern suggests a role in respiratory growth; computational analysis of large-scale protein-protein interaction data suggests a role in ATP/ADP exchange | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum | Mitochondrial matrix protein involved in biogenesis of the iron-sulfur (Fe/S) cluster of Fe/S proteins, isa1 deletion causes loss of mitochondrial DNA and respiratory deficiency; depletion reduces growth on nonfermentable carbon sources | Putative protein of unknown function found in mitochondria; expression is regulated by transcription factors involved in pleiotropic drug resistance, Pdr1p and Yrr1p; YLR346C is not an essential gene | Protein of unknown function; has similarity to Torpedo californica tCTL1p, which is postulated to be a choline transporter, neither null mutation nor overexpression affects choline transport | Putative protein of unknown function; the authentic protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays altered rate of mitochondrial loss and reduced growth rate in minimal glycerol media | Alkaline dihydroceramidase, involved in sphingolipid metabolism; preferentially hydrolyzes dihydroceramide to a free fatty acid and dihydrosphingosine; has a minor reverse activity | Protein of unknown function, localized to the mitochondrial outer membrane | Putative protein of unknown function; expression is activated by transcription factor YRM1/YOR172W; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus |
| 66 | 39 | 436 | -0.323 | 0.530 | 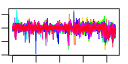 |  | 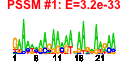 | 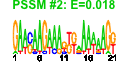 |  | YBR278W YCL067C YCR039C YCR096C YCR097W YDR181C YER029C YER030W YGL095C YGL153W YJR050W YKL023W YKL049C YKL079W YKR014C YKR023W YLR119W YLR218C YLR219W YLR309C YLR423C YLR431C YLR446W YML121W YMR009W YMR028W YMR029C YMR052W YMR193W YMR204C YNL224C YOL108C YOR069W YOR193W YOR256C YOR257W YPL055C YPR148C YPL107W | DPB3 HMLALPHA2 HMRA2 HMRA1 SAS4 SMB1 CHZ1 VPS45 PEX14 ISY1 CSE4 SMY1 YPT52 SRN2 MSC3 IMH1 ATG17 ATG23 GTR1 ADI1 TAP42 FAR8 FAR3 MRPL24 INP1 SQS1 INO4 VPS5 PEX27 TRE2 CDC31 LGE1 | Third-largest subunit of DNA polymerase II (DNA polymerase epsilon), required to maintain fidelity of chromosomal replication and also for inheritance of telomeric silencing; mRNA abundance peaks at the G1/S boundary of the cell cycle | Silenced copy of ALPHA2 at HML; homeobox-domain protein that associates with Mcm1p in haploid cells to repress a-specific gene expression and interacts with a1p in diploid cells to repress haploid-specific gene expression | Silenced copy of a2 at HMR; similarity to Alpha2p; required along with a1p for inhibiting expression of the HO endonuclease in a/alpha HO/HO diploid cells with an active mating-type interconversion system | Silenced copy of a1 at HMR; homeobox corepressor that interacts with Alpha2p to repress haploid-specific gene transcription in diploid cells | Subunit of the SAS complex (Sas2p, Sas4p, Sas5p), which acetylates free histones and nucleosomes and regulates transcriptional silencing; required for the HAT activity of Sas2p | Core Sm protein Sm B; part of heteroheptameric complex (with Smd1p, Smd2p, Smd3p, Sme1p, Smx3p, and Smx2p) that is part of the spliceosomal U1, U2, U4, and U5 snRNPs; homolog of human Sm B and Sm B' | Histone chaperone for Htz1p/H2A-H2B dimer; required for the stabilization of the Chz1p-Htz1-H2B complex; has overlapping function with Nap1p; null mutant displays weak sensitivity to MMS and benomyl; contains a highly conserved CHZ motif | Protein of the Sec1p/Munc-18 family, essential for vacuolar protein sorting; required for the function of Pep12p and the early endosome/late Golgi SNARE Tlg2p; essential for fusion of Golgi-derived vesicles with the prevacuolar compartment | Peroxisomal membrane peroxin that is a central component of the peroxisomal protein import machinery; interacts with both PTS1 (Pex5p) and PTS2 (Pex7p), peroxisomal matrix protein signal recognition factors and membrane receptor Pex13p | Component of the spliceosome complex involved in pre-mRNA splicing, auxiliary splicing factor that may modulate Syf1p activity and help optimize splicing; isy1 syf2 double mutation activates the spindle checkpoint, causing cell cycle arrest | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Centromere protein that resembles histones, required for proper kinetochore function; homolog of human CENP-A | Protein that interacts with Myo2p, proposed to be involved in exocytosis; N-terminal domain is related to the motor domain of kinesins | GTPase, similar to Ypt51p and Ypt53p and to mammalian rab5; required for vacuolar protein sorting and endocytosis | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Component of the ESCRT-I complex, which is involved in ubiquitin-dependent sorting of proteins into the endosome; suppressor of rna1-1 mutation; may be involved in RNA export from nucleus | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR218C is not essential; mutants exhibit glycogen storage defects and growth defects on a non-fermentable carbon source | Protein of unknown function, green fluorescent protein (GFP)-fusion protein localizes to the cell periphery; msc3 mutants are defective in directing meiotic recombination events to homologous chromatids; potential Cdc28p substrate | Protein involved in vesicular transport, mediates transport between an endosomal compartment and the Golgi, contains a Golgi-localization (GRIP) domain that interacts with activated Arl1p-GTP to localize Imh1p to the Golgi | Scaffold potein responsible for pre-autophagosomal structure organization; interacts with and is required for activation of Apg1p protein kinase; involved in autophagy but not in the Cvt (cytoplasm to vacuole targeting) pathway | Peripheral membrane protein, required for autophagy and for the cytoplasm-to-vacuole targeting (Cvt) pathway | Putative protein of unknown function with similarity to hexokinases; transcript is upregulated during sporulation and the unfolded protein response; YLR446W is not an essential gene | Cytoplasmic GTP binding protein and negative regulator, with homolog Gtr2p, of the Ran/Tc4 GTPase cycle; component of GSE complex, which is required for sorting of Gap1p; involved in phosphate transport; has homology to human RagA and RagB | Acireductone dioxygenease involved in the methionine salvage pathway; ortholog of human MTCBP-1; transcribed with YMR010W and regulated post-transcriptionally by RNase III (Rnt1p) cleavage; ADI1 mRNA is induced in heat shock conditions | Essential protein involved in the TOR signaling pathway; physically associates with the protein phosphatase 2A and the SIT4 protein phosphatase catalytic subunits | Protein involved in G1 cell cycle arrest in response to pheromone, in a pathway different from the Far1p-dependent pathway; interacts with Far3p, Far7p, Far9p, Far10p, and Far11p | Protein involved in G1 cell cycle arrest in response to pheromone, in a pathway different from the Far1p-dependent pathway; interacts with Far7p, Far8p, Far9p, Far10p, and Far11p | Mitochondrial ribosomal protein of the large subunit | Peripheral membrane protein of peroxisomes involved in peroxisomal inheritance | Protein of unknown function; overexpression antagonizes the suppression of splicing defects by spp382 mutants; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Transcription factor required for derepression of inositol-choline-regulated genes involved in phospholipid synthesis; forms a complex, with Ino2p, that binds the inositol-choline-responsive element through a basic helix-loop-helix domain | Nexin-1 homolog required for localizing membrane proteins from a prevacuolar/late endosomal compartment back to the late Golgi apparatus; structural component of the retromer membrane coat complex; forms a retromer subcomplex with Vps17p | Peripheral peroxisomal membrane protein involved in controlling peroxisome size and number, interacts with homologous protein Pex25p | Protein that functions with Tre1p to regulate ubiquitylation and vacuolar degradation of the metal transporter Smf1p; has similarity to transferrin receptors; inviability of null mutant in systematic studies is due to proximity to CDC31 | Calcium-binding component of the spindle pole body (SPB) half-bridge, required for SPB duplication in mitosis and meiosis II; homolog of mammalian centrin; interacts with Kar1p | Protein of unknown function; null mutant forms abnormally large cells | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YPL107W is not an essential gene |
| 125 | 46 | 431 | -0.322 | 0.496 | 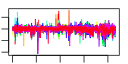 |  | 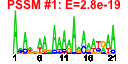 | 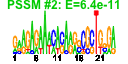 |  | YBL026W YCRX04W YDL157C YDR086C YGL101W YHR107C YHR167W YIL004C YIL016W YIR004W YJL179W YJR032W YJR054W YJR072C YJR076C YLR143W YLR145W YLR147C YLR148W YLR181C YLR210W YML094W YML096W YML097C YML105C YMR130W YMR321C YNL188W YOR159C YOR241W YOR269W YPL051W YPL235W YPR101W YER031C YER032W YHR201C YLR243W YMR091C YNL147W YNL153C YNL206C YOR253W YOR281C YOR305W YPR060C | LSM2 SSS1 CDC12 THP2 BET1 SNL1 DJP1 PFD1 CPR7 NPA3 CDC11 RMP1 SMD3 PEP3 VTA1 CLB4 GIM5 VPS9 SEC65 KAR1 IME2 MET7 PAC1 ARL3 RVB2 SNT309 YPT31 FIR1 PPX1 NPL6 LSM7 GIM3 RTT106 NAT5 PLP2 ARO7 | Lsm (Like Sm) protein; part of heteroheptameric complexes (Lsm2p-7p and either Lsm1p or 8p): cytoplasmic Lsm1p complex involved in mRNA decay; nuclear Lsm8p complex part of U6 snRNP and possibly involved in processing tRNA, snoRNA, and rRNA | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Subunit of the Sec61p translocation complex (Sec61p-Sss1p-Sbh1p) that forms a channel for passage of secretory proteins through the endoplasmic reticulum membrane, and of the Ssh1p complex (Ssh1p-Sbh2p-Sss1p); interacts with Ost4p and Wbp1p | Putative protein of unknown function; non-essential gene with similarity to YBR242W; interacts with the DNA helicase Hpr5p | Component of the septin ring of the mother-bud neck that is required for cytokinesis; septins recruit proteins to the neck and can act as a barrier to diffusion at the membrane, and they comprise the 10nm filaments seen with EM | Subunit of the THO complex, which connects transcription elongation and mitotic recombination, and of the TREX complex, which is recruited to activated genes and couples transcription to mRNA export; involved in telomere maintenance | Type II membrane protein required for vesicular transport between the endoplasmic reticulum and Golgi complex; v-SNARE with similarity to synaptobrevins | Protein of unknown function proposed to be involved in nuclear pore complex biogenesis and maintenance as well as protein folding; has similarity to the mammalian BAG-1 protein | Cytosolic J-domain-containing protein, required for peroxisomal protein import and involved in peroxisome assembly, homologous to E. coli DnaJ | Subunit of heterohexameric prefoldin, which binds cytosolic chaperonin and transfers target proteins to it; involved in the biogenesis of actin and of alpha- and gamma-tubulin | Peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; binds to Hsp82p and contributes to chaperone activity | Vacuolar protein of unknown function; potential Cdc28p substrate | Essential, conserved, cytoplasmic ATPase; phosphorylated by the Pcl1p-Pho85p kinase complex | Putative protein of unknown function; green fluorescent protein (GFP)-tagged protein localizes to the cytoplasm; YLR143W is not an essential gene | Subunit of RNase MRP, which processes pre-rRNA and has a role in cell cycle-regulated degradation of daughter cell-specific mRNAs; unlike most subunits, not shared between RNase MRP and nuclear RNase P | Core Sm protein Sm D3; part of heteroheptameric complex (with Smb1p, Smd1p, Smd2p, Sme1p, Smx3p, and Smx2p) that is part of the spliceosomal U1, U2, U4, and U5 snRNPs; homolog of human Sm D3 | Component of CORVET tethering complex; vacuolar peripheral membrane protein that promotes vesicular docking/fusion reactions in conjunction with SNARE proteins, required for vacuolar biogenesis | Multivesicular body (MVB) protein involved in endosomal protein sorting; regulates Vps4p activity by promoting its oligimeriztion; binds to Vps4p, Vps20p, and Vps60p; may act at a late step in MVB formation | B-type cyclin involved in cell cycle progression; activates Cdc28p to promote the G2/M transition; may be involved in DNA replication and spindle assembly; accumulates during S phase and G2, then targeted for ubiquitin-mediated degradation | Subunit of the heterohexameric cochaperone prefoldin complex which binds specifically to cytosolic chaperonin and transfers target proteins to it | Putative protein of unknown function with similarity to asparagine synthetases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YML096W is not an essential gene and partially overlaps the verified gene RAD10 | A guanine nucleotide exchange factor involved in vesicle-mediated vacuolar protein transport; specifically stimulates the intrinsic guanine nucleotide exchange activity of Vps21p/Rab5: similar to mammalian ras inhibitors; binds ubiquitin | Subunit of the signal recognition particle (SRP), involved in protein targeting to the ER; interacts with Srp54p; homolog of mammalian SRP19 | Putative protein of unknown function; YMR130W is not an essential gene | Putative protein of unknown function | Essential protein involved in karyogamy during mating and in spindle pole body duplication during mitosis, localizes to the half-bridge of the spindle pole body, interacts with Spc72p during karyogamy, also interacts with Cdc31p | Serine/threonine protein kinase involved in activation of meiosis, associates with Ime1p and mediates its stability, activates Ndt80p; IME2 expression is positively regulated by Ime1p | Folylpolyglutamate synthetase, catalyzes extension of the glutamate chains of the folate coenzymes, required for methionine synthesis and for maintenance of mitochondrial DNA | Protein involved in nuclear migration, part of the dynein/dynactin pathway; targets dynein to microtubule tips, which is necessary for sliding of microtubules along bud cortex; synthetic lethal with bni1; homolog of human LIS1 | GTPase of the Ras superfamily required to recruit Arl1p to the Golgi; similar to ADP-ribosylation factor | Essential protein involved in transcription regulation; component of chromatin remodeling complexes; required for assembly and function of the INO80 complex; member of the RUVB-like protein family | Component of NineTeen complex (NTC) containing Prp19p involved in mRNA splicing, interacts physically and genetically with Prp19p | GTPase of the Ypt/Rab family, very similar to Ypt32p; involved in the exocytic pathway; mediates intra-Golgi traffic or the budding of post-Golgi vesicles from the trans-Golgi | Protein involved in 3' mRNA processing, interacts with Ref2p; potential Cdc28p substrate | Exopolyphosphatase, hydrolyzes inorganic polyphosphate (poly P) into Pi residues; located in the cytosol, plasma membrane, and mitochondrial matrix | Putative protein of unknown function; YLR243W is an essential gene | Component of the RSC chromatin remodeling complex; interacts with Rsc3p, Rsc30p, Ldb7p, and Htl1p to form a module important for a broad range of RSC functions; involved in nuclear protein import and maintenance of proper telomere length | Protein with a role in regulation of Ty1 transposition | Subunit of the N-terminal acetyltransferase NatA (Nat1p, Ard1p, Nat5p); N-terminally acetylates many proteins, which influences multiple processes such as the cell cycle, heat-shock resistance, mating, sporulation, and telomeric silencing | Essential protein with similarity to phosducins, which are G-protein regulators; lethality of null mutation is functionally complemented by expression of mouse phosducin-like protein MgcPhLP | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the mitochondrion; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO); YOR305W is not an essential gene | Chorismate mutase, catalyzes the conversion of chorismate to prephenate to initiate the tyrosine/phenylalanine-specific branch of aromatic amino acid biosynthesis |
| 92 | 43 | 436 | -0.317 | 0.515 |  |  |  |  |  | YBL041W YBR173C YCL057W YDL007W YDL029W YDL097C YDL126C YDL147W YDR394W YER012W YER094C YFR010W YFR024C YFR050C YFR052W YGL011C YGL048C YGR048W YGR132C YGR135W YGR231C YGR253C YJL001W YKL117W YKL145W YKL210W YLR387C YML007W YML092C YMR314W YNL155W YOL038W YOR007C YOR157C YOR259C YOR261C YPR103W YPR108W YBL058W YBR170C YIL075C YMR186W YOR117W | PRE7 UMP1 PRD1 RPT2 ARP2 RPN6 CDC48 RPN5 RPT3 PRE1 PUP3 UBP6 LSB3 PRE4 RPN12 SCL1 RPT6 UFD1 PHB1 PRE9 PHB2 PUP2 PRE3 SBA1 RPT1 UBA1 REH1 YAP1 PRE8 PRE5 PRE6 SGT2 PUP1 RPT4 RPN8 PRE2 RPN7 SHP1 NPL4 RPN2 HSC82 RPT5 | 20S proteasome beta-type subunit | Short-lived chaperone required for correct maturation of the 20S proteasome; may inhibit premature dimerization of proteasome half-mers; degraded by proteasome upon completion of its assembly | Zinc metalloendopeptidase, found in the cytoplasm and intermembrane space of mitochondria; with Cym1p, involved in degradation of mitochondrial proteins and of presequence peptides cleaved from imported proteins | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; required for normal peptide hydrolysis by the core 20S particle | Essential component of the Arp2/3 complex, which is a highly conserved actin nucleation center required for the motility and integrity of actin patches; involved in endocytosis and membrane growth and polarity | Essential, non-ATPase regulatory subunit of the 26S proteasome lid required for the assembly and activity of the 26S proteasome; the human homolog (S9 protein) partially rescues Rpn6p depletion | ATPase in ER, nuclear membrane and cytosol with homology to mammalian p97; in a complex with Npl4p and Ufd1p participates in retrotranslocation of ubiquitinated proteins from the ER into the cytosol for degradation by the proteasome | Essential, non-ATPase regulatory subunit of the 26S proteasome lid, similar to mammalian p55 subunit and to another S. cerevisiae regulatory subunit, Rpn7p | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; substrate of N-acetyltransferase B | 20S proteasome beta-type subunit; localizes to the nucleus throughout the cell cycle | Beta subunit of the 20S proteasome involved in ubiquitin-dependent catabolism; human homolog is subunit C10 | Ubiquitin-specific protease situated in the base subcomplex of the 26S proteasome, releases free ubiquitin from branched polyubiquitin chains; deletion causes hypersensitivity to cycloheximide and other toxic compounds | Protein containing a C-terminal SH3 domain; binds Las17p, which is a homolog of human Wiskott-Aldrich Syndrome protein involved in actin patch assembly and actin polymerization | Subunit of the 19S regulatory particle of the 26S proteasome lid; synthetically lethal with RPT1, which is an ATPase component of the 19S regulatory particle; physically interacts with Nob1p and Rpn3p | Alpha subunit of the 20S core complex of the 26S proteasome involved in the degradation of ubiquitinated substrates; essential for growth; detected in the mitochondria | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; bound by ubiquitin-protein ligases Ubr1p and Ufd4p; localized mainly to the nucleus throughout the cell cycle | Protein that interacts with Cdc48p and Npl4p, involved in recognition of polyubiquitinated proteins and their presentation to the 26S proteasome for degradation; involved in transporting proteins from the ER to the cytosol | Subunit of the prohibitin complex (Phb1p-Phb2p), a 1.2 MDa ring-shaped inner mitochondrial membrane chaperone that stabilizes newly synthesized proteins; determinant of replicative life span; involved in mitochondrial segregation | 20S proteasome beta-type subunit; the only nonessential 20S subunit | Alpha subunit of the 20S proteasome involved in ubiquitin-dependent catabolism; human homolog is subunit zeta | 20S proteasome beta-type subunit, responsible for cleavage after acidic residues in peptides | Co-chaperone that binds to and regulates Hsp90 family chaperones; important for pp60v-src activity in yeast; homologous to the mammalian p23 proteins and like p23 can regulate telomerase activity | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; required for optimal CDC20 transcription; interacts with Rpn12p and the E3 ubiquitin-protein ligase Ubr1p | Ubiquitin activating enzyme (E1), involved in ubiquitin-mediated protein degradation and essential for viability | Protein of unknown function, similar to Rei1p but not involved in bud growth; contains dispersed C2H2 zinc finger domains | Basic leucine zipper (bZIP) transcription factor required for oxidative stress tolerance; activated by H2O2 through the multistep formation of disulfide bonds and transit from the cytoplasm to the nucleus; mediates resistance to cadmium | 20S proteasome alpha-type subunit | Putative protein of unknown function, contains DHHC domain, also predicted to have thiol-disulfide oxidoreductase active site | 20S proteasome alpha-type subunit; green fluorescent protein (GFP)-fusion protein relocates from the cytosol to the mitochondrial outer surface upon oxidative stress | Glutamine-rich cytoplasmic protein of unknown function; contains tetratricopeptide (TPR) repeats, which often mediate protein-protein interactions; has similarity to human SGT, which is a cochaperone that negatively regulates Hsp70 | Endopeptidase with trypsin-like activity that cleaves after basic residues; beta-type subunit of 20S proteasome synthesized as a proprotein before being proteolytically processed for assembly into 20S particle; human homolog is subunit Z | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; required for spindle pole body duplication; localized mainly to the nucleus throughout the cell cycle | Essential, non-ATPase regulatory subunit of the 26S proteasome; has similarity to the human p40 proteasomal subunit and to another S. cerevisiae regulatory subunit, Rpn11p | 20S proteasome beta-type subunit, responsible for the chymotryptic activity of the proteasome | Essential, non-ATPase regulatory subunit of the 26S proteasome, similar to another S. cerevisiae regulatory subunit, Rpn5p, as well as to mammalian proteasome subunits | UBX (ubiquitin regulatory X) domain-containing protein that regulates Glc7p phosphatase activity and interacts with Cdc48p; interacts with ubiquitylated proteins in vivo and is required for degradation of a ubiquitylated model substrate | Endoplasmic reticulum and nuclear membrane protein, forms a complex with Cdc48p and Ufd1p that recognizes ubiquitinated proteins in the endoplasmic reticulum and delivers them to the proteasome for degradation | Subunit of the 26S proteasome, substrate of the N-acetyltransferase Nat1p | Cytoplasmic chaperone of the Hsp90 family, redundant in function and nearly identical with Hsp82p, and together they are essential; expressed constitutively at 10-fold higher basal levels than HSP82 and induced 2-3 fold by heat shock | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; recruited to the GAL1-10 promoter region upon induction of transcription |
| 130 | 38 | 438 | -0.312 | 0.506 |  |  | 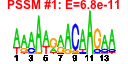 | 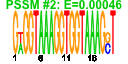 |  | YDR302W YEL040W YHR026W YHR143W YIL118W YIL123W YJR105W YML052W YPL050C YPR053C YPR119W YBL002W YBL003C YBL032W YBR009C YBR010W YBR158W YDL055C YDL179W YDR224C YDR225W YER124C YGL021W YGL028C YGL101W YGR234W YIL158W YLR049C YLR286C YNL030W YNL031C YNL066W YNL111C YNL327W YNR067C YOR248W YOR264W YOR315W | GPI11 UTR2 IPP1 DSE2 RHO3 SIM1 ADO1 SUR7 MNN9 CLB2 HTB2 HTA2 HEK2 HHF1 HHT1 AMN1 PSA1 PCL9 HTB1 HTA1 DSE1 ALK1 SCW11 YHB1 AIM20 CTS1 HHF2 HHT2 SUN4 CYB5 EGT2 DSE4 DSE3 SFG1 | ER membrane protein involved in a late step of glycosylphosphatidylinositol (GPI) anchor assembly; involved in the addition of phosphoethanolamine to the multiply mannosylated GPI intermediate; human PIG-Fp is a functional homolog | Cell wall protein that functions in the transfer of chitin to beta(1-6)glucan; putative chitin transglycosidase; glycosylphosphatidylinositol (GPI)-anchored protein localized to the bud neck; has a role in cell wall maintenance | Cytoplasmic inorganic pyrophosphatase (PPase), catalyzes the rapid exchange of oxygens from Pi with water, highly expressed and essential for viability, active-site residues show identity to those from E. coli PPase | Daughter cell-specific secreted protein with similarity to glucanases, degrades cell wall from the daughter side causing daughter to separate from mother; expression is repressed by cAMP | Non-essential small GTPase of the Rho/Rac subfamily of Ras-like proteins involved in the establishment of cell polarity; GTPase activity positively regulated by the GTPase activating protein (GAP) Rgd1p | Protein of the SUN family (Sim1p, Uth1p, Nca3p, Sun4p) that may participate in DNA replication, promoter contains SCB regulation box at -300 bp indicating that expression may be cell cycle-regulated | Adenosine kinase, required for the utilization of S-adenosylmethionine (AdoMet); may be involved in recycling adenosine produced through the methyl cycle | Putative integral membrane protein; component of eisosomes; associated with endocytosis, along with Pil1p and Lsp1p; sporulation and plasma membrane sphingolipid content are altered in mutants | Subunit of Golgi mannosyltransferase complex also containing Anp1p, Mnn10p, Mnn11p, and Hoc1p that mediates elongation of the polysaccharide mannan backbone; forms a separate complex with Van1p that is also involved in backbone elongation | B-type cyclin involved in cell cycle progression; activates Cdc28p to promote the transition from G2 to M phase; accumulates during G2 and M, then targeted via a destruction box motif for ubiquitin-mediated degradation by the proteasome | One of two nearly identical (see HTB1) histone H2B subtypes required for chromatin assembly and chromosome function; Rad6p-Bre1p-Lge1p mediated ubiquitination regulates transcriptional activation, meiotic DSB formation and H3 methylation | One of two nearly identical (see also HTA1) histone H2A subtypes; core histone required for chromatin assembly and chromosome function; DNA damage-dependent phosphorylation by Mec1p facilitates DNA repair; acetylated by Nat4p | RNA binding protein with similarity to hnRNP-K that localizes to the cytoplasm and to subtelomeric DNA; required for the proper localization of ASH1 mRNA; involved in the regulation of telomere position effect and telomere length | One of two identical histone H4 proteins (see also HHF2); core histone required for chromatin assembly and chromosome function; contributes to telomeric silencing; N-terminal domain involved in maintaining genomic integrity | One of two identical histone H3 proteins (see also HHT2); core histone required for chromatin assembly, involved in heterochromatin-mediated telomeric and HM silencing; regulated by acetylation, methylation, and mitotic phosphorylation | Protein required for daughter cell separation, multiple mitotic checkpoints, and chromosome stability; contains 12 degenerate leucine-rich repeat motifs; expression is induced by the Mitotic Exit Network (MEN) | GDP-mannose pyrophosphorylase (mannose-1-phosphate guanyltransferase), synthesizes GDP-mannose from GTP and mannose-1-phosphate in cell wall biosynthesis; required for normal cell wall structure | Cyclin, forms a functional kinase complex with Pho85p cyclin-dependent kinase (Cdk), expressed in late M/early G1 phase, activated by Swi5p | One of two nearly identical (see HTB2) histone H2B subtypes required for chromatin assembly and chromosome function; Rad6p-Bre1p-Lge1p mediated ubiquitination regulates transcriptional activation, meiotic DSB formation and H3 methylation | One of two nearly identical (see also HTA2) histone H2A subtypes; core histone required for chromatin assembly and chromosome function; DNA damage-dependent phosphorylation by Mec1p facilitates DNA repair; acetylated by Nat4p | Daughter cell-specific protein, may participate in pathways regulating cell wall metabolism; deletion affects cell separation after division and sensitivity to drugs targeted against the cell wall | Protein kinase; accumulation and phosphorylation are periodic during the cell cycle; phosphorylated in response to DNA damage; contains characteristic motifs for degradation via the APC pathway; similar to Alk2p and to mammalian haspins | Cell wall protein with similarity to glucanases; may play a role in conjugation during mating based on its regulation by Ste12p | Putative protein of unknown function; non-essential gene with similarity to YBR242W; interacts with the DNA helicase Hpr5p | Nitric oxide oxidoreductase, flavohemoglobin involved in nitric oxide detoxification; plays a role in the oxidative and nitrosative stress responses | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Putative protein of unknown function | Endochitinase, required for cell separation after mitosis; transcriptional activation during late G and early M cell cycle phases is mediated by transcription factor Ace2p | One of two identical histone H4 proteins (see also HHF1); core histone required for chromatin assembly and chromosome function; contributes to telomeric silencing; N-terminal domain involved in maintaining genomic integrity | One of two identical histone H3 proteins (see also HHT1); core histone required for chromatin assembly, involved in heterochromatin-mediated telomeric and HM silencing; regulated by acetylation, methylation, and mitotic phosphorylation | Cell wall protein related to glucanases, possibly involved in cell wall septation; member of the SUN family | Cytochrome b5, involved in the sterol and lipid biosynthesis pathways; required for sterol C5-6 and fatty acid desaturation | Glycosylphosphatidylinositol (GPI)-anchored cell wall endoglucanase required for proper cell separation after cytokinesis, expression is activated by Swi5p and tightly regulated in a cell cycle-dependent manner | Daughter cell-specific secreted protein with similarity to glucanases, degrades cell wall from the daughter side causing daughter to separate from mother | Daughter cell-specific protein, may help establish daughter fate | Nuclear protein, putative transcription factor required for growth of superficial pseudohyphae (which do not invade the agar substrate) but not for invasive pseudohyphal growth; may act together with Phd1p; potential Cdc28p substrate |
| 46 | 29 | 441 | -0.305 | 0.471 |  |  | 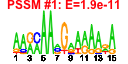 | 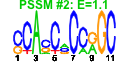 |  | YBR298C YDR247W YDR277C YER101C YFL021W YFL052W YGL193C YGR046W YGR066C YGR143W YGR289C YIL050W YIL056W YLR297W YMR056C YMR081C YMR133W YMR148W YNL144C YPL109C YPL147W YPR098C YGL192W YGL218W YGR023W YGR288W YKL171W YPL054W YPR150W | MAL31 VHS1 MTH1 AST2 GAT1 TAM41 SKN1 MAL11 PCL7 VHR1 AAC1 ISF1 REC114 PXA1 IME4 MTL1 MAL13 LEE1 | Maltose permease, high-affinity maltose transporter (alpha-glucoside transporter); encoded in the MAL3 complex locus; member of the 12 transmembrane domain superfamily of sugar transporters; functional in genomic reference strain S288C | Cytoplasmic serine/threonine protein kinase; identified as a high-copy suppressor of the synthetic lethality of a sis2 sit4 double mutant, suggesting a role in G1/S phase progression; homolog of Sks1p | Negative regulator of the glucose-sensing signal transduction pathway, required for repression of transcription by Rgt1p; interacts with Rgt1p and the Snf3p and Rgt2p glucose sensors; phosphorylated by Yck1p, triggering Mth1p degradation | Protein that may have a role in targeting of plasma membrane [H+]ATPase (Pma1p) to the plasma membrane, as suggested by analysis of genetic interactions | Transcriptional activator of genes involved in nitrogen catabolite repression; contains a GATA-1-type zinc finger DNA-binding motif; activity and localization regulated by nitrogen limitation and Ure2p | Putative zinc cluster protein that contains a DNA binding domain; null mutant sensitive to calcofluor white, low osmolarity and heat, suggesting a role for YFL052Wp in cell wall integrity | Haploid-specific gene repressed by a1-alpha2, turned off in sir3 null strains, absence enhances the sensitivity of rad52-327 cells to campothecin almost 100-fold | Mitochondrial protein involved in protein import into the mitochondrial matrix; maintains the functional integrity of the TIM23 protein translocator complex; viability of null mutant is strain-dependent; mRNA is targeted to the bud | Putative protein of unknown function | Protein involved in sphingolipid biosynthesis; type II membrane protein with similarity to Kre6p | Maltose permease, inducible high-affinity maltose transporter (alpha-glucoside transporter); encoded in the MAL1 complex locus; member of the 12 transmembrane domain superfamily of sugar transporters | Pho85p cyclin of the Pho80p subfamily, forms a functional kinase complex with Pho85p which phosphorylates Mmr1p and is regulated by Pho81p; involved in glycogen metabolism, expression is cell-cycle regulated | Transcriptional activator, required for the vitamin H-responsive element (VHRE) mediated induction of VHT1 (Vitamin H transporter) and BIO5 (biotin biosynthesis intermediate transporter) in response to low biotin concentrations | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; YLR297W is not an essential gene | Mitochondrial inner membrane ADP/ATP translocator, exchanges cytosolic ADP for mitochondrially synthesized ATP; phosphorylated; Aac1p is a minor isoform while Pet9p is the major ADP/ATP translocator | Serine-rich, hydrophilic protein with similarity to Mbr1p; overexpression suppresses growth defects of hap2, hap3, and hap4 mutants; expression is under glucose control; cotranscribed with NAM7 in a cyp1 mutant | Protein involved in early stages of meiotic recombination; possibly involved in the coordination of recombination and meiotic division; mutations lead to premature initiation of the first meiotic division | Putative protein of unknown function; predicted to contain a transmembrane domain; YMR148W is not an essential gene | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YNL144C is not an essential gene | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Subunit of a heterodimeric peroxisomal ATP-binding cassette transporter complex (Pxa1p-Pxa2p), required for import of long-chain fatty acids into peroxisomes; similarity to human adrenoleukodystrophy transporter and ALD-related proteins | Protein of unknown function, localized to the mitochondrial outer membrane | Probable mRNA N6-adenosine methyltransferase that is required for IME1 transcript accumulation and for sporulation; expression is induced in starved MATa/MAT alpha diploid cells | Protein with both structural and functional similarity to Mid2p, which is a plasma membrane sensor required for cell integrity signaling during pheromone-induced morphogenesis; suppresses rgd1 null mutations | MAL-activator protein, part of complex locus MAL1; nonfunctional in genomic reference strain S288C | Putative protein of unknown function; predicted protein kinase; implicated in proteasome function; epitope-tagged protein localizes to the cytoplasm | Zinc-finger protein of unknown function |
| 220 | 30 | 447 | -0.263 | 0.559 |  |  | 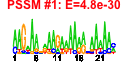 |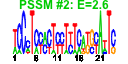  |  | YCR038C YGR249W YHR202W YIL071C YIL071W YJL042W YNL012W YPL087W YCL068C YDR275W YEL057C YER038C YER096W YER098W YER158C YGL205W YGR066C YGR146C YIL042C YJL132W YJR035W YLL025W YLR128W YLR218C YLR219W YMR020W YMR155W YMR182C YNL077W YPL109C | BUD5 MGA1 YIH1 PCI8 MHP1 SPO1 YDC1 BSC2 KRE29 SHC1 UBP9 POX1 PKP1 RAD26 PAU17 DCN1 MSC3 FMS1 RGM1 APJ1 | GTP/GDP exchange factor for Rsr1p (Bud1p) required for both axial and bipolar budding patterns; mutants exhibit random budding in all cell types | Protein similar to heat shock transcription factor; multicopy suppressor of pseudohyphal growth defects of ammonium permease mutants | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole, while HA-tagged protein is found in the soluble fraction, suggesting cytoplasmic localization | Protein that inhibits activation of Gcn2p, an eIF2 alpha subunit protein kinase, by competing for Gcn1p binding, thus impacting gene expression in response to starvation; has sequence and functional similarity to the mouse IMPACT gene | Possible shared subunit of Cop9 signalosome (CSN) and eIF3, binds eIF3b subunit Prt1p, has possible dual functions in transcriptional and translational control, contains a PCI (Proteasome-COP9 signalosome (CSN)-eIF3) domain | Microtubule-associated protein involved in assembly and stabilization of microtubules; overproduction results in cell cycle arrest at G2 phase; similar to Drosophila protein MAP and to mammalian MAP4 proteins | Meiosis-specific protein with similarity to phospholipase B, required for meiotic spindle pole body duplication and separation; required for spore formation | Alkaline dihydroceramidase, involved in sphingolipid metabolism; preferentially hydrolyzes dihydroceramide to a free fatty acid and dihydrosphingosine; has a minor reverse activity | Putative protein of unknown function | Protein of unknown function, ORF exhibits genomic organization compatible with a translational readthrough-dependent mode of expression | Protein of unknown function involved in telomere maintenance; target of UME6 regulation | Essential subunit of the Mms21-Smc5-Smc6 complex; protein of unknown function; required for growth and DNA repair; heterozygous mutant shows haploinsufficiency in K1 killer toxin resistance | Sporulation-specific activator of Chs3p (chitin synthase III), required for the synthesis of the chitosan layer of ascospores; has similarity to Skt5p, which activates Chs3p during vegetative growth; transcriptionally induced at alkaline pH | Ubiquitin carboxyl-terminal hydrolase, ubiquitin-specific protease that cleaves ubiquitin-protein fusions | Protein of unknown function, has similarity to Afr1p; potentially phosphorylated by Cdc28p | Fatty-acyl coenzyme A oxidase, involved in the fatty acid beta-oxidation pathway; localized to the peroxisomal matrix | Putative protein of unknown function; induced by iron homeostasis transcription factor Aft2p; multicopy suppressor of a temperature sensitive hsf1 mutant | Mitochondrial kinase, phosphorylates pyruvate dehydrogenase alpha subunit Pda1p | Putative protein of unknown function; localizes to the membrane fraction; possible Zap1p-regulated target gene induced by zinc deficiency; YJL132W is a non-essential gene | Protein involved in transcription-coupled repair nucleotide excision repair of UV-induced DNA lesions; homolog of human CSB protein | Putative protein of unknown function; YLL025W is not an essential gene | Putative Nedd8 ligase; binds Nedd8; involved in cullin neddylation; not essential; similar to C.elegans DCN-1; contains UBA-like ubiquitin-binding domain and a DUF298 domain | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR218C is not essential; mutants exhibit glycogen storage defects and growth defects on a non-fermentable carbon source | Protein of unknown function, green fluorescent protein (GFP)-fusion protein localizes to the cell periphery; msc3 mutants are defective in directing meiotic recombination events to homologous chromatids; potential Cdc28p substrate | Polyamine oxidase, converts spermine to spermidine, which is required for the essential hypusination modification of translation factor eIF-5A; also involved in pantothenic acid biosynthesis | Putative protein of unknown function; identified as interacting with Hsp82p in a high-throughput two-hybrid screen | Putative transcriptional repressor with proline-rich zinc fingers; overproduction impairs cell growth | Putative chaperone of the HSP40 (DNAJ) family; overexpression interferes with propagation of the [Psi+] prion; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
| 264 | 30 | 434 | -0.214 | 0.514 |  |  | 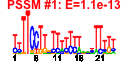 | 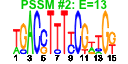 |  | YER054C YGR287C YHR171W YBL101C YCR024C YCR029C YCR030C YDL025C YDR244W YDR277C YDR293C YDR423C YHR082C YIL036W YIL071C YIL071W YIR007W YKL034W YKL062W YKL121W YKL124W YKR089C YLL029W YMR008C YMR041C YMR136W YMR148W YNL025C YNL125C YPR003C | GIP2 ATG7 ECM21 SLM5 SYP1 PEX5 MTH1 SSD1 CAD1 KSP1 CST6 YIH1 PCI8 TUL1 MSN4 SSH4 TGL4 RUP2 PLB1 ARA2 SCT1 SSN8 ESBP6 | Putative regulatory subunit of the protein phosphatase Glc7p, involved in glycogen metabolism; contains a conserved motif (GVNK motif) that is also found in Gac1p, Pig1p, and Pig2p | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; has similarity to alpha-D-glucosidase (maltase); authentic, non-tagged protein detected in purified mitochondria in high-throughput studies | Autophagy-related protein and dual specificity member of the E1 family of ubiquitin-activating enzymes; mediates the conjugation of Atg12p with Atg5p and Atg8p with phosphatidylethanolamine, required steps in autophagosome formation | Non-essential protein of unknown function; promoter contains several Gcn4p binding elements | Mitochondrial asparaginyl-tRNA synthetase | Protein with a potential role in actin cytoskeletal organization; overexpression suppresses a pfy1 (profilin) null mutation | Putative protein kinase, potentially phosphorylated by Cdc28p; YDL025C is not an essential gene | Peroxisomal membrane signal receptor for the C-terminal tripeptide signal sequence (PTS1) of peroxisomal matrix proteins, required for peroxisomal matrix protein import; also proposed to have PTS1-receptor independent functions | Negative regulator of the glucose-sensing signal transduction pathway, required for repression of transcription by Rgt1p; interacts with Rgt1p and the Snf3p and Rgt2p glucose sensors; phosphorylated by Yck1p, triggering Mth1p degradation | Protein with a role in maintenance of cellular integrity, interacts with components of the TOR pathway; ssd1 mutant of a clinical S. cerevisiae strain displays elevated virulence | AP-1-like bZIP transcriptional activator involved in stress responses, iron metabolism, and pleiotropic drug resistance; controls a set of genes involved in stabilizing proteins; binds consensus sequence TTACTAA; 5' UTR contains uORFs | Nonessential putative serine/threonine protein kinase of unknown cellular role; overproduction causes allele-specific suppression of the prp20-10 mutation | Basic leucine zipper (bZIP) transcription factor of the ATF/CREB family, activates transcription of genes involved in utilization of non-optimal carbon sources; involved in telomere maintenance | Protein that inhibits activation of Gcn2p, an eIF2 alpha subunit protein kinase, by competing for Gcn1p binding, thus impacting gene expression in response to starvation; has sequence and functional similarity to the mouse IMPACT gene | Possible shared subunit of Cop9 signalosome (CSN) and eIF3, binds eIF3b subunit Prt1p, has possible dual functions in transcriptional and translational control, contains a PCI (Proteasome-COP9 signalosome (CSN)-eIF3) domain | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YIR007W is a non-essential gene | Golgi-localized RING-finger ubiquitin ligase (E3), involved in ubiquitinating and sorting membrane proteins that contain polar transmembrane domains to multivesicular bodies for delivery to the vacuole for quality control purposes | Transcriptional activator related to Msn2p; activated in stress conditions, which results in translocation from the cytoplasm to the nucleus; binds DNA at stress response elements of responsive genes, inducing gene expression | Putative protein of unknown function | Vacuolar protein that presumably functions within the endosomal-vacuolar trafficking pathway, affecting events that determine whether plasma membrane proteins are degraded or routed to the plasma membrane | Triacylglycerol lipase involved in triacylglycerol mobilization and degradation; found in lipid particles; potential Cdc28p substrate | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLL029W is not an essential gene but mutant is defective in spore formation | Phospholipase B (lysophospholipase) involved in lipid metabolism, required for deacylation of phosphatidylcholine and phosphatidylethanolamine but not phosphatidylinositol | NAD-dependent arabinose dehydrogenase, involved in biosynthesis of erythroascorbic acid; similar to plant L-galactose dehydrogenase | Glycerol 3-phosphate/dihydroxyacetone phosphate dual substrate-specific sn-1 acyltransferase of the glycerolipid biosynthesis pathway, prefers 16-carbon fatty acids, similar to Gpt2p, gene is constitutively transcribed | Putative protein of unknown function; predicted to contain a transmembrane domain; YMR148W is not an essential gene | Cyclin-like component of the RNA polymerase II holoenzyme, involved in phosphorylation of the RNA polymerase II C-terminal domain; involved in glucose repression and telomere maintenance | Protein with similarity to monocarboxylate permeases, appears not to be involved in transport of monocarboxylates such as lactate, pyruvate or acetate across the plasma membrane | Putative sulfate permease; physically interacts with Hsp82p; green fluorescent protein (GFP)-fusion protein localizes to the ER; YPR003C is not an essential gene |
| 260 | 45 | 437 | -0.196 | 0.495 |  |  | 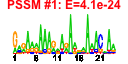 | 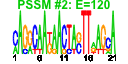 |  | YKR055W YBR114W YBR255W YBR273C YGL090W YGL098W YGL163C YGR099W YJL112W YKL017C YKL022C YKL208W YKR029C YKR051W YKR064W YLR170C YLR260W YLR266C YLR362W YLR369W YLR408C YLR456W YML112W YMR067C YMR073C YMR075W YMR077C YMR098C YMR284W YMR294W YNL127W YNL146W YNR010W YOL043C YOL134C YOR005C YOR076C YOR148C YOR256C YOR257W YOR258W YPL060W YPL070W YPL071C YPL138C | RHO4 RAD16 UBX7 LIF1 USE1 RAD54 TEL2 MDV1 HCS1 CDC16 CBT1 SET3 CDC14 APS1 LCB5 PDR8 STE11 SSQ1 CTK3 UBX4 IRC21 RCO1 VPS20 LRC4 YKU70 JNM1 FAR11 CSE2 NTG2 DNL4 SKI7 SPP2 TRE2 CDC31 HNT3 LPE10 MUK1 SPP1 | Non-essential small GTPase of the Rho/Rac subfamily of Ras-like proteins, likely to be involved in the establishment of cell polarity | Protein that recognizes and binds damaged DNA in an ATP-dependent manner (with Rad7p) during nucleotide excision repair; subunit of Nucleotide Excision Repair Factor 4 (NEF4) and the Elongin-Cullin-Socs (ECS) ligase complex | Protein of unknown function, required for normal growth rate at 15 degrees C; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | UBX (ubiquitin regulatory X) domain-containing protein that interacts with Cdc48p | Component of the DNA ligase IV complex that mediates nonhomologous end joining in DNA double-strand break repair; physically interacts with Dnl4p and Nej1p; homologous to mammalian XRCC4 protein | Essential SNARE protein localized to the ER, involved in retrograde traffic from the Golgi to the ER; forms a complex with the SNAREs Sec22p, Sec20p and Ufe1p | DNA-dependent ATPase, stimulates strand exchange by modifying the topology of double-stranded DNA; involved in the recombinational repair of double-strand breaks in DNA during vegetative growth and meiosis; member of the SWI/SNF family | Essential DNA-binding protein specific to single-stranded yeast telomeric DNA repeats, required for telomere length regulation and telomere position effect | Peripheral protein of the cytosolic face of the mitochondrial outer membrane, required for mitochondrial fission; interacts with Fis1p and with the dynamin-related GTPase Dnm1p; contains WD repeats | Hexameric DNA polymerase alpha-associated DNA helicase A involved in lagging strand DNA synthesis; contains single-stranded DNA stimulated ATPase and dATPase activities; replication protein A stimulates helicase and ATPase activities | Subunit of the anaphase-promoting complex/cyclosome (APC/C), which is a ubiquitin-protein ligase required for degradation of anaphase inhibitors, including mitotic cyclins, during the metaphase/anaphase transition; required for sporulation | Protein involved in 5' end processing of mitochondrial COB, 15S_rRNA, and RPM1 transcripts; may also have a role in 3' end processing of the COB pre-mRNA; displays genetic interaction with cell cycle-regulated kinase Dbf2p | Defining member of the SET3 histone deacetylase complex which is a meiosis-specific repressor of sporulation genes; necessary for efficient transcription by RNAPII; one of two yeast proteins that contains both SET and PHD domains | Putative protein of unknown function | Protein phosphatase required for mitotic exit; located in the nucleolus until liberated by the FEAR and Mitotic Exit Network in anaphase, enabling it to act on key substrates to effect a decrease in CDK/B-cyclin activity and mitotic exit | Small subunit of the clathrin-associated adaptor complex AP-1, which is involved in protein sorting at the trans-Golgi network; homolog of the sigma subunit of the mammalian clathrin AP-1 complex | Minor sphingoid long-chain base kinase, paralog of Lcb4p responsible for few percent of the total activity, possibly involved in synthesis of long-chain base phosphates, which function as signaling molecules | Transcription factor; targets include ATP-binding cassette (ABC) transporters, major facilitator superfamily transporters, and other genes involved in the pleiotropic drug resistance (PDR) phenomenon | Signal transducing MEK kinase involved in pheromone response and pseudohyphal/invasive growth pathways where it phosphorylates Ste7p, and the high osmolarity response pathway, via phosphorylation of Pbs2p; regulated by Ste20p and Ste50p | Mitochondrial hsp70-type molecular chaperone, required for assembly of iron/sulfur clusters into proteins at a step after cluster synthesis, and for maturation of Yfh1p, which is a homolog of human frataxin implicated in Friedreich's ataxia | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endosome; YLR408C is not an essential gene | Gamma subunit of C-terminal domain kinase I (CTDK-I), which phosphorylates the C-terminal repeated domain of the RNA polymerase II large subunit (Rpo21p) to affect both transcription and pre-mRNA 3' end processing | Putative protein of unknown function; proposed to be involved in resistance to carboplatin and cisplatin; shares similarity to a human cytochrome oxidoreductase; null mutant displays increased levels of spontaneous Rad52 foci | Essential subunit of the histone deacetylase Rpd3S complex; interacts with Eaf3p | Myristoylated subunit of ESCRTIII, the endosomal sorting complex required for transport of transmembrane proteins into the multivesicular body pathway to the lysosomal/vacuolar lumen; cytoplasmic protein recruited to endosomal membranes | Protein of unknown function; non-tagged protein is detected in highly purified mitochondria; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Subunit of the telomeric Ku complex (Yku70p-Yku80p), involved in telomere length maintenance, structure and telomere position effect; relocates to sites of double-strand cleavage to promote nonhomologous end joining during DSB repair | Component of the yeast dynactin complex, consisting of Nip100p, Jnm1p, and Arp1p; required for proper nuclear migration and spindle partitioning during mitotic anaphase B | Protein involved in G1 cell cycle arrest in response to pheromone, in a pathway different from the Far1p-dependent pathway; interacts with Far3p, Far7p, Far8p, Far9p, and Far10p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum; YNL146W is not an essential gene | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; component of the Med9/10 module; required for regulation of RNA polymerase II activity | DNA N-glycosylase and apurinic/apyrimidinic (AP) lyase involved in base excision repair, localizes to the nucleus | DNA ligase required for nonhomologous end-joining (NHEJ), forms stable heterodimer with required cofactor Lif1p, interacts with Nej1p; involved in meiosis, not essential for vegetative growth | Putative GTPase that mediates interactions via its N-terminus between the exosome and Ski complex (Ski2p, Ski3p, Ski8p) which operate in the 3'-to-5' mRNA-decay pathway; cytoplasmic protein required for degrading nonstop mRNAs | Essential protein that promotes the first step of splicing and is required for the final stages of spliceosome maturation; interacts with Prp2p, which may release Spp2p from the spliceosome following the first cleavage reaction | Protein that functions with Tre1p to regulate ubiquitylation and vacuolar degradation of the metal transporter Smf1p; has similarity to transferrin receptors; inviability of null mutant in systematic studies is due to proximity to CDC31 | Calcium-binding component of the spindle pole body (SPB) half-bridge, required for SPB duplication in mitosis and meiosis II; homolog of mammalian centrin; interacts with Kar1p | Member of the third branch of the histidine triad (HIT) superfamily of nucleotide-binding proteins; similar to Aprataxin, a Hint related protein that is mutated in individuals with ataxia with oculomotor apraxia | Mitochondrial inner membrane magnesium transporter, involved in maintenance of magnesium concentrations inside mitochondria; indirectly affects splicing of group II introns; functionally and structurally related to Mrs2p | Cytoplasmic protein of unknown function; computational analysis of large-scale protein-protein interaction data suggests a possible role in transcriptional regulation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Subunit of COMPASS (Set1C), a complex which methylates histone H3 on lysine 4 and is required in telomeric transcriptional silencing; PHD finger domain protein similar to human CGBP, an unmethylated CpG binding protein |
| 69 | 32 | 438 | -0.184 | 0.504 |  |  | 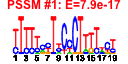 | 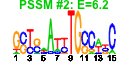 |  | YBR037C YBR111C YBR128C YBR151W YBR177C YCR004C YDL072C YDR517W YFR003C YGL047W YGL198W YGR167W YGR232W YIL034C YIL062C YJR044C YJR125C YLR250W YLR257W YML070W YMR210W YNL045W YOL032W YOR185C YPR079W YBR035C YDR368W YGR136W YKL013C YKL193C YKR014C YPR172W | SCO1 YSA1 ATG14 APD1 EHT1 YCP4 YET3 GRH1 YPI1 ALG13 YIP4 CLC1 NAS6 CAP2 ARC15 VPS55 ENT3 SSP120 DAK1 OPI10 GSP2 MRL1 PDX3 YPR1 LSB1 ARC19 SDS22 YPT52 | Copper-binding protein of the mitochondrial inner membrane, required for cytochrome c oxidase activity and respiration; may function to deliver copper to cytochrome c oxidase; has similarity to thioredoxins | Nudix hydrolase family member with ADP-ribose pyrophosphatase activity | Subunit of an autophagy-specific phosphatidylinositol 3-kinase complex (with Vps34p, Vps15p, and Vps30p) required for organization of a pre-autophagosomal structure; ATG14 transcription is activated by Gln3p during nitrogen starvation | Protein of unknown function, required for normal localization of actin patches and for normal tolerance of sodium ions and hydrogen peroxide; localizes to both cytoplasm and nucleus | Acyl-coenzymeA:ethanol O-acyltransferase that plays a minor role in medium-chain fatty acid ethyl ester biosynthesis; contains esterase activity; localizes to lipid particles and the mitochondrial outer membrane | Protein of unknown function, has sequence and structural similarity to flavodoxins; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein of unknown function; YET3 null mutant decreases the level of secreted invertase; homolog of human BAP31 protein | Acetylated, cis-golgi localized protein involved in ER to Golgi transport; homolog of human GRASP65; forms a complex with the coiled-coil protein Bug1p; mutants are compromised for the fusion of ER-derived vesicles with Golgi membranes | Inhibitor of the type I protein phosphatase Glc7p, which is involved in regulation of a variety of metabolic processes; overproduction causes decreased cellular content of glycogen | Catalytic component of UDP-GlcNAc transferase, required for the second step of dolichyl-linked oligosaccharide synthesis; anchored to the ER membrane via interaction with Alg14p; similar to bacterial and human glycosyltransferases | Protein that interacts with Rab GTPases, localized to late Golgi vesicles; computational analysis of large-scale protein-protein interaction data suggests a possible role in vesicle-mediated transport | Clathrin light chain, subunit of the major coat protein involved in intracellular protein transport and endocytosis; thought to regulate clathrin function, two Clathrin heavy chains (CHC1) form the clathrin triskelion structural component | Regulatory, non-ATPase subunit of the 26S proteasome; homolog of the human oncoprotein gankyrin, which interacts with the retinoblastoma tumor suppressor (Rb) and cyclin-dependent kinase 4/6 | Beta subunit of the capping protein (CP) heterodimer (Cap1p and Cap2p) which binds to the barbed ends of actin filaments preventing further polymerization; localized predominantly to cortical actin patches | Subunit of the ARP2/3 complex, which is required for the motility and integrity of cortical actin patches | Late endosomal protein involved in late endosome to vacuole trafficking; functional homolog of human obesity receptor gene-related protein (OB-RGRP) | Protein containing an N-terminal epsin-like domain involved in clathrin recruitment and traffic between the Golgi and endosomes; associates with the clathrin adaptor Gga2p | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Putative protein of unknown function | Dihydroxyacetone kinase, required for detoxification of dihydroxyacetone (DHA); involved in stress adaptation | Putative acyltransferase with similarity to Eeb1p and Eht1p, has a minor role in medium-chain fatty acid ethyl ester biosynthesis; may be involved in lipid metabolism and detoxification | Leucyl aminopeptidase (leukotriene A4 hydrolase) with epoxide hydrolase activity, metalloenzyme containing one zinc atom; role in vivo is not defined; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Protein with a possible role in phospholipid biosynthesis, based on inositol-excreting phenotype of the null mutant and its suppression by exogenous choline | GTP binding protein (mammalian Ranp homolog) involved in the maintenance of nuclear organization, RNA processing and transport; interacts with Kap121p, Kap123p and Pdr6p (karyophilin betas); Gsp1p homolog that is not required for viability | Membrane protein with similarity to mammalian mannose-6-phosphate receptors, possibly functions as a sorting receptor in the delivery of vacuolar hydrolases | Pyridoxine (pyridoxamine) phosphate oxidase, has homologs in E. coli and Myxococcus xanthus; transcription is under the general control of nitrogen metabolism | 2-methylbutyraldehyde reductase, may be involved in isoleucine catabolism | Protein containing an N-terminal SH3 domain; binds Las17p, which is a homolog of human Wiskott-Aldrich Syndrome protein involved in actin patch assembly and actin polymerization | Conserved nuclear regulatory subunit of Glc7p type 1 protein serine-threonine phosphatase (PP1), functions positively with Glc7p to promote dephosphorylation of nuclear substrates required for chromosome transmission during mitosis | GTPase, similar to Ypt51p and Ypt53p and to mammalian rab5; required for vacuolar protein sorting and endocytosis | Protein of unknown function, transcriptionally activated by Yrm1p along with genes involved in multidrug resistance |
| 288 | 31 | 447 | -0.183 | 0.528 |  |  | 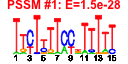 | 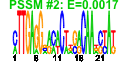 |  | YFR049W YJL045W YJR106W YMR103C YPR093C YPR192W YBR018C YBR020W YBR299W YDR125C YER015W YER054C YFL054C YGR292W YHR210C YIL120W YIL166C YJR108W YLR329W YML091C YMR056C YMR257C YNL019C YNL033W YNL117W YNL144C YOR178C YPL113C YPL119C YPR200C YPR201W | YMR31 ECM27 ASR1 AQY1 GAL7 GAL1 MAL32 ECM18 FAA2 GIP2 MAL12 QDR1 ABM1 REC102 RPM2 AAC1 PET111 MLS1 GAC1 YPL119C-A ARR2 ARR3 | Mitochondrial ribosomal protein of the small subunit, has similarity to human mitochondrial ribosomal protein MRP-S36 | Minor succinate dehydrogenase isozyme; homologous to Sdh1p, the major isozyme reponsible for the oxidation of succinate and transfer of electrons to ubiquinone; induced during the diauxic shift in a Cat8p-dependent manner | Non-essential protein of unknown function | Protein involved in a putative alcohol-responsive signaling pathway; accumulates in the nucleus under alcohol stress; contains a Ring/PHD finger domain | Spore-specific water channel that mediates the transport of water across cell membranes, developmentally controlled; may play a role in spore maturation, probably by allowing water outflow, may be involved in freeze tolerance | Galactose-1-phosphate uridyl transferase, synthesizes glucose-1-phosphate and UDP-galactose from UDP-D-glucose and alpha-D-galactose-1-phosphate in the second step of galactose catabolism | Galactokinase, phosphorylates alpha-D-galactose to alpha-D-galactose-1-phosphate in the first step of galactose catabolism; expression regulated by Gal4p | Maltase (alpha-D-glucosidase), inducible protein involved in maltose catabolism; encoded in the MAL3 complex locus; functional in genomic reference strain S288C | Protein of unknown function, similar to Rlp24p | Long chain fatty acyl-CoA synthetase; accepts a wider range of acyl chain lengths than Faa1p, preferring C9:0-C13:0; involved in the activation of endogenous pools of fatty acids | Putative regulatory subunit of the protein phosphatase Glc7p, involved in glycogen metabolism; contains a conserved motif (GVNK motif) that is also found in Gac1p, Pig1p, and Pig2p | Putative channel-like protein; similar to Fps1p; mediates passive diffusion of glycerol in the presence of ethanol | Maltase (alpha-D-glucosidase), inducible protein involved in maltose catabolism; encoded in the MAL1 complex locus | Putative protein of unknown function; non-essential gene; highly expressed under anaeorbic conditions; sequence similarity to aldose 1-epimerases such as GAL10 | Multidrug transporter of the major facilitator superfamily, required for resistance to quinidine, ketoconazole, fluconazole, and barban | Putative protein with similarity to the allantoate permease (Dal5p) subfamily of the major facilitator superfamily; mRNA expression is elevated by sulfur limitation; YIL166C is a non-essential gene | Protein of unknown function, required for normal microtubule organization | Protein involved in early stages of meiotic recombination; required for chromosome synapsis; forms a complex with Rec104p and Spo11p necessary during the initiation of recombination | Protein subunit of mitochondrial RNase P, has roles in nuclear transcription, cytoplasmic and mitochondrial RNA processing, and mitochondrial translation; distributed to mitochondria, cytoplasmic processing bodies, and the nucleus | Mitochondrial inner membrane ADP/ATP translocator, exchanges cytosolic ADP for mitochondrially synthesized ATP; phosphorylated; Aac1p is a minor isoform while Pet9p is the major ADP/ATP translocator | Mitochondrial translational activator specific for the COX2 mRNA; located in the mitochondrial inner membrane | Putative protein of unknown function | Malate synthase, enzyme of the glyoxylate cycle, involved in utilization of non-fermentable carbon sources; expression is subject to carbon catabolite repression; localizes in peroxisomes during growth in oleic acid medium | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YNL144C is not an essential gene | Regulatory subunit for Glc7p type-1 protein phosphatase (PP1), tethers Glc7p to Gsy2p glycogen synthase, binds Hsf1p heat shock transcription factor, required for induction of some HSF-regulated genes under heat shock | Glyoxylate reductase; acts on glyoxylate and hydroxypyruvate substrates; YPL113C is not an essential gene | Putative protein of unknown function; identified by expression profiling and mass spectrometry | Arsenate reductase required for arsenate resistance; converts arsenate to arsenite which can then be exported from cells by Arr3p | Arsenite transporter of the plasma membrane, required for resistance to arsenic compounds; transcription is activated by Arr1p in the presence of arsenite |
| 13 | 27 | 437 | -0.156 | 0.481 | 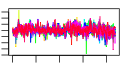 |  | 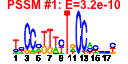 |  |  | YBR062C YBR204C YCL035C YDL027C YDR058C YDR059C YDR230W YDR368W YEL012W YGL248W YGR127W YGR141W YJL057C YJL068C YLR271W YLR283W YML004C YMR311C YNL011C YNL100W YOL048C YOL071W YOR121C YOR245C YER182W YOR173W YOR285W | GRX1 TGL2 UBC5 YPR1 UBC8 PDE1 VPS62 IKS1 GLO1 GLC8 AIM37 EMI5 DGA1 FMP10 DCS2 | hypothetical protein | Serine hydrolase; YBR204C is not an essential gene | Hydroperoxide and superoxide-radical responsive heat-stable glutathione-dependent disulfide oxidoreductase with active site cysteine pair; protects cells from oxidative damage | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YDL027C is not an essential gene | Protein with lipolytic activity towards triacylglycerols and diacylglycerols when expressed in E. coli; role in yeast lipid degradation is unclear | Ubiquitin-conjugating enzyme that mediates selective degradation of short-lived and abnormal proteins, central component of the cellular stress response; expression is heat inducible | 2-methylbutyraldehyde reductase, may be involved in isoleucine catabolism | Ubiquitin-conjugating enzyme that negatively regulates gluconeogenesis by mediating the glucose-induced ubiquitination of fructose-1,6-bisphosphatase (FBPase); cytoplasmic enzyme that catalyzes the ubiquitination of histones in vitro | Low-affinity cyclic AMP phosphodiesterase, controls glucose and intracellular acidification-induced cAMP signaling, target of the cAMP-protein kinase A (PKA) pathway; glucose induces transcription and inhibits translation | Putative protein of unknown function; expression is regulated by Msn2p/Msn4p, indicating a possible role in stress response | Vacuolar protein sorting (VPS) protein required for cytoplasm to vacuole targeting of proteins | Putative serine/threonine kinase; expression is induced during mild heat stress; deletion mutants are hypersensitive to copper sulphate and resistant to sorbate; interacts with an N-terminal fragment of Sst2p | Non-essential intracellular esterase that can function as an S-formylglutathione hydrolase; may be involved in the detoxification of formaldehyde, which can be metabolized to S-formylglutathione; similar to human esterase D | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and the nucleus and is induced in response to the DNA-damaging agent MMS | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YLR283W is not an essential gene | Monomeric glyoxalase I, catalyzes the detoxification of methylglyoxal (a by-product of glycolysis) via condensation with glutathione to produce S-D-lactoylglutathione; expression regulated by methylglyoxal levels and osmotic stress | Regulatory subunit of protein phosphatase 1 (Glc7p), involved in glycogen metabolism and chromosome segregation; proposed to regulate Glc7p activity via conformational alteration; ortholog of the mammalian protein phosphatase inhibitor 2 | Putative protein of unknown function; YNL011C is not an essential gene | Putative protein of unknown function; non-tagged protein is detected in purified mitochondria; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Putative protein of unknown function | Non-essential protein of unknown function required for transcriptional induction of the early meiotic-specific transcription factor IME1, also required for sporulation | Diacylglycerol acyltransferase, catalyzes the terminal step of triacylglycerol (TAG) formation, acylates diacylglycerol using acyl-CoA as an acyl donor, localized to lipid particles | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Non-essential, stress induced regulatory protein containing a HIT (histidine triad) motif; modulates m7G-oligoribonucleotide metabolism; inhibits Dcs1p; regulated by Msn2p, Msn4p, and the Ras-cAMP-cAPK signaling pathway, similar to Dcs1p. | Protein of unknown function, localized to the mitochondrial outer membrane |
| 155 | 38 | 437 | -0.082 | 0.551 | 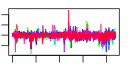 |  | 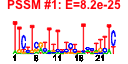 | 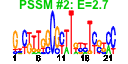 |  | YBR164C YBR254C YCR090C YDL070W YDR297W YDR331W YDR339C YDR381W YER146W YGL221C YIL020C YIL021W YIL040W YJL133W YJL167W YJL192C YJR024C YJR068W YJR133W YLR229C YNL217W YOL128C YOR246C YOR283W YPL227C YDR071C YDR367W YGR033C YHL014C YHR026W YJL193W YKL008C YLR147C YLR285W YLR447C YMR292W YOL149W YOR159C | ARL1 TRS20 BDF2 SUR2 GPI8 FCF1 YRA1 LSM5 NIF3 HIS6 RPB3 APQ12 MRS3 ERG20 SOP4 MDE1 RFC2 XPT1 CDC42 YGK3 ALG5 PAA1 TIM21 YLF2 IPP1 MTH1 SMD3 NNT1 VMA6 GOT1 DCP1 IME2 | Soluble GTPase with a role in regulation of membrane traffic; regulates potassium influx; G protein of the Ras superfamily, similar to ADP-ribosylation factor | One of 10 subunits of the transport protein particle (TRAPP) complex of the cis-Golgi which mediates vesicle docking and fusion; mutations in the human homolog cause the spondyloepiphyseal dysplasia tarda (SEDL) disorder | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YCR090C is not an essential gene | Protein involved in transcription initiation at TATA-containing promoters; associates with the basal transcription factor TFIID; contains two bromodomains; corresponds to the C-terminal region of mammalian TAF1; redundant with Bdf1p | Sphinganine C4-hydroxylase, catalyses the conversion of sphinganine to phytosphingosine in sphingolipid biosyntheis | ER membrane glycoprotein subunit of the glycosylphosphatidylinositol transamidase complex that adds glycosylphosphatidylinositol (GPI) anchors to newly synthesized proteins; human PIG-K protein is a functional homolog | Essential nucleolar protein that is a component of the SSU (small subunit) processome involved in the pre-rRNA processing steps of 40S ribosomal subunit biogenesis; contains a PINc domain; copurifies with Faf1p | Nuclear protein that binds to RNA and to Mex67p, required for export of poly(A)+ mRNA from the nucleus; member of the REF (RNA and export factor binding proteins) family; another family member, Yra2p, can substitute for Yra1p function | Lsm (Like Sm) protein; part of heteroheptameric complexes (Lsm2p-7p and either Lsm1p or 8p): cytoplasmic Lsm1p complex involved in mRNA decay; nuclear Lsm8p complex part of U6 snRNP and possibly involved in processing tRNA, snoRNA, and rRNA | Protein of unknown function, similar to Listeria monocytogenes major sigma factor (rpoD gene product); the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Phosphoribosyl-5-amino-1-phosphoribosyl-4- imidazolecarboxiamide isomerase, catalyzes the fourth step in histidine biosynthesis; mutations cause histidine auxotrophy and sensitivity to Cu, Co, and Ni salts | RNA polymerase II third largest subunit B44, part of central core; similar to prokaryotic alpha subunit | Protein involved in nucleocytoplasmic transport of mRNA | Mitochondrial iron transporter of the mitochondrial carrier family (MCF), very similar to and functionally redundant with Mrs4p; functions under low-iron conditions; may transport other cations in addition to iron | Farnesyl pyrophosphate synthetase, has both dimethylallyltranstransferase and geranyltranstransferase activities; catalyzes the formation of C15 farnesyl pyrophosphate units for isoprenoid and sterol biosynthesis | ER-membrane protein; suppressor of pma1-7, deletion of SOP4 slows down the export of wild-type Pma1p and Pma1-7 from the ER | Putative methylthio-ribulose-1-phosphate dehydratase; acts in the methionine salvage pathway; potential Smt3p sumoylation substrate; expression downregulated by caspofungin and deletion mutant is caspofungin resistant | Subunit of heteropentameric Replication factor C (RF-C), which is a DNA binding protein and ATPase that acts as a clamp loader of the proliferating cell nuclear antigen (PCNA) processivity factor for DNA polymerases delta and epsilon | Xanthine-guanine phosphoribosyl transferase, required for xanthine utilization and for optimal utilization of guanine | Small rho-like GTPase, essential for establishment and maintenance of cell polarity; mutants have defects in the organization of actin and septins | hypothetical protein | Protein kinase related to mammalian glycogen synthase kinases of the GSK-3 family; GSK-3 homologs (Mck1p, Rim11p, Mrk1p, Ygk3p) are involved in control of Msn2p-dependent transcription of stress responsive genes and in protein degradation | Protein with similarity to oxidoreductases, found in lipid particles; required for replication of Brome mosaic virus in S. cerevisiae, which is a model system for studying replication of positive-strand RNA viruses in their natural hosts | Putative protein of unknown function with some similarity to GPM1/YKL152C, a phosphoglycerate mutase; YOR283W is not an essential gene | UDP-glucose:dolichyl-phosphate glucosyltransferase, involved in asparagine-linked glycosylation in the endoplasmic reticulum | Polyamine acetyltransferase; acetylates polyamines (e.g. putrescine, spermidine, spermine) and also aralkylamines (e.g. tryptamine, phenylethylamine); may be involved in transcription and/or DNA replication | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Constituent of the mitochondrial inner membrane presequence translocase (TIM23 complex); may regulate protein import by binding to both the translocase of the outer membrane (TOM) and presequence-associated motor (PAM) complexes | Protein of unknown function, has weak similarity to E. coli GTP-binding protein gtp1; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Cytoplasmic inorganic pyrophosphatase (PPase), catalyzes the rapid exchange of oxygens from Pi with water, highly expressed and essential for viability, active-site residues show identity to those from E. coli PPase | Putative protein of unknown function, predicted to encode a triose phosphate transporter subfamily member based on phylogenetic analysis; similar to YOR307C/SLY41; deletion mutant has a respiratory growth defect | Negative regulator of the glucose-sensing signal transduction pathway, required for repression of transcription by Rgt1p; interacts with Rgt1p and the Snf3p and Rgt2p glucose sensors; phosphorylated by Yck1p, triggering Mth1p degradation | Core Sm protein Sm D3; part of heteroheptameric complex (with Smb1p, Smd1p, Smd2p, Sme1p, Smx3p, and Smx2p) that is part of the spliceosomal U1, U2, U4, and U5 snRNPs; homolog of human Sm D3 | Putative nicotinamide N-methyltransferase, has a role in rDNA silencing and in lifespan determination | Subunit d of the five-subunit V0 integral membrane domain of vacuolar H+-ATPase (V-ATPase), an electrogenic proton pump found in the endomembrane system; stabilizes VO subunits; required for V1 domain assembly on the vacuolar membrane | Evolutionarily conserved non-essential protein present in early Golgi cisternae that may be involved in ER-Golgi transport at a step after vesicle tethering to Golgi membranes, exhibits membrane topology similar to that of Sft2p | Subunit of the Dcp1p-Dcp2p decapping enzyme complex, which removes the 5' cap structure from mRNAs prior to their degradation; enhances the activity of catalytic subunit Dcp2p; regulated by DEAD box protein Dhh1p | Serine/threonine protein kinase involved in activation of meiosis, associates with Ime1p and mediates its stability, activates Ndt80p; IME2 expression is positively regulated by Ime1p |
| 204 | 50 | 438 | -0.077 | 0.507 |  |  | 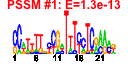 | 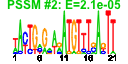 |  | YBR102C YGL232W YGR186W YGR196C YJL061W YKL072W YLR067C YLR095C YLR305C YLR306W YLR363C YLR418C YLR426W YML059C YMR073C YMR287C YMR288W YNL068C YNL272C YOR129C YPL009C YPL199C YPL202C YBL010C YER029C YER030W YJL004C YJL201W YJL203W YJL204C YJR060W YKL049C YKL089W YLR010C YLR115W YLR182W YLR187W YLR226W YML062C YMR172W YNL152W YNL260C YNL267W YNL316C YOR279C YOR308C YPL001W YPL011C YPL051W YPL103C | EXO84 TAN1 TFG1 FYV8 NUP82 STB6 PET309 IOC2 STT4 UBC12 NMD4 CDC73 NTE1 IRC21 SEM1 HSH155 FKH2 SEC2 AFT2 SMB1 CHZ1 SYS1 ECM25 PRP21 RCY1 CBF1 CSE4 MAS2 TEN1 CFT2 SWI6 SKG3 BUR2 MFT1 HOT1 PIK1 PHA2 RFM1 SNU66 HAT1 TAF3 ARL3 FMP30 | Essential protein with dual roles in spliceosome assembly and exocytosis; the exocyst complex (Sec3p, Sec5p, Sec6p, Sec8p, Sec10p, Sec15p, Exo70p, and Exo84p) mediates polarized targeting of secretory vesicles to active sites of exocytosis | Putative tRNA acetyltransferase, RNA-binding protein required for the formation of the modified nucleoside N(4)-acetylcytidine in serine and leucine tRNAs but not required for the same modification in 18S rRNA | TFIIF (Transcription Factor II) largest subunit; involved in both transcription initiation and elongation of RNA polymerase II; homologous to human RAP74 | Protein of unknown function, required for survival upon exposure to K1 killer toxin | Nucleoporin, subunit of the nuclear pore complex (NPC); forms a subcomplex with Nup159p and Nsp1p, interacts with Nup116p and is required for proper localization of Nup116p in the NPC | Protein that binds Sin3p in a two-hybrid assay | Specific translational activator for the COX1 mRNA, also influences stability of intron-containing COX1 primary transcripts; located in the mitochondrial inner membrane | Member of a complex (Isw1b) with Isw1p and Ioc4p that exhibits nucleosome-stimulated ATPase activity and acts within coding regions to coordinate transcription elongation with termination and processing, contains a PHD finger motif | Phosphatidylinositol-4-kinase that functions in the Pkc1p protein kinase pathway; required for normal vacuole morphology, cell wall integrity, and actin cytoskeleton organization | Enzyme that mediates the conjugation of Rub1p, a ubiquitin-like protein, to other proteins; related to E2 ubiquitin-conjugating enzymes | Protein interacting with Nam7p, may be involved in the nonsense-mediated mRNA decay pathway | Constituent of Paf1 complex with RNA polymerase II, Paf1p, Hpr1p, Ctr9, Leo1, Rtf1 and Ccr4p, distinct from Srb-containing Pol II complexes; required for expression of certain genes, modification of some histones, and telomere maintenance | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; proposed to be involved in resistance to mechlorethamine and streptozotocin | Serine esterase that deacylates exogenous lysophospholipids, homolog of human neuropathy target esterase (NTE); mammalian NTE1 deacylates phosphatidylcholine to glycerophosphocholine | Putative protein of unknown function; proposed to be involved in resistance to carboplatin and cisplatin; shares similarity to a human cytochrome oxidoreductase; null mutant displays increased levels of spontaneous Rad52 foci | Component of the lid subcomplex of the regulatory subunit of the 26S proteasome; ortholog of human DSS1 | U2-snRNP associated splicing factor that forms extensive associations with the branch site-3' splice site-3' exon region upon prespliceosome formation; similarity to the mammalian U2 snRNP-associated splicing factor SAP155 | Forkhead family transcription factor with a major role in the expression of G2/M phase genes; positively regulates transcriptional elongation; negative role in chromatin silencing at HML and HMR; substrate of the Cdc28p/Clb5p kinase | Guanyl-nucleotide exchange factor for the small G-protein Sec4p, located on cytoplasmic vesicles; essential for post-Golgi vesicle transport | Putative component of the outer plaque of the spindle pole body; may be involved in cation homeostasis or multidrug resistance | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YPL009C is not an essential gene | hypothetical protein | Iron-regulated transcriptional activator; activates genes involved in intracellular iron use and required for iron homeostasis and resistance to oxidative stress; similar to Aft1p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein colocalizes with clathrin-coated vesicles | Core Sm protein Sm B; part of heteroheptameric complex (with Smd1p, Smd2p, Smd3p, Sme1p, Smx3p, and Smx2p) that is part of the spliceosomal U1, U2, U4, and U5 snRNPs; homolog of human Sm B and Sm B' | Histone chaperone for Htz1p/H2A-H2B dimer; required for the stabilization of the Chz1p-Htz1-H2B complex; has overlapping function with Nap1p; null mutant displays weak sensitivity to MMS and benomyl; contains a highly conserved CHZ motif | Integral membrane protein of the Golgi required for targeting of the Arf-like GTPase Arl3p to the Golgi; multicopy suppressor of ypt6 null mutation | Non-essential protein of unknown function; promoter contains a consensus binding sequence for factor Abf1p | Subunit of the SF3a splicing factor complex, required for spliceosome assembly | F-box protein involved in recycling plasma membrane proteins internalized by endocytosis; localized to sites of polarized growth | Helix-loop-helix protein that binds the motif CACRTG, which is present at several sites including MET gene promoters and centromere DNA element I (CDEI); required for nucleosome positioning at this motif; targets Isw1p to DNA | Centromere protein that resembles histones, required for proper kinetochore function; homolog of human CENP-A | Larger subunit of the mitochondrial processing protease (MPP), essential processing enzyme that cleaves the N-terminal targeting sequences from mitochondrially imported proteins | Protein that regulates telomeric length; protects telomeric ends in a complex with Cdc13p and Stn1p | Subunit of the mRNA cleavage and polyadenlylation factor (CPF); required for pre-mRNA cleavage, polyadenylation and poly(A) site recognition, 43% similarity with the mammalian CPSF-100 protein. | Transcription cofactor, forms complexes with DNA-binding proteins Swi4p and Mbp1p to regulate transcription at the G1/S transition; involved in meiotic gene expression; localization regulated by phosphorylation; potential Cdc28p substrate | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery, cytoplasm, bud, and bud neck; potential Cdc28p substrate; similar to Caf120p and Skg4p | Cyclin for the Sgv1p (Bur1p) protein kinase; Sgv1p and Bur2p comprise a CDK-cyclin complex involved in transcriptional regulation through its phosphorylation of the carboxy-terminal domain of the largest subunit of RNA polymerase II | Subunit of the THO complex, which is a nuclear complex comprised of Hpr1p, Mft1p, Rlr1p, and Thp2p, that is involved in transcription elongation and mitotic recombination; involved in telomere maintenance | Transcription factor required for the transient induction of glycerol biosynthetic genes GPD1 and GPP2 in response to high osmolarity; targets Hog1p to osmostress responsive promoters; has similarity to Msn1p and Gcr1p | Putative protein of unknown function; cytoplasmically localized protein proposed to function in phospholipid binding; YBL107C is an essential gene | Putative protein of unknown function with similarity to a human protein overexpressed in oral cancers; localizes to the nucleus and cytoplasm; YNL260C is an essential gene | Phosphatidylinositol 4-kinase; catalyzes first step in the biosynthesis of phosphatidylinositol-4,5-biphosphate; may control cytokineses through the actin cytoskeleton | Prephenate dehydratase, catalyzes the conversion of prephanate to phenylpyruvate, which is a step in the phenylalanine biosynthesis pathway | DNA-binding protein required for vegetative repression of middle sporulation genes; specificity factor that directs the Hst1p histone deacetylase to some of the promoters regulated by Sum1p; involved in telomere maintenance | Component of the U4/U6.U5 snRNP complex involved in pre-mRNA splicing via spliceosome; has homology to human SART-1 and to an S. pombe protein; snu66 null mutation confers cold-sensitivity but is not lethal at normal growth temperatures | Catalytic subunit of the Hat1p-Hat2p histone acetyltransferase complex that uses the cofactor acetyl coenzyme A, to acetylate free nuclear and cytoplasmic histone H4; involved in telomeric silencing and DNA double-strand break repair | TFIID subunit (47 kDa), involved in promoter binding and RNA polymerase II transcription initiation | GTPase of the Ras superfamily required to recruit Arl1p to the Golgi; similar to ADP-ribosylation factor | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
| 87 | 44 | 437 | -0.075 | 0.540 |  |  |  |  |  | YDL054C YDR105C YDR244W YDR259C YDR435C YER041W YGL073W YGR016W YGR080W YHL006C YHL018W YHL019C YHR034C YIL002C YJL099W YJL149W YJR036C YJR052W YKL022C YKL219W YKL222C YKR064W YKR085C YLL006W YLR001C YLR113W YLR289W YMR025W YMR053C YMR201C YOL133W YPL257W YBL074C YGL035C YGR143W YGR212W YGR223C YJR131W YKR015C YKR086W YLR271W YLR403W YMR279C YOL060C | MCH1 TMS1 PEX5 YAP6 PPM1 YEN1 HSF1 TWF1 SHU1 APM2 PIH1 INP51 CHS6 DAS1 HUL4 RAD7 CDC16 COS9 CDC14 MRPL20 YLL006W-A HOG1 GUF1 CSI1 STB2 RAD14 TEC1 YPL257W-B AAR2 MIG1 SKN1 SLI1 HSV2 MNS1 PRP16 SFP1 MAM3 | Protein with similarity to mammalian monocarboxylate permeases, which are involved in transport of monocarboxylic acids across the plasma membrane; mutant is not deficient in monocarboxylate transport | Vacuolar membrane protein of unknown function that is conserved in mammals; predicted to contain eleven transmembrane helices; interacts with Pdr5p, a protein involved in multidrug resistance | Peroxisomal membrane signal receptor for the C-terminal tripeptide signal sequence (PTS1) of peroxisomal matrix proteins, required for peroxisomal matrix protein import; also proposed to have PTS1-receptor independent functions | Putative basic leucine zipper (bZIP) transcription factor; overexpression increases sodium and lithium tolerance | Carboxyl methyl transferase, methylates the C terminus of the protein phosphatase 2A catalytic subunit (Pph21p or Pph22p), which is important for complex formation with regulatory subunits | Protein of unknown function, has similarity to endonuclease Rth1p; potentially phosphorylated by Cdc28p | Trimeric heat shock transcription factor, activates multiple genes in response to stresses that include hyperthermia; recognizes variable heat shock elements (HSEs) consisting of inverted NGAAN repeats; posttranslationally regulated | Putative protein of unknown function | Twinfilin, highly conserved actin monomer-sequestering protein involved in regulation of the cortical actin cytoskeleton, composed of two cofilin-like regions, localizes actin monomers to sites of rapid filament assembly | Protein of unassigned function involved in mutation suppression, important for error-free repair of spontaneous and induced DNA lesions to protect the genome from mutation; associates with Shu2p, Psy3p, and Csm2p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria and is induced in response to the DNA-damaging agent MMS | Protein of unknown function, homologous to the medium chain of mammalian clathrin-associated protein complex; involved in vesicular transport | Protein of unresolved function; may function in protein folding and/or rRNA processing, interacts with a chaperone (Hsp82p), two chromatin remodeling factors (Rvb1p, Rvb2p) and two rRNA processing factors (Rrp43p, Nop58p) | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase, synaptojanin-like protein with an N-terminal Sac1 domain, plays a role in phosphatidylinositol 4,5-bisphosphate homeostasis and in endocytosis; null mutation confers cold-tolerant growth | Member of the ChAPs family of proteins (Chs5p-Arf1p-binding proteins: Bch1p, Bch2p, Bud7p, Chs6p), that forms the exomer complex with Chs5p to mediate export of specific cargo proteins, including Chs3p, from the Golgi to the plasma membrane | Putative SCF ubiquitin ligase F-box protein; interacts physically with both Cdc53p and Skp1 and genetically with CDC34; similar to putative F-box protein YDR131C; null mutant suppresses dst1delta sensitivity for 6-azauracil | Protein with similarity to hect domain E3 ubiquitin-protein ligases, not essential for viability | Protein that recognizes and binds damaged DNA in an ATP-dependent manner (with Rad16p) during nucleotide excision repair; subunit of Nucleotide Excision Repair Factor 4 (NEF4) and the Elongin-Cullin-Socs (ECS) ligase complex | Subunit of the anaphase-promoting complex/cyclosome (APC/C), which is a ubiquitin-protein ligase required for degradation of anaphase inhibitors, including mitotic cyclins, during the metaphase/anaphase transition; required for sporulation | Protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; similar to transcriptional regulators from the zinc cluster (binuclear cluster) family; null mutant is sensitive to caffeine | Protein phosphatase required for mitotic exit; located in the nucleolus until liberated by the FEAR and Mitotic Exit Network in anaphase, enabling it to act on key substrates to effect a decrease in CDK/B-cyclin activity and mitotic exit | Mitochondrial ribosomal protein of the large subunit | Putative protein of unknown function; identified by fungal homology and RT-PCR | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YLR001C is not an essential gene | Mitogen-activated protein kinase involved in osmoregulation via three independent osmosensors; mediates the recruitment and activation of RNA Pol II at Hot1p-dependent promoters; localization regulated by Ptp2p and Ptp3p | Mitochondrial GTPase of unknown function, similar to E. coli elongation factor-type GTP-binding protein LepA and to LK1236.1 from Caenorhabditis elegans | Subunit of the Cop9 signalosome, which is required for deneddylation, or removal of the ubiquitin-like protein Rub1p from Cdc53p (cullin); involved in adaptation to pheromone signaling | Protein that interacts with Sin3p in a two-hybrid assay and is part of a large protein complex with Sin3p and Stb1p | Protein that recognizes and binds damaged DNA during nucleotide excision repair; subunit of Nucleotide Excision Repair Factor 1 (NEF1); contains zinc finger motif; homolog of human XPA protein | Transcription factor required for full Ty1 epxression, Ty1-mediated gene activation, and haploid invasive and diploid pseudohyphal growth; TEA/ATTS DNA-binding domain family member | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Component of the U5 snRNP, required for splicing of U3 precursors; originally described as a splicing factor specifically required for splicing pre-mRNA of the MATa1 cistron | Transcription factor involved in glucose repression; sequence specific DNA binding protein containing two Cys2His2 zinc finger motifs; regulated by the SNF1 kinase and the GLC7 phosphatase | Protein involved in sphingolipid biosynthesis; type II membrane protein with similarity to Kre6p | N-acetyltransferase, confers resistance to the sphingolipid biosynthesis inhibitor myriocin (ISP-1) by converting it into N-acetyl-myriocin, co-operates with Ypk1p in mediating resistance to myriocin | Phosphatidylinositol 3,5-bisphosphate-binding protein, predicted to fold as a seven-bladed beta-propeller; displays punctate cytoplasmic localization | Alpha-1,2-mannosidase involved in ER quality control; catalyzes the removal of one mannose residue from Man9GlcNAc to produce a single isomer of Man8GlcNAc in N-linked oligosaccharide biosynthesis; integral to ER membrane | RNA helicase in the DEAH-box family involved in the second catalytic step of splicing, exhibits ATP-dependent RNA unwinding activity | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and the nucleus and is induced in response to the DNA-damaging agent MMS | Transcription factor that controls expression of many ribosome biogenesis genes in response to nutrients and stress, regulates G2/M transitions during mitotic cell cycle and DNA-damage response, involved in cell size modulation | Putative protein of unknown function; identified as a heat-induced gene in a high-throughout screen; YMR279C is not an essential gene | Protein required for normal mitochondrial morphology, has similarity to hemolysins |
| 201 | 25 | 445 | -0.068 | 0.464 |  |  | 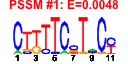 |  |  | YBR252W YCRX16C YDR152W YDR413C YDR429C YGR158C YHR127W YIL110W YNL244C YNL255C YOR021C YOR169C YOR210W YPL238C YBR164C YCR087W YDL050C YDR045C YDR321W YDR339C YGL070C YGR078C YIL008W YOR277C YPR187W | DUT1 GIR2 TIF35 MTR3 MNI1 SUI1 GIS2 RPB10 ARL1 RPC11 ASP1 FCF1 RPB9 PAC10 URM1 RPO26 | dUTPase, catalyzes hydrolysis of dUTP to dUMP and PPi, thereby preventing incorporation of uracil into DNA during replication; critical for the maintenance of genetic stability | Highly-acidic cytoplasmic RWD domain-containing protein of unknown function, sensitive to proteolysis, N-terminal region has high content of acidic amino acid residues, putative IUP (intrinsically unstructured protein) | Subunit of the core complex of translation initiation factor 3(eIF3), which is essential for translation | 3'5' exoribonuclease, exosome subunit; nucleolar protein involved in export of mRNA and ribosomal subunits; homologous to the E. coli exonuclease RNase PH | Protein of unknown function; localizes to the nucleus | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; deletion mutant exhibits a weak vacuolar protein sorting defect, enhanced resistance to caspofungin, and is synthetically lethal with MEN mutants | Translation initiation factor eIF1; component of a complex involved in recognition of the initiator codon; modulates translation accuracy at the initiation phase | Protein with seven cysteine-rich CCHC zinc-finger motifs, similar to human CNBP, proposed to be involved in the RAS/cAMP signaling pathway | Putative protein of unknown function; YOR021C is not an essential gene | RNA polymerase subunit ABC10-beta, common to RNA polymerases I, II, and III | Soluble GTPase with a role in regulation of membrane traffic; regulates potassium influx; G protein of the Ras superfamily, similar to ADP-ribosylation factor | RNA polymerase III subunit C11; mediates pol III RNA cleavage activity and is important for termination of transcription; homologous to TFIIS | Cytosolic L-asparaginase, involved in asparagine catabolism | Essential nucleolar protein that is a component of the SSU (small subunit) processome involved in the pre-rRNA processing steps of 40S ribosomal subunit biogenesis; contains a PINc domain; copurifies with Faf1p | RNA polymerase II subunit B12.6; contacts DNA; mutations affect transcription start site; involved in telomere maintenance | Part of the heteromeric co-chaperone GimC/prefoldin complex, which promotes efficient protein folding | Ubiquitin-like protein with weak sequence similarity to ubiquitin; depends on the E1-like activating enzyme Uba4p; molecular function of the Urm1p pathway is unknown, but it is required for normal growth, particularly at high temperature | RNA polymerase subunit ABC23, common to RNA polymerases I, II, and III; part of central core; similar to bacterial omega subunit |
| 216 | 41 | 431 | -0.058 | 0.564 |  |  | 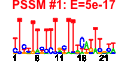 | 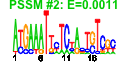 |  | YLR056W YLR183C YLR300W YOL030W YBR029C YBR078W YBR088C YCR065W YDR144C YDR309C YER003C YGR014W YGR054W YGR155W YHR042W YHR149C YIL040W YIL123W YIL140W YJL158C YKR013W YKR042W YKR093W YLR110C YLR342W YLR379W YML052W YML085C YML124C YMR199W YNL238W YNL283C YNL300W YOL007C YOL019W YOR209C YOR247W YPL032C YPL163C YPL256C YPR159W | ERG3 TOS4 EXG1 GAS5 CDS1 ECM33 POL30 HCM1 MKC7 GIC2 PMI40 MSB2 CYS4 CPR1 SKG6 APQ12 SIM1 AXL2 CIS3 PRY2 UTH1 PTR2 CCW12 FKS1 SUR7 TUB1 TUB3 CLN1 PSA1 WSC2 CSI2 YOL019W-A NPT1 SRL1 SVL3 SVS1 CLN2 KRE6 | C-5 sterol desaturase, catalyzes the introduction of a C-5(6) double bond into episterol, a precursor in ergosterol biosynthesis; mutants are viable, but cannot grow on non-fermentable carbon sources | Transcription factor that binds to a number of promoter regions, particularly promoters of some genes involved in pheromone response and cell cycle; potential Cdc28p substrate; expression is induced in G1 by bound SBF | Major exo-1,3-beta-glucanase of the cell wall, involved in cell wall beta-glucan assembly; exists as three differentially glycosylated isoenzymes | 1,3-beta-glucanosyltransferase, has similarity to Gas1p; localizes to the cell wall | Phosphatidate cytidylyltransferase (CDP-diglyceride synthetase); an enzyme that catalyzes that conversion of CTP + phosphate into diphosphate + CDP-diaclglyerol, a critical step in the synthesis of all major yeast phospholipids | GPI-anchored protein of unknown function, has a possible role in apical bud growth; GPI-anchoring on the plasma membrane crucial to function; phosphorylated in mitochondria; similar to Sps2p and Pst1p | Proliferating cell nuclear antigen (PCNA), functions as the sliding clamp for DNA polymerase delta; may function as a docking site for other proteins required for mitotic and meiotic chromosomal DNA replication and for DNA repair | Forkhead transcription factor that drives S-phase specific expression of genes involved in chromosome segregation, spindle dynamics, and budding; suppressor of calmodulin mutants with specific SPB assembly defects; telomere maintenance role | GPI-anchored aspartyl protease (yapsin) involved in protein processing; shares functions with Yap3p and Kex2p | Protein of unknown function involved in initiation of budding and cellular polarization, interacts with Cdc42p via the Cdc42/Rac-interactive binding (CRIB) domain | Mannose-6-phosphate isomerase, catalyzes the interconversion of fructose-6-P and mannose-6-P; required for early steps in protein mannosylation | Mucin family member involved in the Cdc42p- and MAP kinase-dependent filamentous growth signaling pathway; also functions as an osmosensor in parallel to the Sho1p-mediated pathway; potential Cdc28p substrate | Eukaryotic initiation factor (eIF) 2A; associates specifically with both 40S subunits and 80 S ribosomes, and interacts genetically with both eIF5b and eIF4E; homologous to mammalian eIF2A | Cystathionine beta-synthase, catalyzes the synthesis of cystathionine from serine and homocysteine, the first committed step in cysteine biosynthesis | Cytoplasmic peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; binds the drug cyclosporin A | Integral membrane protein that localizes primarily to growing sites such as the bud tip or the cell periphery; potential Cdc28p substrate; Skg6p interacts with Zds1p and Zds2p | Protein involved in nucleocytoplasmic transport of mRNA | Protein of the SUN family (Sim1p, Uth1p, Nca3p, Sun4p) that may participate in DNA replication, promoter contains SCB regulation box at -300 bp indicating that expression may be cell cycle-regulated | Integral plasma membrane protein required for axial budding in haploid cells, localizes to the incipient bud site and bud neck; glycosylated by Pmt4p; potential Cdc28p substrate | Mannose-containing glycoprotein constituent of the cell wall; member of the PIR (proteins with internal repeats) family | Protein of unknown function, has similarity to Pry1p and Pry3p and to the plant PR-1 class of pathogen related proteins | Mitochondrial outer membrane and cell wall localized SUN family member required for mitochondrial autophagy; involved in the oxidative stress response, life span during starvation, mitochondrial biogenesis, and cell death | Integral membrane peptide transporter, mediates transport of di- and tri-peptides; conserved protein that contains 12 transmembrane domains; PTR2 expression is regulated by the N-end rule pathway via repression by Cup9p | Cell wall mannoprotein, mutants are defective in mating and agglutination, expression is downregulated by alpha-factor | Catalytic subunit of 1,3-beta-D-glucan synthase, functionally redundant with alternate catalytic subunit Gsc2p; binds to regulatory subunit Rho1p; involved in cell wall synthesis and maintenance; localizes to sites of cell wall remodeling | Putative integral membrane protein; component of eisosomes; associated with endocytosis, along with Pil1p and Lsp1p; sporulation and plasma membrane sphingolipid content are altered in mutants | Alpha-tubulin; associates with beta-tubulin (Tub2p) to form tubulin dimer, which polymerizes to form microtubules | Alpha-tubulin; associates with beta-tubulin (Tub2p) to form tubulin dimer, which polymerizes to form microtubules; expressed at lower level than Tub1p | G1 cyclin involved in regulation of the cell cycle; activates Cdc28p kinase to promote the G1 to S phase transition; late G1 specific expression depends on transcription factor complexes, MBF (Swi6p-Mbp1p) and SBF (Swi6p-Swi4p) | GDP-mannose pyrophosphorylase (mannose-1-phosphate guanyltransferase), synthesizes GDP-mannose from GTP and mannose-1-phosphate in cell wall biosynthesis; required for normal cell wall structure | Partially redundant sensor-transducer of the stress-activated PKC1-MPK1 signaling pathway involved in maintenance of cell wall integrity and recovery from heat shock; secretory pathway Wsc2p is required for the arrest of secretion response | Glycosylphosphatidylinositol-dependent cell wall protein, expression is periodic and decreases in respone to ergosterol perturbation or upon entry into stationary phase; depletion increases resistance to lactic acid | Protein of unknown function; green fluorescent protein (GFP)- fusion protein localizes to the mother side of the bud neck and the vacuole; YOL007C is not an essential gene | Identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching | Nicotinate phosphoribosyltransferase, acts in the salvage pathway of NAD+ biosynthesis; required for silencing at rDNA and telomeres and has a role in silencing at mating-type loci; localized to the nucleus | Mannoprotein that exhibits a tight association with the cell wall, required for cell wall stability in the absence of GPI-anchored mannoproteins; has a high serine-threonine content; expression is induced in cell wall mutants | Protein of unknown function, mutant phenotype suggests a potential role in vacuolar function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery, cytoplasm, bud, and bud neck | Cell wall and vacuolar protein, required for wild-type resistance to vanadate | Protein required for beta-1,6 glucan biosynthesis; putative beta-glucan synthase; appears functionally redundant with Skn1p |
| 63 | 30 | 441 | -0.054 | 0.521 | 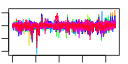 |  | 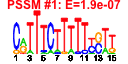 | 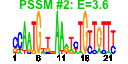 |  | YAR071W YBL068W YBR092C YBR093C YBR242W YDL050C YDL062W YDL111C YDR280W YDR454C YEL048C YER049W YER110C YGR177C YGR234W YHR215W YKL110C YMR292W YNL010W YOL061W YOL124C YPL019C YPL117C YPL207W YPL252C YPR016C YDR346C YFL004W YML106W YOR051C | PHO11 PRS4 PHO3 PHO5 RRP42 RRP45 GUK1 TPA1 KAP123 ATF2 YHB1 PHO12 KTI12 GOT1 PRS5 TRM11 VTC3 IDI1 TYW1 YAH1 TIF6 SVF1 VTC2 URA5 | One of three repressible acid phosphatases, a glycoprotein that is transported to the cell surface by the secretory pathway; induced by phosphate starvation and coordinately regulated by PHO4 and PHO2 | 5-phospho-ribosyl-1(alpha)-pyrophosphate synthetase, involved in nucleotide, histidine, and tryptophan biosynthesis; one of five related enzymes, which are active as heteromultimeric complexes | Constitutively expressed acid phosphatase similar to Pho5p; brought to the cell surface by transport vesicles; hydrolyzes thiamin phosphates in the periplasmic space, increasing cellular thiamin uptake; expression is repressed by thiamin | Repressible acid phosphatase (1 of 3) that also mediates extracellular nucleotide-derived phosphate hydrolysis; secretory pathway derived cell surface glycoprotein; induced by phosphate starvation and coordinately regulated by PHO4 and PHO2 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YBR242W is not an essential gene | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex with Rrp4p, Rrp41p, Rrp43p and Dis3p | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex | Guanylate kinase, converts GMP to GDP; required for growth and mannose outer chain elongation of cell wall N-linked glycoproteins | Putative protein of unknown function; synthetic lethal with gcs1delta | Protein of unknown function; interacts with Sup45p (eRF1), Sup35p (eRF3) and Pab1p; has a role in translation termination efficiency, mRNA poly(A) tail length and mRNA stability | Karyopherin beta, mediates nuclear import of ribosomal proteins prior to assembly into ribosomes and import of histones H3 and H4; localizes to the nuclear pore, nucleus, and cytoplasm; exhibits genetic interactions with RAI1 | Alcohol acetyltransferase, may play a role in steroid detoxification; forms volatile esters during fermentation, which is important in brewing | Nitric oxide oxidoreductase, flavohemoglobin involved in nitric oxide detoxification; plays a role in the oxidative and nitrosative stress responses | One of three repressible acid phosphatases, a glycoprotein that is transported to the cell surface by the secretory pathway; nearly identical to Pho11p; upregulated by phosphate starvation | Protein that plays a role, with Elongator complex, in modification of wobble nucleosides in tRNA; involved in sensitivity to G1 arrest induced by zymocin; interacts with chromatin throughout the genome; also interacts with Cdc19p | Evolutionarily conserved non-essential protein present in early Golgi cisternae that may be involved in ER-Golgi transport at a step after vesicle tethering to Golgi membranes, exhibits membrane topology similar to that of Sft2p | Putative protein of unknown function with similarity to phosphoserine phosphatases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; homozygous diploid mutant shows an increase in glycogen accumulation | Catalytic subunit of an adoMet-dependent tRNA methyltransferase complex (Trm11p-Trm112p), required for the methylation of the guanosine nucleotide at position 10 (m2G10) in tRNAs; contains a THUMP domain and a methyltransferase domain | Vacuolar membrane protein involved in vacuolar polyphosphate accumulation; functions as a regulator of vacuolar H+-ATPase activity and vacuolar transporter chaperones; involved in non-autophagic vacuolar fusion | Isopentenyl diphosphate:dimethylallyl diphosphate isomerase (IPP isomerase), catalyzes an essential activation step in the isoprenoid biosynthetic pathway; required for viability | Protein required for the synthesis of wybutosine, a modified guanosine found at the 3'-position adjacent to the anticodon of phenylalanine tRNA which supports reading frame maintenance by stabilizing codon-anticodon interactions | Ferredoxin of the mitochondrial matrix required for formation of cellular iron-sulfur proteins; involved in heme A biosynthesis; homologous to human adrenodoxin | Constituent of 66S pre-ribosomal particles, has similarity to human translation initiation factor 6 (eIF6); may be involved in the biogenesis and or stability of 60S ribosomal subunits | Protein with a potential role in cell survival pathways, required for the diauxic growth shift; expression in mammalian cells increases survival under conditions inducing apoptosis | Vacuolar membrane protein involved in vacuolar polyphosphate accumulation; functions as a regulator of vacuolar H+-ATPase activity and vacuolar transporter chaperones; involved in protein localization and non-autophagic vacuolar fusion | One of two orotate phosphoribosyltransferase isozymes (see also URA10) that catalyze the fifth enzymatic step in de novo biosynthesis of pyrimidines, converting orotate into orotidine-5'-phosphate | Nuclear protein that inhibits replication of Brome mosaic virus in S. cerevisiae, which is a model system for studying replication of positive-strand RNA viruses in their natural hosts |
| 30 | 51 | 432 | -0.053 | 0.538 | 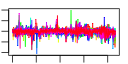 |  |  |  |  | YBL061C YBR007C YBR060C YCR052W YDR013W YDR049W YFL013C YFR009W YHR045W YJL085W YJL087C YJL201W YJR131W YKL073W YKL074C YLR085C YLR226W YLR277C YLR321C YML071C YML088W YML112W YMR091C YMR212C YMR270C YNL004W YNL059C YNL147W YNL148C YNL152W YNL153C YNL164C YNL236W YNL238W YNL264C YOL028C YOL115W YOR112W YOR123C YOR205C YOR281C YOR305W YPL083C YPL112C YPL195W YPL209C YPR018W YPR126C YPR130C YPR174C YPR175W | SKT5 DSF2 ORC2 RSC6 PSF1 IES1 GCN20 EXO70 TRL1 ECM25 MNS1 LHS1 MUD2 ARP6 BUR2 YSH1 YKL091C COG8 UFO1 CTK3 NPL6 EFR3 RRN9 HRB1 ARP5 LSM7 ALF1 GIM3 IBD2 SIN4 PSA1 PDR17 YAP7 PAP2 CEX1 LEO1 LRC5 PLP2 SEN54 PEX25 APL5 IPL1 RLF2 DPB2 | Activator of Chs3p (chitin synthase III), recruits Chs3p to the bud neck via interaction with Bni4p; has similarity to Shc1p, which activates Chs3p during sporulation | Deletion suppressor of mpt5 mutation | Subunit of the origin recognition complex, which directs DNA replication by binding to replication origins and is also involved in transcriptional silencing; phosphorylated by Cdc28p | Component of the RSC chromatin remodeling complex; essential for mitotic growth; homolog of SWI/SNF subunit Swp73p | Subunit of the GINS complex (Sld5p, Psf1p, Psf2p, Psf3p), which is localized to DNA replication origins and implicated in assembly of the DNA replication machinery | Zinc finger protein; putative transcription factor that may interact with proteins involved in histone acetylation or deacetylation; may be involved in altering acetylation on histone lysines | Subunit of the INO80 chromatin remodeling complex | Positive regulator of the Gcn2p kinase activity, forms a complex with Gcn1p; proposed to stimulate Gcn2p activation by an uncharged tRNA | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum | Essential 70kDa subunit of the exocyst complex (Sec3p, Sec5p, Sec6p, Sec8p, Sec10p, Sec15p, Exo70p, and Exo84p), which has the essential function of mediating polarized targeting of secretory vesicles to active sites of exocytosis | tRNA ligase, required for tRNA splicing; composed of three essential domains containing the phosphodiesterase, polynucleotide kinase, and ligase activities required for ligation; localized at the inner membrane of the nuclear envelope | Non-essential protein of unknown function; promoter contains a consensus binding sequence for factor Abf1p | Alpha-1,2-mannosidase involved in ER quality control; catalyzes the removal of one mannose residue from Man9GlcNAc to produce a single isomer of Man8GlcNAc in N-linked oligosaccharide biosynthesis; integral to ER membrane | Molecular chaperone of the endoplasmic reticulum lumen, involved in polypeptide translocation and folding; member of the Hsp70 family; localizes to the lumen of the ER; regulated by the unfolded protein response pathway | Protein involved in early pre-mRNA splicing; component of the pre-mRNA-U1 snRNP complex, the commitment complex; interacts with Msl5p/BBP splicing factor and Sub2p; similar to metazoan splicing factor U2AF65 | Actin-related protein that binds nucleosomes; a component of the SWR1 complex, which exchanges histone variant H2AZ (Htz1p) for chromatin-bound histone H2A | Cyclin for the Sgv1p (Bur1p) protein kinase; Sgv1p and Bur2p comprise a CDK-cyclin complex involved in transcriptional regulation through its phosphorylation of the carboxy-terminal domain of the largest subunit of RNA polymerase II | Putative endoribonuclease, subunit of the mRNA cleavage and polyadenylation specificity complex required for 3' processing of mRNAs | Putative homolog of Sec14p, which is a phosphatidylinositol/phosphatidylcholine transfer protein involved in lipid metabolism; localizes to the nucleus | Component of the conserved oligomeric Golgi complex (Cog1p through Cog8p), a cytosolic tethering complex that functions in protein trafficking to mediate fusion of transport vesicles to Golgi compartments | F-box receptor protein, subunit of the Skp1-Cdc53-F-box receptor (SCF) E3 ubiquitin ligase complex; binds to phosphorylated Ho endonuclease, allowing its ubiquitylation by SCF and subsequent degradation | Gamma subunit of C-terminal domain kinase I (CTDK-I), which phosphorylates the C-terminal repeated domain of the RNA polymerase II large subunit (Rpo21p) to affect both transcription and pre-mRNA 3' end processing | Component of the RSC chromatin remodeling complex; interacts with Rsc3p, Rsc30p, Ldb7p, and Htl1p to form a module important for a broad range of RSC functions; involved in nuclear protein import and maintenance of proper telomere length | Non-essential protein of unknown function; exhibits synthetic lethal genetic interactions with PHO85; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies and is phosphorylated | Protein involved in promoting high level transcription of rDNA, subunit of UAF (upstream activation factor) for RNA polymerase I | Poly(A+) RNA-binding protein, involved in the export of mRNAs from the nucleus to the cytoplasm; similar to Gbp2p and Npl3p | Nuclear actin-related protein involved in chromatin remodeling, component of chromatin-remodeling enzyme complexes | Lsm (Like Sm) protein; part of heteroheptameric complexes (Lsm2p-7p and either Lsm1p or 8p): cytoplasmic Lsm1p complex involved in mRNA decay; nuclear Lsm8p complex part of U6 snRNP and possibly involved in processing tRNA, snoRNA, and rRNA | Alpha-tubulin folding protein, similar to mammalian cofactor B; Alf1p-GFP localizes to cytoplasmic microtubules; required for the folding of alpha-tubulin and may play an additional role in microtubule maintenance | Putative protein of unknown function; cytoplasmically localized protein proposed to function in phospholipid binding; YBL107C is an essential gene | Subunit of the heterohexameric cochaperone prefoldin complex which binds specifically to cytosolic chaperonin and transfers target proteins to it | Component of the BUB2-dependent spindle checkpoint pathway, interacts with Bfa1p and functions upstream of Bub2p and Bfa1p | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; contributes to both postive and negative transcriptional regulation; dispensible for basal transcription | GDP-mannose pyrophosphorylase (mannose-1-phosphate guanyltransferase), synthesizes GDP-mannose from GTP and mannose-1-phosphate in cell wall biosynthesis; required for normal cell wall structure | Phosphatidylinositol transfer protein (PITP), downregulates Plb1p-mediated turnover of phosphatidylcholine, found in the cytosol and microsomes, homologous to Pdr16p, deletion affects phospholipid composition | Putative basic leucine zipper (bZIP) transcription factor | Catalytic subunit of TRAMP (Trf4/Pap2p-Mtr4p-Air1p/2p), a nuclear poly (A) polymerase complex involved in RNA quality control; catalyzes polyadenylation of unmodified tRNAs, and snoRNA and rRNA precursors; disputed role as a DNA polymerase | Cytoplasmic component of the nuclear aminoacylation-dependent tRNA export pathway; interacts with nuclear pore component Nup116p; copurifies with tRNA export receptors Los1p and Msn5p, as well as eIF-1a and the RAN GTPase Gsp1p | Component of the Paf1 complex, which associates with RNA polymerase II and is involved in histone methylation | Protein of unknown function; non-tagged protein is detected in purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss and severe growth defect in minimal glycerol media | Essential protein with similarity to phosducins, which are G-protein regulators; lethality of null mutation is functionally complemented by expression of mouse phosducin-like protein MgcPhLP | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the mitochondrion; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO); YOR305W is not an essential gene | Subunit of the tRNA splicing endonuclease, which is composed of Sen2p, Sen15p, Sen34p, and Sen54p | Peripheral peroxisomal membrane peroxin required for the regulation of peroxisome size and maintenance, recruits GTPase Rho1p to peroxisomes, induced by oleate, interacts with homologous protein Pex27p | Delta adaptin-like subunit of the clathrin associated protein complex (AP-3); functions in transport of alkaline phosphatase to the vacuole via the alternate pathway, suppressor of loss of casein kinase 1 function | Aurora kinase involved in regulating kinetochore-microtubule attachments; helps maintain condensed chromosomes during anaphase and early telophase; associates with Sli15p, which promotes Ipl1p association with mitotic spindle | Largest subunit (p90) of the Chromatin Assembly Complex (CAF-1) with Cac2p and Msi1p that assembles newly synthesized histones onto recently replicated DNA; involved in the maintenance of transcriptionally silent chromatin | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nuclear periphery; potential Cdc28p substrate | Second largest subunit of DNA polymerase II (DNA polymerase epsilon), required for normal yeast chromosomal replication; expression peaks at the G1/S phase boundary; potential Cdc28p substrate |
| 20 | 42 | 445 | -0.026 | 0.555 |  |  |  |  |  | YBR158W YDL084W YDL179W YDR033W YDR211W YDR346C YER031C YER032W YER124C YFL026W YGL008C YGL021W YGL028C YGL116W YGR108W YHR152W YHR201C YIL158W YJL051W YJL111W YJL183W YJR047C YKL164C YKL165C YKL185W YLR049C YLR153C YLR286C YLR447C YMR123W YNL058C YNL066W YNL145W YNL216W YNL327W YOR025W YOR264W YOR315W YPL256C YPR113W YGR229C YLR190W | AMN1 SUB2 PCL9 MRH1 GCD6 SVF1 YPT31 FIR1 DSE1 STE2 PMA1 ALK1 SCW11 CDC20 CLB1 SPO12 PPX1 AIM20 IRC8 CCT7 MNN11 ANB1 PIR1 MCD4 ASH1 ACS2 CTS1 VMA6 PKR1 SUN4 MFA2 RAP1 EGT2 HST3 DSE3 SFG1 CLN2 PIS1 SMI1 MMR1 | Protein required for daughter cell separation, multiple mitotic checkpoints, and chromosome stability; contains 12 degenerate leucine-rich repeat motifs; expression is induced by the Mitotic Exit Network (MEN) | Component of the TREX complex required for nuclear mRNA export; member of the DEAD-box RNA helicase superfamily and is involved in early and late steps of spliceosome assembly; homolog of the human splicing factor hUAP56 | Cyclin, forms a functional kinase complex with Pho85p cyclin-dependent kinase (Cdk), expressed in late M/early G1 phase, activated by Swi5p | Protein that localizes primarily to the plasma membrane, also found at the nuclear envelope; the authentic, non-tagged protein is detected in mitochondria in a phosphorylated state; has similarity to Hsp30p and Yro2p | Catalytic epsilon subunit of the translation initiation factor eIF2B, the guanine-nucleotide exchange factor for eIF2; activity subsequently regulated by phosphorylated eIF2; first identified as a negative regulator of GCN4 expression | Protein with a potential role in cell survival pathways, required for the diauxic growth shift; expression in mammalian cells increases survival under conditions inducing apoptosis | GTPase of the Ypt/Rab family, very similar to Ypt32p; involved in the exocytic pathway; mediates intra-Golgi traffic or the budding of post-Golgi vesicles from the trans-Golgi | Protein involved in 3' mRNA processing, interacts with Ref2p; potential Cdc28p substrate | Daughter cell-specific protein, may participate in pathways regulating cell wall metabolism; deletion affects cell separation after division and sensitivity to drugs targeted against the cell wall | Receptor for alpha-factor pheromone; seven transmembrane-domain GPCR that interacts with both pheromone and a heterotrimeric G protein to initiate the signaling response that leads to mating between haploid a and alpha cells | Plasma membrane H+-ATPase, pumps protons out of the cell; major regulator of cytoplasmic pH and plasma membrane potential; part of the P2 subgroup of cation-transporting ATPases | Protein kinase; accumulation and phosphorylation are periodic during the cell cycle; phosphorylated in response to DNA damage; contains characteristic motifs for degradation via the APC pathway; similar to Alk2p and to mammalian haspins | Cell wall protein with similarity to glucanases; may play a role in conjugation during mating based on its regulation by Ste12p | Cell-cycle regulated activator of anaphase-promoting complex/cyclosome (APC/C), which is required for metaphase/anaphase transition; directs ubiquitination of mitotic cyclins, Pds1p, and other anaphase inhibitors; potential Cdc28p substrate | B-type cyclin involved in cell cycle progression; activates Cdc28p to promote the transition from G2 to M phase; accumulates during G2 and M, then targeted via a destruction box motif for ubiquitin-mediated degradation by the proteasome | Nucleolar protein of unknown function, positive regulator of exit from mitosis; involved in regulating the release of Cdc14p from the nucleolus in early anaphase; proposed to play similar role in meiosis | Exopolyphosphatase, hydrolyzes inorganic polyphosphate (poly P) into Pi residues; located in the cytosol, plasma membrane, and mitochondrial matrix | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Bud tip localized protein of unknown function; mRNA is targeted to the bud by a She2p dependent transport system; mRNA is cell cycle regulated via Fkh2p, peaking in G2/M phase; null mutant displays increased levels of spontaneous Rad52 foci | Subunit of the cytosolic chaperonin Cct ring complex, related to Tcp1p, required for the assembly of actin and tubulins in vivo | Subunit of a Golgi mannosyltransferase complex that also contains Anp1p, Mnn9p, Mnn10p, and Hoc1p, and mediates elongation of the polysaccharide mannan backbone; has homology to Mnn10p | Translation initiation factor eIF-5A, promotes formation of the first peptide bond; similar to and functionally redundant with Hyp2p; undergoes an essential hypusination modification; expressed under anaerobic conditions | O-glycosylated protein required for cell wall stability; attached to the cell wall via beta-1,3-glucan; mediates mitochondrial translocation of Apn1p; expression regulated by the cell integrity pathway and by Swi5p during the cell cycle | Protein involved in glycosylphosphatidylinositol (GPI) anchor synthesis; multimembrane-spanning protein that localizes to the endoplasmic reticulum; highly conserved among eukaryotes | Zinc-finger inhibitor of HO transcription; mRNA is localized and translated in the distal tip of anaphase cells, resulting in accumulation of Ash1p in daughter cell nuclei and inhibition of HO expression; potential Cdc28p substrate | Putative protein of unknown function | Acetyl-coA synthetase isoform which, along with Acs1p, is the nuclear source of acetyl-coA for histone acetylation; mutants affect global transcription; required for growth on glucose; expressed under anaerobic conditions | Endochitinase, required for cell separation after mitosis; transcriptional activation during late G and early M cell cycle phases is mediated by transcription factor Ace2p | Subunit d of the five-subunit V0 integral membrane domain of vacuolar H+-ATPase (V-ATPase), an electrogenic proton pump found in the endomembrane system; stabilizes VO subunits; required for V1 domain assembly on the vacuolar membrane | V-ATPase assembly factor, functions with other V-ATPase assembly factors in the ER to efficiently assemble the V-ATPase membrane sector (V0); overproduction confers resistance to Pichia farinosa killer toxin | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; YNL058C is not an essential gene | Cell wall protein related to glucanases, possibly involved in cell wall septation; member of the SUN family | Mating pheromone a-factor, made by a cells; interacts with alpha cells to induce cell cycle arrest and other responses leading to mating; biogenesis involves C-terminal modification, N-terminal proteolysis, and export; also encoded by MFA1 | DNA-binding protein involved in either activation or repression of transcription, depending on binding site context; also binds telomere sequences and plays a role in telomeric position effect (silencing) and telomere structure | Glycosylphosphatidylinositol (GPI)-anchored cell wall endoglucanase required for proper cell separation after cytokinesis, expression is activated by Swi5p and tightly regulated in a cell cycle-dependent manner | Member of the Sir2 family of NAD(+)-dependent protein deacetylases; involved along with Hst4p in telomeric silencing, cell cycle progression, radiation resistance, genomic stability and short-chain fatty acid metabolism | Daughter cell-specific protein, may help establish daughter fate | Nuclear protein, putative transcription factor required for growth of superficial pseudohyphae (which do not invade the agar substrate) but not for invasive pseudohyphal growth; may act together with Phd1p; potential Cdc28p substrate | G1 cyclin involved in regulation of the cell cycle; activates Cdc28p kinase to promote the G1 to S phase transition; late G1 specific expression depends on transcription factor complexes, MBF (Swi6p-Mbp1p) and SBF (Swi6p-Swi4p) | Phosphatidylinositol synthase, required for biosynthesis of phosphatidylinositol, which is a precursor for polyphosphoinositides, sphingolipids, and glycolipid anchors for some of the plasma membrane proteins | Protein involved in the regulation of cell wall synthesis; proposed to be involved in coordinating cell cycle progression with cell wall integrity | Phosphorylated protein of the mitochondrial outer membrane, localizes only to mitochondria of the bud; interacts with Myo2p to mediate mitochondrial distribution to buds; mRNA is targeted to the bud via the transport system involving She2p |
| 163 | 24 | 448 | -0.012 | 0.496 |  |  |  |  |  | YBR001C YDR001C YDR358W YFL054C YJR039W YKL188C YKR058W YLR267W YMR084W YMR104C YBR046C YBR147W YDL199C YDL215C YFR015C YGR141W YGR205W YIL107C YMR196W YNL009W YOL153C YPL165C YPL166W YPR026W | NTH2 NTH1 GGA1 PAT1 GLG1 BOP2 YPK2 ZTA1 GDH2 GSY1 VPS62 PFK26 IDP3 SET6 ATG29 ATH1 | Putative neutral trehalase, required for thermotolerance and may mediate resistance to other cellular stresses | Neutral trehalase, degrades trehalose; required for thermotolerance and may mediate resistance to other cellular stresses; may be phosphorylated by Cdc28p | Golgi-localized protein with homology to gamma-adaptin, interacts with and regulates Arf1p and Arf2p in a GTP-dependent manner in order to facilitate traffic through the late Golgi | Putative channel-like protein; similar to Fps1p; mediates passive diffusion of glycerol in the presence of ethanol | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Topoisomerase II-associated deadenylation-dependent mRNA-decapping factor; also required for faithful chromosome transmission, maintenance of rDNA locus stability, and protection of mRNA 3'-UTRs from trimming; functionally linked to Pab1p | Self-glucosylating initiator of glycogen synthesis, also glucosylates n-dodecyl-beta-D-maltoside; similar to mammalian glycogenin | Protein of unknown function, overproduction suppresses a pam1 slv3 double null mutation | Putative protein of unknown function; YMR084W and adjacent ORF YMR085W are merged in related strains | Protein kinase with similarityto serine/threonine protein kinase Ypk1p; functionally redundant with YPK1 at the genetic level; participates in a signaling pathway required for optimal cell wall integrity; homolog of mammalian kinase SGK | Zeta-crystallin homolog, found in the cytoplasm and nucleus; has similarity to E. coli quinone oxidoreductase and to human zeta-crystallin, which has quinone oxidoreductase activity | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays fluconazole resistance | Putative transporter, member of the sugar porter family | NAD(+)-dependent glutamate dehydrogenase, degrades glutamate to ammonia and alpha-ketoglutarate; expression sensitive to nitrogen catabolite repression and intracellular ammonia levels | Glycogen synthase with similarity to Gsy2p, the more highly expressed yeast homolog; expression induced by glucose limitation, nitrogen starvation, environmental stress, and entry into stationary phase | Vacuolar protein sorting (VPS) protein required for cytoplasm to vacuole targeting of proteins | ATP-binding protein of unknown function; crystal structure resembles that of E.coli pantothenate kinase and other small kinases | 6-phosphofructo-2-kinase, inhibited by phosphoenolpyruvate and sn-glycerol 3-phosphate, has negligible fructose-2,6-bisphosphatase activity, transcriptional regulation involves protein kinase A | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YMR196W is not an essential gene | Peroxisomal NADP-dependent isocitrate dehydrogenase, catalyzes oxidation of isocitrate to alpha-ketoglutarate with the formation of NADP(H+), required for growth on unsaturated fatty acids | Protein of unknown function; deletion heterozygote is sensitive to compounds that target ergosterol biosynthesis, may be involved in compound availability | Protein specifically required for autophagy; may function in autophagosome formation at the pre-autophagosomal structure in collaboration with other autophagy proteins | Acid trehalase required for utilization of extracellular trehalose |
| 32 | 43 | 436 | -0.010 | 0.495 | 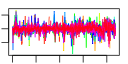 |  | 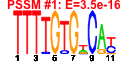 |  |  | YAL018C YBL013W YBR019C YBR180W YBR250W YCL048W YDL087C YDR010C YDR042C YDR048C YDR218C YDR278C YDR522C YER106W YFL012W YFL040W YFR012W YFR032C YGL138C YGL230C YGR107W YGR225W YHL041W YHR125W YHR184W YLR365W YPL200W YBR184W YDL154W YDL186W YDR285W YDR344C YDR374C YER179W YFR023W YGL033W YGL182C YGL183C YGR059W YHR093W YHR124W YIR013C YLL046C | FMT1 GAL10 DTR1 SPO23 YCL048W-A LUC7 SPR28 SPS2 MAM1 AMA1 SSP1 CSM4 MSH5 ZIP1 DMC1 PES4 HOP2 MND1 SPR3 NDT80 GAT4 RNP1 | Putative protein of unknown function | Methionyl-tRNA formyltransferase, catalyzes the formylation of initiator Met-tRNA in mitochondria; potential Cdc28p substrate | UDP-glucose-4-epimerase, catalyzes the interconversion of UDP-galactose and UDP-D-glucose in galactose metabolism; also catalyzes the conversion of alpha-D-glucose or alpha-D-galactose to their beta-anomers | Putative dityrosine transporter, required for spore wall synthesis; expressed during sporulation; member of the major facilitator superfamily (DHA1 family) of multidrug resistance transporters | Protein of unknown function; associates with meiosis-specific protein Spo1p | Essential protein associated with the U1 snRNP complex; splicing factor involved in recognition of 5' splice site; contains two zinc finger motifs; N-terminal zinc finger binds pre-mRNA | Putative protein of unknown function; expression is increased in ssu72-ts69 mutant | Sporulation-specific homolog of the yeast CDC3/10/11/12 family of bud neck microfilament genes; meiotic septin expressed at high levels during meiotic divisions and ascospore formation | Protein expressed during sporulation, redundant with Sps22p for organization of the beta-glucan layer of the spore wall; S. pombe ortholog is a spore wall component | Monopolin, kinetochore associated protein involved in chromosome attachment to meiotic spindle | Putative protein of unknown function; transcribed during sporulation; null mutant exhibits increased resistance to rapamycin | Putative transporter, member of the sugar porter family; YFL040W is not an essential gene | Putative protein of unknown function; transcribed during sporulation; YFR032C is not an essential gene | Putative protein of unknown function; has no significant sequence similarity to any known protein | Putative protein of unknown function; non-essential gene | Activator of meiotic anaphase promoting complex (APC/C); Cdc20p family member; required for initiation of spore wall assembly; required for Clb1p degradation during meiosis | Protein involved in the control of meiotic nuclear division and coordination of meiosis with spore formation; transcription is induced midway through meiosis | Protein required for accurate chromosome segregation during meiosis | Putative protein of unknown function; YBR184W is not an essential gene | Protein of the MutS family, forms a dimer with Msh4p that facilitates crossovers between homologs during meiosis; msh5-Y823H mutation confers tolerance to DNA alkylating agents; homologs present in C. elegans and humans | Putative protein of unknown function; YDL186W is not an essential gene | Transverse filament protein of the synaptonemal complex; required for normal levels of meiotic recombination and pairing between homologous chromosome during meiosis; potential Cdc28p substrate | Meiosis-specific protein required for repair of double-strand breaks and pairing between homologous chromosomes; homolog of Rad51p and the bacterial RecA protein | Poly(A) binding protein, suppressor of DNA polymerase epsilon mutation, similar to Mip6p | Meiosis-specific protein that localizes to chromosomes, preventing synapsis between nonhomologous chromosomes and ensuring synapsis between homologs; complexes with Mnd1p to promote homolog pairing and meiotic double-strand break repair | Protein required for recombination and meiotic nuclear division; forms a complex with Hop2p, which is involved in chromosome pairing and repair of meiotic double-strand breaks | Sporulation-specific homolog of the yeast CDC3/10/11/12 family of bud neck microfilament genes; septin protein involved in sporulation; regulated by ABFI | Meiosis-specific transcription factor required for exit from pachytene and for full meiotic recombination; activates middle sporulation genes; competes with Sum1p for binding to promoters containing middle sporulation elements (MSE) | Protein containing GATA family zinc finger motifs | Ribonucleoprotein that contains two RNA recognition motifs (RRM) |
| 61 | 30 | 458 | 0.000 | 0.520 | 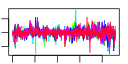 |  | 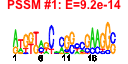 | 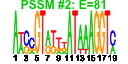 |  | YBL095W YDR223W YDR306C YDR338C YER039C YGL166W YGL205W YGR131W YGR133W YHR180W YIL006W YIL007C YIL160C YJL132W YJR019C YJR020W YLR070C YMR068W YNL116W YNL202W YNL203C YNL270C YPL222W YPR001W YCL038C YDR022C YGL208W YGR225W YIL155C YOR180C | CRF1 HVG1 CUP2 POX1 PEX4 YIA6 NAS2 POT1 TES1 XYL2 AVO2 DMA2 SPS19 APL1 FMP40 CIT3 ATG22 CIS1 SIP2 AMA1 GUT2 DCI1 | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Transcriptional corepressor involved in the regulation of ribosomal protein gene transcription via the TOR signaling pathway and protein kinase A, phosphorylated by activated Yak1p which promotes accumulation of Crf1p in the nucleus | F-box protein of unknown function; interacts with Sgt1p via a Leucine-Rich Repeat (LRR) domain | hypothetical protein | Protein of unknown function, has homology to Vrg4p | Copper-binding transcription factor; activates transcription of the metallothionein genes CUP1-1 and CUP1-2 in response to elevated copper concentrations | Fatty-acyl coenzyme A oxidase, involved in the fatty acid beta-oxidation pathway; localized to the peroxisomal matrix | Protein of unknown function; expression induced in response to ketoconazole; promoter region contains a sterol regulatory element motif, which has been identified as a Upc2p-binding site | Peroxisomal ubiquitin conjugating enzyme required for peroxisomal matrix protein import and peroxisome biogenesis | Mitochondrial NAD+ transporter, involved in the transport of NAD+ into the mitochondria (see also YEA6); member of the mitochondrial carrier subfamily; disputed role as a pyruvate transporter; has putative mouse and human orthologs | Protein with similarity to the p27 subunit of mammalian proteasome modulator; not essential; interacts with Rpn4p | 3-ketoacyl-CoA thiolase with broad chain length specificity, cleaves 3-ketoacyl-CoA into acyl-CoA and acetyl-CoA during beta-oxidation of fatty acids | Putative protein of unknown function; localizes to the membrane fraction; possible Zap1p-regulated target gene induced by zinc deficiency; YJL132W is a non-essential gene | Peroxisomal acyl-CoA thioesterase likely to be involved in fatty acid oxidation rather than fatty acid synthesis; conserved protein also found in human peroxisomes; TES1 mRNA levels increase during growth on fatty acids | Xylitol dehydrogenase, converts xylitol to D-xylulose in the endogenous xylose utilization pathway | Component of a complex containing the Tor2p kinase and other proteins, which may have a role in regulation of cell growth | Protein involved in regulating spindle position and orientation, functionally redundant with Dma1p; homolog of S. pombe Dma1 and H. sapiens Chfr | Peroxisomal 2,4-dienoyl-CoA reductase, auxiliary enzyme of fatty acid beta-oxidation; homodimeric enzyme required for growth and sporulation on petroselineate medium; expression induced during late sporulation and in the presence of oleate | Beta-adaptin, large subunit of the clathrin associated protein complex (AP-2); involved in vesicle mediated transport; similar to mammalian beta-chain of the clathrin associated protein complex | Dual specificity mitochondrial citrate and methylcitrate synthase; catalyzes the condensation of acetyl-CoA and oxaloacetate to form citrate and that of propionyl-CoA and oxaloacetate to form 2-methylcitrate | Protein required for the breakdown of autophagic vesicles in the vacuole during autophagy, putative integral membrane protein that localizes to vacuolar membranes and punctate structures attached to the vacuole | Protein required for autophagosome formation in concert with Atg17p; may be involved in microtubule organization; high-copy suppressor of CIK1 deletion | One of three beta subunits of the Snf1 serine/threonine protein kinase complex involved in the response to glucose starvation; null mutants exhibit accelerated aging; N-myristoylprotein localized to the cytoplasm and the plasma membrane | Activator of meiotic anaphase promoting complex (APC/C); Cdc20p family member; required for initiation of spore wall assembly; required for Clb1p degradation during meiosis | Mitochondrial glycerol-3-phosphate dehydrogenase; expression is repressed by both glucose and cAMP and derepressed by non-fermentable carbon sources in a Snf1p, Rsf1p, Hap2/3/4/5 complex dependent manner | Peroxisomal delta(3,5)-delta(2,4)-dienoyl-CoA isomerase, involved in fatty acid metabolism, contains peroxisome targeting signals at amino and carboxy termini |
| 147 | 43 | 430 | 0.001 | 0.503 |  |  | 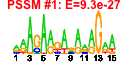 | 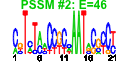 |  | YGL023C YGL090W YHR207C YJL065C YJL218W YKL017C YKL105C YKL129C YKR019C YKR020W YKR062W YLR260W YLR376C YLR424W YMR067C YMR111C YNL027W YOL112W YOR005C YOR077W YOR090C YOR279C YPL077C YPL151C YIL089W YKL079W YLL058W YLR035C YLR417W YML107C YML121W YMR009W YMR028W YMR124W YMR213W YNL011C YNL116W YNR005C YOL066C YOL070C YOL114C YPR177C YPR179C | PIB2 LIF1 SET5 DLS1 HCS1 MYO3 IRS4 VPS51 TFA2 LCB5 PSY3 SPP382 UBX4 CRZ1 MSB4 DNL4 RTS2 PTC5 RFM1 PRP46 SMY1 MLH2 VPS36 PML39 GTR1 ADI1 TAP42 CEF1 DMA2 RIB2 NBA1 HDA3 | Protein binding phosphatidylinositol 3-phosphate, involved in telomere-proximal repression of gene expression; similar to Fab1 and Vps27 | Component of the DNA ligase IV complex that mediates nonhomologous end joining in DNA double-strand break repair; physically interacts with Dnl4p and Nej1p; homologous to mammalian XRCC4 protein | Zinc-finger protein of unknown function, contains one canonical and two unusual fingers in unusual arrangements; deletion enhances replication of positive-strand RNA virus | Subunit of ISW2/yCHRAC chromatin accessibility complex along with Itc1p, Isw2p, and Dpb4p; involved in inheritance of telomeric silencing | Putative protein of unknown function, similar to bacterial galactoside O-acetyltransferases; induced by oleate in an OAF1/PIP2-dependent manner; promoter contains an oleate response element consensus sequence; non-essential gene | Hexameric DNA polymerase alpha-associated DNA helicase A involved in lagging strand DNA synthesis; contains single-stranded DNA stimulated ATPase and dATPase activities; replication protein A stimulates helicase and ATPase activities | Putative protein of unknown function | One of two type I myosins; localizes to actin cortical patches; deletion of MYO3 has little effect on growth, but myo3 myo5 double deletion causes severe defects in growth and actin cytoskeleton organization | Protein involved in regulation of phosphatidylinositol 4,5-bisphosphate concentrations; Irs4p and Tax4p bind and activate the phosphatase Inp51p; mutation confers an increase in rDNA silencing | Component of the GARP (Golgi-associated retrograde protein) complex, Vps51p-Vps52p-Vps53p-Vps54p, which is required for the recycling of proteins from endosomes to the late Golgi; links the (VFT/GARP) complex to the SNARE Tlg1p | TFIIE small subunit, involved in RNA polymerase II transcription initiation | Minor sphingoid long-chain base kinase, paralog of Lcb4p responsible for few percent of the total activity, possibly involved in synthesis of long-chain base phosphates, which function as signaling molecules | Protein of unknown function; deletion results in a mutator phenotype suggesting a role for this protein as a mutational suppressor; deletion increases sensitivity to anticancer drugs oxaliplatin and cisplatin but not mitomycin C | Essential protein that forms a dimer with Ntr2p; also forms a trimer, with Ntr2p and Prp43p, that is involved in spliceosome disassembly; found also in a multisubunit complex with the splicing factor Clf1p; suppressor of prp38-1 mutation | UBX (ubiquitin regulatory X) domain-containing protein that interacts with Cdc48p | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; YMR111C is not an essential gene | Transcription factor that activates transcription of genes involved in stress response; nuclear localization is positively regulated by calcineurin-mediated dephosphorylation | GTPase-activating protein of the Ras superfamily that acts primarily on Sec4p, localizes to the bud site and bud tip, has similarity to Msb3p; msb3 msb4 double mutation causes defects in secretion and actin organization | DNA ligase required for nonhomologous end-joining (NHEJ), forms stable heterodimer with required cofactor Lif1p, interacts with Nej1p; involved in meiosis, not essential for vegetative growth | Basic zinc-finger protein, similar to human and mouse Kin17 proteins which are chromatin-associated proteins involved in UV response and DNA replication | Mitochondrially localized type 2C protein phosphatase involved in regulation of pyruvate dehydrogenase activity; contains Mg2+/Mn2+-dependent casein phosphatase activity in vitro | DNA-binding protein required for vegetative repression of middle sporulation genes; specificity factor that directs the Hst1p histone deacetylase to some of the promoters regulated by Sum1p; involved in telomere maintenance | Putative protein of unknown function; regulates PIS1 expression; mutant displays spore wall assembly defect in ether sensitivity screen; YPL077C is not an essential gene | Splicing factor that is found in the Cef1p subcomplex of the spliceosome | Protein that interacts with Myo2p, proposed to be involved in exocytosis; N-terminal domain is related to the motor domain of kinesins | Putative protein of unknown function with similarity to Str2p, which is a cystathionine gamma-synthase important in sulfur metabolism; YLL058W is not an essential gene | Protein involved in the mismatch repair of certain frameshift intermediates and involved in meiotic recombination; forms a complex with Mlh1p | Component of the ESCRT-II complex; contains the GLUE (GRAM Like Ubiquitin binding in EAP45) domain which is involved in interactions with ESCRT-I and ubiquitin-dependent sorting of proteins into the endosome | Protein required for nuclear retention of unspliced pre-mRNAs along with Mlp1p and Pml1p; anchored to nuclear pore complex via Mlp1p and Mlp2p; found with the subset of nuclear pores farthest from the nucleolus; may interact with ribosomes | Cytoplasmic GTP binding protein and negative regulator, with homolog Gtr2p, of the Ran/Tc4 GTPase cycle; component of GSE complex, which is required for sorting of Gap1p; involved in phosphate transport; has homology to human RagA and RagB | Acireductone dioxygenease involved in the methionine salvage pathway; ortholog of human MTCBP-1; transcribed with YMR010W and regulated post-transcriptionally by RNase III (Rnt1p) cleavage; ADI1 mRNA is induced in heat shock conditions | Essential protein involved in the TOR signaling pathway; physically associates with the protein phosphatase 2A and the SIT4 protein phosphatase catalytic subunits | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery, cytoplasm, bud, and bud neck; interacts with Crm1p in two-hybrid assay; YMR124W is not an essential gene | Essential splicing factor; associated with Prp19p and the spliceosome, contains an N-terminal c-Myb DNA binding motif necessary for cell viability but not for Prp19p association, evolutionarily conserved and homologous to S. pombe Cdc5p | Putative protein of unknown function; YNL011C is not an essential gene | Protein involved in regulating spindle position and orientation, functionally redundant with Dma1p; homolog of S. pombe Dma1 and H. sapiens Chfr | DRAP deaminase, catalyzes the third step of the riboflavin biosynthesis pathway; cytoplasmic tRNA pseudouridine synthase involved in pseudouridylation of cytoplasmic tRNAs at position 32 | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery, cytoplasm, and bud neck; potential Cdc28p substrate | Putative protein of unknown function with similarity to human ICT1 and prokaryotic factors that may function in translation termination; YOL114C is not an essential gene | Subunit of a possibly tetrameric trichostatin A-sensitive class II histone deacetylase complex that contains an Hda1p homodimer and an Hda2p-Hda3p heterodimer; required for the activity of the complex; has similarity to Hda2p |
| 173 | 41 | 433 | 0.008 | 0.558 | 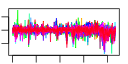 |  | 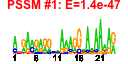 | 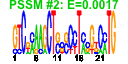 |  | YDL013W YDR275W YGR028W YGR239C YIL143C YJL004C YJR046W YJR049C YJR135C YLR010C YLR102C YLR323C YMR034C YMR035W YMR126C YMR237W YNL042W YOL110W YOL111C YPR037C YBR278W YDR013W YEL038W YFR036W YHL025W YIL144W YIL161W YJL065C YJR050W YKL005C YKL192C YLR011W YLR091W YLR375W YMR165C YNL063W YNL076W YNL097C YNL133C YOR193W YPR040W | SLX5 BSC2 MSP1 PEX21 SSL2 SYS1 TAH11 UTR1 MCM22 TEN1 APC9 CWC24 IMP2' DLT1 BCH1 YNL042W-B SHR5 MDY2 ERV2 DPB3 PSF1 UTR4 CDC26 SNF6 TID3 DLS1 ISY1 BYE1 ACP1 LOT6 STP3 PAH1 MTQ1 MKS1 YNL097C-B FYV6 PEX27 TIP41 | Subunit of the Slx5-Slx8 substrate-specific ubiquitin ligase complex; stimulated by prior attachment of SUMO to the substrate | Protein of unknown function, ORF exhibits genomic organization compatible with a translational readthrough-dependent mode of expression | Mitochondrial protein involved in sorting of proteins in the mitochondria; putative membrane-spanning ATPase | Peroxin required for targeting of peroxisomal matrix proteins containing PTS2; interacts with Pex7p; partially redundant with Pex18p | Component of the holoenzyme form of RNA polymerase transcription factor TFIIH, has DNA-dependent ATPase/helicase activity and is required, with Rad3p, for unwinding promoter DNA; involved in DNA repair; homolog of human ERCC3 | Integral membrane protein of the Golgi required for targeting of the Arf-like GTPase Arl3p to the Golgi; multicopy suppressor of ypt6 null mutation | DNA replication licensing factor, required for pre-replication complex assembly | ATP-NADH kinase; phosphorylates both NAD and NADH; active as a hexamer; enhances the activity of ferric reductase (Fre1p) | Protein involved in minichromosome maintenance; component of the kinetochore; binds to centromeric DNA in a Ctf19p-dependent manner | Protein that regulates telomeric length; protects telomeric ends in a complex with Cdc13p and Stn1p | Subunit of the Anaphase-Promoting Complex/Cyclosome (APC/C), which is a ubiquitin-protein ligase required for degradation of anaphase inhibitors, including mitotic cyclins, during the metaphase/anaphase transition | Essential protein, component of a complex containing Cef1p; has similarity to S. pombe Cwf24p | Putative transporter, member of the SLC10 carrier family; identified in a transposon mutagenesis screen as a gene involved in azole resistance; YMR034C is not an essential gene | Transcriptional activator involved in maintenance of ion homeostasis and protection against DNA damage caused by bleomycin and other oxidants, contains a C-terminal leucine-rich repeat | Protein of unknown function, mutant sensitive to 6-azauracil (6AU) and mycophenolic acid (MPA) | Member of the ChAPs family (Chs5p-Arf1p-binding proteins: Bch1p, Bch2p, Bud7p, Chs6p), that forms the exomer complex with Chs5p to mediate export of specific cargo proteins from the Golgi to the plasma membrane; may interact with ribosomes | Putative protein of unknown function | Subunit of a palmitoyltransferase, composed of Shr5p and Erf2p, that adds a palmitoyl lipid moiety to heterolipidated substrates such as Ras1p and Ras2p through a thioester linkage; palmitoylation is required for Ras2p membrane localization | Protein required for efficient mating; involved in shmoo formation and nuclear migration in the pre-zygote; associates with ribosomes and interacts with YOR164C; contains a ubiquitin-like (UBL) domain | Flavin-linked sulfhydryl oxidase localized to the endoplasmic reticulum lumen, involved in disulfide bond formation within the ER | Third-largest subunit of DNA polymerase II (DNA polymerase epsilon), required to maintain fidelity of chromosomal replication and also for inheritance of telomeric silencing; mRNA abundance peaks at the G1/S boundary of the cell cycle | Subunit of the GINS complex (Sld5p, Psf1p, Psf2p, Psf3p), which is localized to DNA replication origins and implicated in assembly of the DNA replication machinery | Protein of unknown function with sequence similarity to 2,3-diketo-5-methylthiopentyl-1-phosphate enolase-phosphatases, found in both the cytoplasm and nucleus | Subunit of the SWI/SNF chromatin remodeling complex involved in transcriptional regulation; functions interdependently in transcriptional activation with Snf2p and Snf5p | Component of the evolutionarily conserved kinetochore-associated Ndc80 complex (Ndc80p-Nuf2p-Spc24p-Spc25p); conserved coiled-coil protein involved in chromosome segregation, spindle checkpoint activity, kinetochore assembly and clustering | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; mRNA is enriched in Scp160p-associated mRNPs; YIL161W is a non-essential gene | Subunit of ISW2/yCHRAC chromatin accessibility complex along with Itc1p, Isw2p, and Dpb4p; involved in inheritance of telomeric silencing | Component of the spliceosome complex involved in pre-mRNA splicing, auxiliary splicing factor that may modulate Syf1p activity and help optimize splicing; isy1 syf2 double mutation activates the spindle checkpoint, causing cell cycle arrest | Negative regulator of transcription elongation, contains a TFIIS-like domain and a PHD finger, multicopy suppressor of temperature-sensitive ess1 mutations, probably binds RNA polymerase II large subunit | Mitochondrial matrix acyl carrier protein, involved in biosynthesis of octanoate, which is a precursor to lipoic acid; activated by phosphopantetheinylation catalyzed by Ppt2p | FMN-dependent NAD(P)H:quinone reductase that may be involved in quinone detoxification; gene expression increases in cultures shifted to a lower temperature | Protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YLR091W is not an esssential gene | Zinc-finger protein of unknown function, possibly involved in pre-tRNA splicing and in uptake of branched-chain amino acids | Mg2+-dependent phosphatidate (PA) phosphatase, catalyzes the dephosphorylation of PA to yield diacylglycerol and Pi, responsible for de novo lipid synthesis; homologous to mammalian lipin 1 | S-adenosylmethionine-dependent methyltransferase; methylates translational release factor Mrf1p; similar to E.coli PrmC; is not an essential gene | Pleiotropic negative transcriptional regulator involved in Ras-CAMP and lysine biosynthetic pathways and nitrogen regulation; involved in retrograde (RTG) mitochondria-to-nucleus signaling | Protein of unknown function, required for survival upon exposure to K1 killer toxin; proposed to regulate double-strand break repair via non-homologous end-joining | Peripheral peroxisomal membrane protein involved in controlling peroxisome size and number, interacts with homologous protein Pex25p | Protein that interacts physically and genetically with Tap42p, which regulates protein phosphatase 2A; component of the TOR (target of rapamycin) signaling pathway |
| 29 | 39 | 436 | 0.011 | 0.543 | 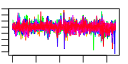 |  | 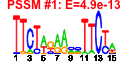 | 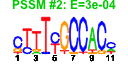 |  | YAL004W YBL058W YBR082C YBR101C YBR170C YCL050C YDL020C YDR214W YER103W YER142C YFL016C YGL062W YGR136W YGR137W YGR142W YIL075C YJR045C YLL024C YLR216C YLR217W YLR259C YML130C YMR186W YNL006W YNL007C YNL064C YNL077W YNL193W YNL281W YOR020C YOR027W YOR117W YPL106C YPL240C YAL005C YDR155C YER143W YGR211W YPR158W | SHP1 UBC4 FES1 NPL4 APA1 RPN4 AHA1 SSA4 MAG1 MDJ1 PYC1 LSB1 BTN2 RPN2 PMR1 SSA2 CPR6 HSP60 ERO1 HSC82 LST8 SIS1 YDJ1 APJ1 HCH1 HSP10 STI1 RPT5 SSE1 SSA1 CPR1 DDI1 ZPR1 | UBX (ubiquitin regulatory X) domain-containing protein that regulates Glc7p phosphatase activity and interacts with Cdc48p; interacts with ubiquitylated proteins in vivo and is required for degradation of a ubiquitylated model substrate | Ubiquitin-conjugating enzyme (E2), mediates degradation of short-lived and abnormal proteins; interacts with E3-CaM in ubiquitinating calmodulin; interacts with many SCF ubiquitin protein ligases; component of the cellular stress response | Hsp70 (Ssa1p) nucleotide exchange factor, cytosolic homolog of Sil1p, which is the nucleotide exchange factor for BiP (Kar2p) in the endoplasmic reticulum | Endoplasmic reticulum and nuclear membrane protein, forms a complex with Cdc48p and Ufd1p that recognizes ubiquitinated proteins in the endoplasmic reticulum and delivers them to the proteasome for degradation | Diadenosine 5',5''-P1,P4-tetraphosphate phosphorylase I (AP4A phosphorylase), involved in catabolism of bis(5'-nucleosidyl) tetraphosphates; has similarity to Apa2p | Transcription factor that stimulates expression of proteasome genes; Rpn4p levels are in turn regulated by the 26S proteasome in a negative feedback control mechanism; RPN4 is transcriptionally regulated by various stress responses | Co-chaperone that binds to Hsp82p and activates its ATPase activity; similar to Hch1p; expression is regulated by stresses such as heat shock | Heat shock protein that is highly induced upon stress; plays a role in SRP-dependent cotranslational protein-membrane targeting and translocation; member of the HSP70 family; cytoplasmic protein that concentrates in nuclei upon starvation | 3-methyl-adenine DNA glycosylase involved in protecting DNA against alkylating agents; initiates base excision repair by removing damaged bases to create abasic sites that are subsequently repaired | Protein involved in folding of mitochondrially synthesized proteins in the mitochondrial matrix; localizes to the mitochondrial inner membrane; member of the DnaJ family of molecular chaperones | Pyruvate carboxylase isoform, cytoplasmic enzyme that converts pyruvate to oxaloacetate; highly similar to isoform Pyc2p but differentially regulated; mutations in the human homolog are associated with lactic acidosis | Protein containing an N-terminal SH3 domain; binds Las17p, which is a homolog of human Wiskott-Aldrich Syndrome protein involved in actin patch assembly and actin polymerization | v-SNARE binding protein that facilitates specific protein retrieval from a late endosome to the Golgi; modulates arginine uptake, possible role in mediating pH homeostasis between the vacuole and plasma membrane H(+)-ATPase | Subunit of the 26S proteasome, substrate of the N-acetyltransferase Nat1p | High affinity Ca2+/Mn2+ P-type ATPase required for Ca2+ and Mn2+ transport into Golgi; involved in Ca2+ dependent protein sorting and processing; mutations in human homolog ATP2C1 cause acantholytic skin condition Hailey-Hailey disease | ATP binding protein involved in protein folding and vacuolar import of proteins; member of heat shock protein 70 (HSP70) family; associated with the chaperonin-containing T-complex; present in the cytoplasm, vacuolar membrane and cell wall | Peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; binds to Hsp82p and contributes to chaperone activity | Tetradecameric mitochondrial chaperonin required for ATP-dependent folding of precursor polypeptides and complex assembly; prevents aggregation and mediates protein refolding after heat shock; role in mtDNA transmission; phosphorylated | Thiol oxidase required for oxidative protein folding in the endoplasmic reticulum | Cytoplasmic chaperone of the Hsp90 family, redundant in function and nearly identical with Hsp82p, and together they are essential; expressed constitutively at 10-fold higher basal levels than HSP82 and induced 2-3 fold by heat shock | Protein required for the transport of amino acid permease Gap1p from the Golgi to the cell surface; component of the TOR signaling pathway; associates with both Tor1p and Tor2p; contains a WD-repeat | Type II HSP40 co-chaperone that interacts with the HSP70 protein Ssa1p; not functionally redundant with Ydj1p due to due to substrate specificity; shares similarity with bacterial DnaJ proteins | Protein chaperone involved in regulation of the HSP90 and HSP70 functions; involved in protein translocation across membranes; member of the DnaJ family | Putative chaperone of the HSP40 (DNAJ) family; overexpression interferes with propagation of the [Psi+] prion; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; exhibits a two-hybrid interaction with Yhr151cp in a large-scale analysis | Heat shock protein regulator that binds to Hsp90p and may stimulate ATPase activity; originally identified as a high-copy number suppressor of a HSP90 loss-of-function mutation; GFP-fusion protein localizes to the cytoplasm and nucleus | Mitochondrial matrix co-chaperonin that inhibits the ATPase activity of Hsp60p, a mitochondrial chaperonin; involved in protein folding and sorting in the mitochondria; 10 kD heat shock protein with similarity to E. coli groES | Hsp90 cochaperone, interacts with the Ssa group of the cytosolic Hsp70 chaperones; activates the ATPase activity of Ssa1p; homolog of mammalian Hop protein | One of six ATPases of the 19S regulatory particle of the 26S proteasome involved in the degradation of ubiquitinated substrates; recruited to the GAL1-10 promoter region upon induction of transcription | ATPase that is a component of the heat shock protein Hsp90 chaperone complex; binds unfolded proteins; member of the heat shock protein 70 (HSP70) family; localized to the cytoplasm | ATPase involved in protein folding and nuclear localization signal (NLS)-directed nuclear transport; member of heat shock protein 70 (HSP70) family; forms a chaperone complex with Ydj1p; localized to the nucleus, cytoplasm, and cell wall | Cytoplasmic peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; binds the drug cyclosporin A | DNA damage-inducible v-SNARE binding protein, contains a ubiquitin-associated (UBA) domain, may act as a negative regulator of constitutive exocytosis, may play a role in S-phase checkpoint control | Essential protein with two zinc fingers, present in the nucleus of growing cells but relocates to the cytoplasm in starved cells via a process mediated by Cpr1p; binds to translation elongation factor eEF-1 (Tef1p) | Putative protein of unknown function |
| 275 | 24 | 440 | 0.015 | 0.443 |  |  |  |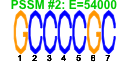  |  | YPR193C YAL034C YBR050C YBR285W YDL024C YDR018C YDR436W YFL030W YFR045W YGL096W YGR053C YGR110W YHR160C YIL045W YJL089W YJR158W YKL093W YKL188C YLR164W YLR174W YNR002C YOR019W YOR031W YPR061C | HPA2 FUN19 REG2 DIA3 PPZ2 AGX1 TOS8 PEX18 PIG2 SIP4 HXT16 MBR1 PAT1 IDP2 ATO2 CRS5 JID1 | Tetrameric histone acetyltransferase with similarity to Gcn5p, Hat1p, Elp3p, and Hpa3p; acetylates histones H3 and H4 in vitro and exhibits autoacetylation activity | Non-essential protein of unknown function | Regulatory subunit of the Glc7p type-1 protein phosphatase; involved with Reg1p, Glc7p, and Snf1p in regulation of glucose-repressible genes, also involved in glucose-induced proteolysis of maltose permease | Putative protein of unknown function; YBR285W is not an essential gene and deletion of YBR285W leads to poor growth on glucose-minimal medium at 15C | Protein of unknown function, involved in invasive and pseudohyphal growth | Probable membrane protein with three predicted transmembrane domains; homologous to Ybr042cp, similar to C. elegans F55A11.5 and maize 1-acyl-glycerol-3-phosphate acyltransferase; null exhibits no apparent phenotype | Serine/threonine protein phosphatase Z, isoform of Ppz1p; involved in regulation of potassium transport, which affects osmotic stability, cell cycle progression, and halotolerance | Alanine:glyoxylate aminotransferase (AGT), catalyzes the synthesis of glycine from glyoxylate, which is one of three pathways for glycine biosynthesis in yeast; has similarity to mammalian and plant alanine:glyoxylate aminotransferases | Putative mitochondrial transport protein; null mutant is viable, exhibits decreased levels of chitin and normal resistance to calcofluor white | Homeodomain-containing transcription factor; SBF regulated target gene that in turn regulates expression of genes involved in G1/S phase events such as bud site selection, bud emergence and cell cycle progression; similarity to Cup9p | Putative protein of unknown function | Putative protein of unknown function; transcription is increased in response to genotoxic stress | Peroxin required for targeting of peroxisomal matrix proteins containing PTS2; interacts with Pex7p; partially redundant with Pex21p | Putative type-1 protein phosphatase targeting subunit that tethers Glc7p type-1 protein phosphatase to Gsy2p glycogen synthase | C6 zinc cluster transcriptional activator that binds to the carbon source-responsive element (CSRE) of gluconeogenic genes; involved in the positive regulation of gluconeogenesis; regulated by Snf1p protein kinase; localized to the nucleus | Protein of unknown function with similarity to hexose transporter family members, expression is repressed by high levels of glucose | Protein involved in mitochondrial functions and stress response; overexpression suppresses growth defects of hap2, hap3, and hap4 mutants | Topoisomerase II-associated deadenylation-dependent mRNA-decapping factor; also required for faithful chromosome transmission, maintenance of rDNA locus stability, and protection of mRNA 3'-UTRs from trimming; functionally linked to Pab1p | Mitochondrial inner membrane of unknown function; similar to Tim18p and Sdh4p; expression induced by nitrogen limitation in a GLN3, GAT1-dependent manner | Cytosolic NADP-specific isocitrate dehydrogenase, catalyzes oxidation of isocitrate to alpha-ketoglutarate; levels are elevated during growth on non-fermentable carbon sources and reduced during growth on glucose | Putative transmembrane protein involved in export of ammonia, a starvation signal that promotes cell death in aging colonies; phosphorylated in mitochondria; member of the TC 9.B.33 YaaH family; homolog of Ady2p and Y. lipolytica Gpr1p | Protein of unknown function that may interact with ribosomes, based on co-purification experiments | Copper-binding metallothionein, required for wild-type copper resistance | Probable Hsp40p co-chaperone, has a DnaJ-like domain and appears to be involved in ER-associated degradation of misfolded proteins containing a tightly folded cytoplasmic domain; inhibits replication of Brome mosaic virus in S. cerevisiae |
| 218 | 44 | 440 | 0.033 | 0.502 |  |  | 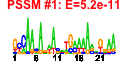 | 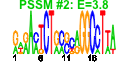 |  | YOL134C YOR041C YPL005W YPL035C YPL071C YPL099C YPR003C YPR128C YFL013C YGR007W YHR060W YJL099W YJR005W YKL010C YKR019C YKR020W YLR207W YLR330W YLR363C YLR371W YLR418C YLR426W YMR004W YMR037C YMR064W YMR066W YNL020C YNL118C YNL214W YNL223W YNL224C YOL063C YOL107W YOL113W YOL133W YOR037W YOR069W YOR141C YOR353C YOR357C YPL077C YPL151C YPR111W YPR131C | AEP3 AIM43 PDR1 IES1 MUQ1 VMA22 CHS6 APL1 UFD4 IRS4 VPS51 HRD3 CHS5 NMD4 ROM2 CDC73 MVP1 MSN2 AEP1 SOV1 ARK1 DCP2 PEX17 ATG4 SQS1 CRT10 SKM1 TEC1 CYC2 VPS5 ARP8 SOG2 SNX3 PRP46 DBF20 NAT3 | Peripheral mitochondrial inner membrane protein, located on the matrix face of the membrane; stabilizes the bicistronic AAP1-ATP6 mRNA encoding subunits 6 and 8 of the ATP synthase complex | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Protein of unknown function;non-tagged protein detected in purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss and severe growth defect in minimal glycerol media | Putative sulfate permease; physically interacts with Hsp82p; green fluorescent protein (GFP)-fusion protein localizes to the ER; YPR003C is not an essential gene | Zinc cluster protein that is a master regulator involved in recruiting other zinc cluster proteins to pleiotropic drug response elements (PDREs) to fine tune the regulation of multidrug resistance genes | Subunit of the INO80 chromatin remodeling complex | Choline phosphate cytidylyltransferase, catalyzes the second step of phosphatidylethanolamine biosynthesis; involved in the maintenance of plasma membrane; similar to mammalian CTP: phosphocholine cytidylyl-transferases | Integral membrane protein that is required for vacuolar H+-ATPase (V-ATPase) function, although not an actual component of the V-ATPase complex; functions in the assembly of the V-ATPase; localized to the yeast endoplasmic reticulum (ER) | Member of the ChAPs family of proteins (Chs5p-Arf1p-binding proteins: Bch1p, Bch2p, Bud7p, Chs6p), that forms the exomer complex with Chs5p to mediate export of specific cargo proteins, including Chs3p, from the Golgi to the plasma membrane | Beta-adaptin, large subunit of the clathrin associated protein complex (AP-2); involved in vesicle mediated transport; similar to mammalian beta-chain of the clathrin associated protein complex | Ubiquitin-protein ligase (E3) that interacts with Rpt4p and Rpt6p, two subunits of the 19S particle of the 26S proteasome; cytoplasmic E3 involved in the degradation of ubiquitin fusion proteins | Protein involved in regulation of phosphatidylinositol 4,5-bisphosphate concentrations; Irs4p and Tax4p bind and activate the phosphatase Inp51p; mutation confers an increase in rDNA silencing | Component of the GARP (Golgi-associated retrograde protein) complex, Vps51p-Vps52p-Vps53p-Vps54p, which is required for the recycling of proteins from endosomes to the late Golgi; links the (VFT/GARP) complex to the SNARE Tlg1p | Resident protein of the ER membrane that plays a central role in ER-associated protein degradation (ERAD), forms HRD complex with Hrd1p and ERAD determinants that engages in lumen to cytosol communication and coordination of ERAD events | Component of the exomer complex, which also contains Csh6p, Bch1p, Bch2p, and Bud7p and is involved in export of selected proteins, such as chitin synthase Chs3p, from the Golgi to the plasma membrane | Protein interacting with Nam7p, may be involved in the nonsense-mediated mRNA decay pathway | GDP/GTP exchange protein (GEP) for Rho1p and Rho2p; mutations are synthetically lethal with mutations in rom1, which also encodes a GEP | Constituent of Paf1 complex with RNA polymerase II, Paf1p, Hpr1p, Ctr9, Leo1, Rtf1 and Ccr4p, distinct from Srb-containing Pol II complexes; required for expression of certain genes, modification of some histones, and telomere maintenance | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; proposed to be involved in resistance to mechlorethamine and streptozotocin | Protein required for sorting proteins to the vacuole; overproduction of Mvp1p suppresses several dominant VPS1 mutations; Mvp1p and Vps1p act in concert to promote membrane traffic to the vacuole | Transcriptional activator related to Msn4p; activated in stress conditions, which results in translocation from the cytoplasm to the nucleus; binds DNA at stress response elements of responsive genes, inducing gene expression | Protein required for expression of the mitochondrial OLI1 gene encoding subunit 9 of F1-F0 ATP synthase | Mitochondrial protein of unknown function | Serine/threonine protein kinase involved in regulation of the cortical actin cytoskeleton; involved in control of endocytosis | Catalytic subunit of the Dcp1p-Dcp2p decapping enzyme complex, which removes the 5' cap structure from mRNAs prior to their degradation; member of the Nudix hydrolase family | Peroxisomal membrane peroxin and subunit of the docking complex that facilitates the import of peroxisomal matrix proteins; required for peroxisome biogenesis | Cysteine protease required for autophagy; cleaves Atg8p to a form required for autophagosome and Cvt vesicle generation; mediates attachment of autophagosomes to microtubules through interactions with Tub1p and Tub2p | Protein of unknown function; overexpression antagonizes the suppression of splicing defects by spp382 mutants; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Protein involved in transcriptional regulation of RNR2 and RNR3; expression of the gene is induced by DNA damage and null mutations confer increased resistance to hydroxyurea; N-terminal region has a leucine repeat and a WD40 repeat | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and colocalizes in a punctate pattern with the early golgi/COPI vesicles; YOL107W is not an essential protein | Member of the PAK family of serine/threonine protein kinases with similarity to Ste20p and Cla4p; proposed to be a downstream effector of Cdc42p during polarized growth | Transcription factor required for full Ty1 epxression, Ty1-mediated gene activation, and haploid invasive and diploid pseudohyphal growth; TEA/ATTS DNA-binding domain family member | Mitochondrial peripheral inner membrane protein, contains a FAD cofactor in a domain exposed in the intermembrane space; exhibits redox activity in vitro; likely participates in ligation of heme to acytochromes c and c1 (Cyc1p and Cyt1p) | Nexin-1 homolog required for localizing membrane proteins from a prevacuolar/late endosomal compartment back to the late Golgi apparatus; structural component of the retromer membrane coat complex; forms a retromer subcomplex with Vps17p | Nuclear actin-related protein involved in chromatin remodeling, component of chromatin-remodeling enzyme complexes | Key component of the RAM signaling network, required for proper cell morphogenesis and cell separation after mitosis | Sorting nexin required to maintain late-Golgi resident enzymes in their proper location by recycling molecules from the prevacuolar compartment; contains a PX domain and sequence similarity to human Snx3p | Putative protein of unknown function; regulates PIS1 expression; mutant displays spore wall assembly defect in ether sensitivity screen; YPL077C is not an essential gene | Splicing factor that is found in the Cef1p subcomplex of the spliceosome | Ser/Thr kinase involved in late nuclear division, one of the mitotic exit network (MEN) proteins; necessary for the execution of cytokinesis | Catalytic subunit of the NatB N-terminal acetyltransferase, which catalyzes acetylation of the amino-terminal methionine residues of all proteins beginning with Met-Asp or Met-Glu and of some proteins beginning with Met-Asn or Met-Met |
| 39 | 48 | 438 | 0.042 | 0.541 |  |  | 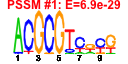 | 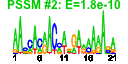 |  | YBR073W YBR245C YCL024W YCL060C YCL061C YDR097C YDR144C YDR279W YDR325W YFR005C YJL074C YJL095W YJL168C YJL181W YJR043C YJR092W YKL042W YKL045W YKL101W YKL113C YLL021W YLR103C YLR154C YLR383W YLR410W YML021C YML060W YML061C YML065W YML109W YMR076C YMR078C YMR179W YNL082W YNL088W YNL102W YNL126W YNL166C YNL312W YNR009W YOR144C YOR368W YPL032C YPL255W YPL267W YPR120C YPR186C YDR507C | RDH54 ISW1 KCC4 MRC1 MSH6 MKC7 RNH202 YCG1 SAD1 SMC3 BCK1 SET2 POL32 BUD4 SPC42 PRI2 HSL1 FEN1 SPA2 CDC45 RNH203 SMC6 VIP1 UNG1 OGG1 PIF1 ORC1 ZDS2 PDS5 CTF18 SPT21 PMS1 TOP2 POL1 SPC98 BNI5 RFA2 NRM1 ELG1 RAD17 SVL3 BBP1 ACM1 CLB5 PZF1 GIN4 | DNA-dependent ATPase, stimulates strand exchange by modifying the topology of double-stranded DNA; involved in recombinational repair of DNA double-strand breaks during mitosis and meiosis; proposed to be involved in crossover interference | Member of the imitation-switch (ISWI) class of ATP-dependent chromatin remodeling complexes; ATPase that forms a complex with Ioc2p and Ioc4p to regulate transcription elongation, and a complex with Ioc3p to repress transcription initiation | Protein kinase of the bud neck involved in the septin checkpoint, associates with septin proteins, negatively regulates Swe1p by phosphorylation, shows structural homology to bud neck kinases Gin4p and Hsl1p | S-phase checkpoint protein found at replication forks, required for DNA replication; also required for Rad53p activation during DNA replication stress, where it forms a replication-pausing complex with Tof1p and is phosphorylated by Mec1p | Protein required for mismatch repair in mitosis and meiosis, forms a complex with Msh2p to repair both single-base & insertion-deletion mispairs; potentially phosphorylated by Cdc28p | GPI-anchored aspartyl protease (yapsin) involved in protein processing; shares functions with Yap3p and Kex2p | Ribonuclease H2 subunit, required for RNase H2 activity | Non-SMC subunit of the condensin complex (Smc2p-Smc4p-Ycs4p-Brn1p-Ycg1p); required for establishment and maintenance of chromosome condensation, chromosome segregation and for chromatin binding of the condensin complex | Conserved zinc-finger domain protein involved in pre-mRNA splicing, required for assembly of U4 snRNA into the U4/U6 particle | Subunit of the multiprotein cohesin complex required for sister chromatid cohesion in mitotic cells; also required, with Rec8p, for cohesion and recombination during meiosis; phylogenetically conserved SMC chromosomal ATPase family member | Mitogen-activated protein (MAP) kinase kinase kinase acting in the protein kinase C signaling pathway, which controls cell integrity; upon activation by Pkc1p phosphorylates downstream kinases Mkk1p and Mkk2p | Histone methyltransferase with a role in transcriptional elongation, methylates a lysine residue of histone H3; associates with the C-terminal domain of Rpo21p; histone methylation activity is regulated by phosphorylation status of Rpo21p | Putative protein of unknown function; expression is cell-cycle regulated as shown by microarray analysis | Third subunit of DNA polymerase delta, involved in chromosomal DNA replication; required for error-prone DNA synthesis in the presence of DNA damage and processivity; interacts with Hys2p, PCNA (Pol30p), and Pol1p | Protein involved in bud-site selection and required for axial budding pattern; localizes with septins to bud neck in mitosis and may constitute an axial landmark for next round of budding; potential Cdc28p substrate | Central plaque component of spindle pole body (SPB); involved in SPB duplication, may facilitate attachment of the SPB to the nuclear membrane | Subunit of DNA primase, which is required for DNA synthesis and double-strand break repair | Nim1p-related protein kinase that regulates the morphogenesis and septin checkpoints; associates with the assembled septin filament; required along with Hsl7p for bud neck recruitment, phosphorylation, and degradation of Swe1p | Fatty acid elongase, involved in sphingolipid biosynthesis; acts on fatty acids of up to 24 carbons in length; mutations have regulatory effects on 1,3-beta-glucan synthase, vacuolar ATPase, and the secretory pathway | Component of the polarisome, which functions in actin cytoskeletal organization during polarized growth; acts as a scaffold for Mkk1p and Mpk1p cell wall integrity signaling components; potential Cdc28p substrate | DNA replication initiation factor; recruited to MCM pre-RC complexes at replication origins; promotes release of MCM from Mcm10p, recruits elongation machinery; mutants in human homolog may cause velocardiofacial and DiGeorge syndromes | Protein involved in structural maintenance of chromosomes; essential subunit of Mms21-Smc5-Smc6 complex; required for growth, DNA repair, interchromosomal and sister chromatid recombination; homologous to S. pombe rad18 | Inositol hexakisphosphate (IP6) and inositol heptakisphosphate (IP7) kinase; generation of IP7 by Vip1p is important for phosphate signaling; likely involved in cortical actin cytoskeleton function, by analogy with S. pombe ortholog asp1 | Uracil-DNA glycosylase, required for repair of uracil in DNA formed by spontaneous cytosine deamination, not required for strand-specific mismatch repair, cell-cycle regulated, expressed in late G1, localizes to mitochondria and nucleus | Mitochondrial glycosylase/lyase that specifically excises 7,8-dihydro-8-oxoguanine residues located opposite cytosine or thymine residues in DNA, repairs oxidative damage to mitochondrial DNA | DNA helicase involved in telomere formation and elongation; acts as a catalytic inhibitor of telomerase; also plays a role in repair and recombination of mitochondrial DNA | Largest subunit of the origin recognition complex, which directs DNA replication by binding to replication origins and is also involved in transcriptional silencing; exhibits ATPase activity | Protein that interacts with silencing proteins at the telomere, involved in transcriptional silencing; paralog of Zds1p | Protein required for establishment and maintenance of sister chromatid condensation and cohesion, colocalizes with cohesin on chromosomes in an interdependent manner, may function as a protein-protein interaction scaffold | Subunit of a complex with Ctf8p that shares some subunits with Replication Factor C and is required for sister chromatid cohesion; may have overlapping functions with Rad24p in the DNA damage replication checkpoint | Protein required for normal transcription at several loci including HTA2-HTB2 and HHF2-HHT2, but not required at the other histone loci; functionally related to Spt10p; involved in telomere maintenance | ATP-binding protein required for mismatch repair in mitosis and meiosis; functions as a heterodimer with Mlh1p, binds double- and single-stranded DNA via its N-terminal domain, similar to E. coli MutL | Essential type II topoisomerase, relieves torsional strain in DNA by cleaving and re-sealing the phosphodiester backbone of both positively and negatively supercoiled DNA; cleaves complementary strands; localizes to axial cores in meiosis | Catalytic subunit of the DNA polymerase I alpha-primase complex, required for the initiation of DNA replication during mitotic DNA synthesis and premeiotic DNA synthesis | Component of the microtubule-nucleating Tub4p (gamma-tubulin) complex; interacts with Spc110p at the spindle pole body (SPB) inner plaque and with Spc72p at the SPB outer plaque | Protein involved in organization of septins at the mother-bud neck, may interact directly with the Cdc11p septin, localizes to bud neck in a septin-dependent manner | Subunit of heterotrimeric Replication Protein A (RPA), which is a highly conserved single-stranded DNA binding protein involved in DNA replication, repair, and recombination | Transcriptional co-repressor of MBF (MCB binding factor)-regulated gene expression; Nrm1p associates stably with promoters via MBF to repress transcription upon exit from G1 phase | Protein required for S phase progression and telomere homeostasis, forms an alternative replication factor C complex important for DNA replication and genome integrity; involved in homologous recombination-mediated DNA repair | Checkpoint protein, involved in the activation of the DNA damage and meiotic pachytene checkpoints; with Mec3p and Ddc1p, forms a clamp that is loaded onto partial duplex DNA; homolog of human and S. pombe Rad1 and U. maydis Rec1 proteins | Protein of unknown function, mutant phenotype suggests a potential role in vacuolar function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery, cytoplasm, bud, and bud neck | Protein required for the spindle pole body (SPB) duplication, localized at the central plaque periphery; forms a complex with a nuclear envelope protein Mps2p and SPB components Spc29p and Kar1p; required for mitotic functions of Cdc5p | Cell cycle regulated protein of unknown function; associated with Cdh1p and may supress the APC/C[Cdh1]-mediated proteolysis of mitotic cyclins | B-type cyclin involved in DNA replication during S phase; activates Cdc28p to promote initiation of DNA synthesis; functions in formation of mitotic spindles along with Clb3p and Clb4p; most abundant during late G1 phase | Transcription factor IIIA (TFIIIA), essential protein with nine C2H2 Zn-fingers, binds the 5S rRNA gene through the zinc finger domain and directs assembly of a multiprotein initiation complex for RNA polymerase III; also binds DNA | Protein kinase involved in bud growth and assembly of the septin ring, proposed to have kinase-dependent and kinase-independent activities; undergoes autophosphorylation; similar to Kcc4p and Hsl1p |
| 42 | 48 | 438 | 0.048 | 0.545 |  |  |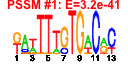  | 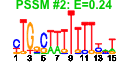 |  | YAL064W YAL066W YCL066W YCR040W YDL032W YDL071C YDL114W YDL162C YDL186W YDR095C YDR446W YDR523C YGL137W YGL158W YGL170C YGR273C YHR015W YHR105W YHR126C YHR185C YIL159W YJL043W YJL160C YJR078W YLR012C YLR308W YLR343W YNL204C YNL318C YOR255W YPR054W YAL018C YBR180W YBR250W YCL048W YDR042C YDR218C YDR402C YDR522C YER106W YFL040W YFR032C YGL015C YGL138C YGR107W YHR184W YLL030C YPR078C | YAL064W-B HMLALPHA1 ECM11 SPS1 SEC27 RCK1 SPO74 MIP6 YPT35 PFS1 BNR1 BNA2 CDA2 GAS2 SPS18 HXT14 OSW1 SMK1 DTR1 SPO23 YCL048W-A SPR28 DIT2 SPS2 MAM1 SSP1 | Fungal-specific protein of unknown function | Silenced copy of ALPHA1 at HML, encoding a transcriptional coactivator involved in the regulation of mating-type alpha-specific gene expression | Putative protein of unknown function with similarity to acyl-carrier-protein reductases; YDL114W is not an essential gene | Putative protein of unknown function; YDL186W is not an essential gene | Non-essential protein apparently involved in meiosis, GFP fusion protein is present in discrete clusters in the nucleus throughout mitosis; may be involved in maintaining chromatin structure | Putative protein serine/threonine kinase expressed at the end of meiosis and localized to the prospore membrane, required for correct localization of enzymes involved in spore wall synthesis | Essential beta'-coat protein of the COPI coatomer, involved in ER-to-Golgi and Golgi-to-ER transport; contains WD40 domains that mediate cargo selective interactions; 45% sequence identity to mammalian beta'-COP | Protein kinase involved in the response to oxidative stress; identified as suppressor of S. pombe cell cycle checkpoint mutations | Component of the meiotic outer plaque of the spindle pole body, involved in modifying the meiotic outer plaque that is required prior to prospore membrane formation | Putative protein of unknown function; deletion mutant has no readily detectable phenotype | Putative RNA-binding protein, interacts with Mex67p, which is a component of the nuclear pore involved in nuclear mRNA export | Endosomal protein of unknown function that contains a phox (PX) homology domain and binds to both phosphatidylinositol-3-phosphate (PtdIns(3)P) and proteins involved in ER-Golgi or vesicular transport | Putative protein of unknown function; transcription dependent upon Azf1p | Sporulation protein required for prospore membrane formation at selected spindle poles, ensures functionality of all four spindle pole bodies during meiosis II; not required for meiotic recombination or meiotic chromosome segregation | Formin, nucleates the formation of linear actin filaments, involved in cell processes such as budding and mitotic spindle orientation which require the formation of polarized actin cables, functionally redundant with BNI1 | Putative protein of unknown function; YJL043W is a non-essential gene | Putative protein of unknown function; member of the PIR (proteins with internal repeats) family of cell wall proteins; non-essential gene that is required for sporulation; mRNA is weakly cell cycle regulated, peaking in mitosis | Tryptophan 2,3-dioxygenase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Putative protein of unknown function; YLR012C is not an essential gene | Chitin deacetylase, together with Cda1p involved in the biosynthesis ascospore wall component, chitosan; required for proper rigidity of the ascospore wall | 1,3-beta-glucanosyltransferase, involved with Gas4p in spore wall assembly; has similarity to Gas1p | Protein of unknown function, contains a putative zinc-binding domain; expressed during sporulation | Protein with similarity to hexose transporter family members, expression is induced in low glucose and repressed in high glucose; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein involved in sporulation; required for the construction of the outer spore wall layers; required for proper localization of Spo14p | Middle sporulation-specific mitogen-activated protein kinase (MAPK) required for production of the outer spore wall layers; negatively regulates activity of the glucan synthase subunit Gsc2p | Putative protein of unknown function | Putative dityrosine transporter, required for spore wall synthesis; expressed during sporulation; member of the major facilitator superfamily (DHA1 family) of multidrug resistance transporters | Protein of unknown function; associates with meiosis-specific protein Spo1p | Putative protein of unknown function; expression is increased in ssu72-ts69 mutant | Sporulation-specific homolog of the yeast CDC3/10/11/12 family of bud neck microfilament genes; meiotic septin expressed at high levels during meiotic divisions and ascospore formation | N-formyltyrosine oxidase, sporulation-specific microsomal enzyme involved in the production of N,N-bisformyl dityrosine required for spore wall maturation, homologous to cytochrome P-450s | Protein expressed during sporulation, redundant with Sps22p for organization of the beta-glucan layer of the spore wall; S. pombe ortholog is a spore wall component | Monopolin, kinetochore associated protein involved in chromosome attachment to meiotic spindle | Putative transporter, member of the sugar porter family; YFL040W is not an essential gene | Putative protein of unknown function; transcribed during sporulation; YFR032C is not an essential gene | hypothetical protein | Putative protein of unknown function; has no significant sequence similarity to any known protein | Protein involved in the control of meiotic nuclear division and coordination of meiosis with spore formation; transcription is induced midway through meiosis | Putative protein of unknown function; possible role in DNA metabolism and/or in genome stability; expression is heat-inducible |
| 178 | 29 | 456 | 0.050 | 0.553 |  |  | 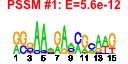 | 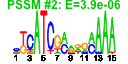 |  | YDL063C YDL131W YGL016W YJR071W YLL012W YLR084C YMR117C YMR301C YOL092W YBR061C YDL166C YDR280W YEL048C YHL013C YJL051W YJR072C YKL021C YLR397C YLR405W YMR080C YMR259C YNL120C YNL312W YNL313C YOL021C YOL141W YOR001W YOR246C YOR271C | LYS21 KAP122 YEH1 RAX2 SPC24 ATM1 TRM7 FAP7 RRP45 OTU2 IRC8 NPA3 MAK11 AFG2 DUS4 NAM7 RFA2 DIS3 PPM2 RRP6 FSF1 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YDL063C is not an essential gene | Homocitrate synthase isozyme, catalyzes the condensation of acetyl-CoA and alpha-ketoglutarate to form homocitrate, which is the first step in the lysine biosynthesis pathway; highly similar to the other isozyme, Lys20p | Karyopherin beta, responsible for import of the Toa1p-Toa2p complex into the nucleus; binds to nucleoporins Nup1p and Nup2p; may play a role in regulation of pleiotropic drug resistance | Steryl ester hydrolase, one of three gene products (Yeh1p, Yeh2p, Tgl1p) responsible for steryl ester hydrolase activity and involved in sterol homeostasis; localized to lipid particle membranes | N-glycosylated protein involved in the maintenance of bud site selection during bipolar budding; localization requires Rax1p | Component of the evolutionarily conserved kinetochore-associated Ndc80 complex (Ndc80p-Nuf2p-Spc24p-Spc25p); involved in chromosome segregation, spindle checkpoint activity and kinetochore clustering | Mitochondrial inner membrane ATP-binding cassette (ABC) transporter, exports mitochondrially synthesized precursors of iron-sulfur (Fe/S) clusters to the cytosol | Putative protein of unknown function; predicted to contain six transmembrane domains and is 58% similar to the uncharacterized ORF YBR147W; deletion mutant has no detectable phenotype | 2'-O-ribose methyltransferase, methylates the 2'-O-ribose of nucleotides at positions 32 and 34 of the tRNA anticodon loop | Essential NTPase required for small ribosome subunit synthesis, mediates processing of the 20S pre-rRNA at site D in the cytoplasm but associates only transiently with 43S preribosomes via Rps14p, may be the endonuclease for site D | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex | Putative protein of unknown function; synthetic lethal with gcs1delta | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; member of the ovarian tumor-like (OTU) superfamily of predicted cysteine proteases; shows cytoplasmic localization | Bud tip localized protein of unknown function; mRNA is targeted to the bud by a She2p dependent transport system; mRNA is cell cycle regulated via Fkh2p, peaking in G2/M phase; null mutant displays increased levels of spontaneous Rad52 foci | Essential, conserved, cytoplasmic ATPase; phosphorylated by the Pcl1p-Pho85p kinase complex | Protein involved in an early, nucleolar step of 60S ribosomal subunit biogenesis; essential for cell growth and replication of killer M1 dsRNA virus; contains four beta-transducin repeats | ATPase of the CDC48/PAS1/SEC18 (AAA) family, forms a hexameric complex; may be involved in degradation of aberrant mRNAs | Dihydrouridine synthase, member of a widespread family of conserved proteins including Smm1p, Dus1p, and Dus3p | ATP-dependent RNA helicase of the SFI superfamily, required for nonsense mediated mRNA decay and for efficient translation termination at nonsense codons; involved in telomere maintenance | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YMR259C is not an essential gene | Subunit of heterotrimeric Replication Protein A (RPA), which is a highly conserved single-stranded DNA binding protein involved in DNA replication, repair, and recombination | Putative protein of unknown function; identified as interacting with Kar2p, Grs1p, and Tub3p in high-throughput TAP-tagging experiments; YNL313C is an essential gene | Catalytic component of the exosome, involved in RNA processing and degradation; binds Gsp1p/Ran and enhances the GEF activity of Srm1p; implicated in mitotic control; homologous to the E. coli RNase R of the RNase II family | Putative carboxymethyltransferase, similar to Ppm1p but biochemical activity not yet demonstrated; required for the synthesis of wybutosine, a modified guanosine found at the 3'-position adjacent to the anticodon of phe-tRNA | Exonuclease component of the nuclear exosome; contributes to the quality-control system that retains and degrades aberrant mRNAs in the nucleus | Protein with similarity to oxidoreductases, found in lipid particles; required for replication of Brome mosaic virus in S. cerevisiae, which is a model system for studying replication of positive-strand RNA viruses in their natural hosts | Putative protein, predicted to be an alpha-isopropylmalate carrier; belongs to the sideroblastic-associated protein family; non-tagged protein is detected in purified mitochondria; likely to play a role in iron homeostasis |
| 132 | 44 | 440 | 0.052 | 0.516 |  |  | 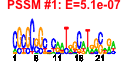 |  |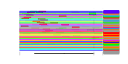  | YBR244W YCR102C YCR107W YCRX21C YDL168W YDL243C YDR132C YDR453C YER130C YFL056C YFL057C YGR007W YGR011W YHR008C YHR106W YHR159W YHR199C YIR018W YJR104C YKL071W YKL086W YKR041W YLL060C YLR460C YML116W YMR023C YMR038C YMR061W YNL260C YNL333W YNL334C YOL165C YOR114W YOR225W YOR226C YBR008C YBR047W YDR353W YIL164C YJL101C YMR027W YMR042W YNL331C YPL171C | GPX2 AAD3 ADH5 AAD4 TSA2 AAD6 AAD16 MUQ1 SOD2 TRR2 AIM46 YAP5 SOD1 SRX1 GTT2 ATR1 MSS1 CCS1 RNA14 SNZ2 SNO2 AAD15 ISU2 FLR1 FMP23 TRR1 NIT1 GSH1 ARG80 AAD14 OYE3 | Phospholipid hydroperoxide glutathione peroxidase induced by glucose starvation that protects cells from phospholipid hydroperoxides and nonphospholipid peroxides during oxidative stress | Putative protein of unknown function; involved in copper metabolism; similar to C.carbonum toxD gene; YCR102C is not an essential gene | Putative aryl-alcohol dehydrogenase with similarity to P. chrysosporium aryl-alcohol dehydrogenase; mutational analysis has not yet revealed a physiological role | Alcohol dehydrogenase isoenzyme V; involved in ethanol production | Putative aryl-alcohol dehydrogenase with similarity to P. chrysosporium aryl-alcohol dehydrogenase, involved in the oxidative stress response | Putative protein of unknown function | Stress inducible cytoplasmic thioredoxin peroxidase; cooperates with Tsa1p in the removal of reactive oxygen, nitrogen and sulfur species using thioredoxin as hydrogen donor; deletion enhances the mutator phenotype of tsa1 mutants | hypothetical protein | Choline phosphate cytidylyltransferase, catalyzes the second step of phosphatidylethanolamine biosynthesis; involved in the maintenance of plasma membrane; similar to mammalian CTP: phosphocholine cytidylyl-transferases | Mitochondrial superoxide dismutase, protects cells against oxygen toxicity; phosphorylated | Mitochondrial thioredoxin reductase involved in protection against oxidative stress, required with Glr1p to maintain the redox state of Trx3p; contains active-site motif (CAVC) present in prokaryotic orthologs; binds NADPH and FAD | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; potential Cdc28p substrate | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Basic leucine zipper (bZIP) transcription factor | Cytosolic superoxide dismutase; some mutations are analogous to those that cause ALS (amyotrophic lateral sclerosis) in humans | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Sulfiredoxin, contributes to oxidative stress resistance by reducing cysteine-sulfinic acid groups in the peroxiredoxins Tsa1p and Ahp1p that are formed upon exposure to oxidants; conserved in higher eukaryotes | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Glutathione S-transferase capable of homodimerization; functional overlap with Gtt2p, Grx1p, and Grx2p | Putative protein of unknown function, possibly up-regulated by iodine | Multidrug efflux pump of the major facilitator superfamily, required for resistance to aminotriazole and 4-nitroquinoline-N-oxide | Mitochondrial protein, forms a heterodimer complex with Mto1p that performs the 5-carboxymethylaminomethyl modification of the wobble uridine base in mitochondrial tRNAs; similar to human GTPBP3 | Copper chaperone for superoxide dismutase Sod1p, involved in oxidative stress protection; Met-X-Cys-X2-Cys motif within the N-terminal portion is involved in insertion of copper into Sod1p under conditions of copper deprivation | Cleavage and polyadenylation factor I (CF I) component involved in cleavage and polyadenylation of mRNA 3' ends; bridges interaction between Rna15p and Hrp1p in the CF I complex | Putative protein of unknown function with similarity to a human protein overexpressed in oral cancers; localizes to the nucleus and cytoplasm; YNL260C is an essential gene | Member of a stationary phase-induced gene family; transcription of SNZ2 is induced prior to diauxic shift, and also in the absence of thiamin in a Thi2p-dependent manner; forms a coregulated gene pair with SNO2; interacts with Thi11p | Protein of unknown function, nearly identical to Sno3p; expression is induced before the diauxic shift and also in the absence of thiamin | Conserved protein of the mitochondrial matrix, required for synthesis of mitochondrial and cytosolic iron-sulfur proteins, performs a scaffolding function in mitochondria during Fe/S cluster assembly; isu1 isu2 double mutant is inviable | Plasma membrane multidrug transporter of the major facilitator superfamily, involved in efflux of fluconazole, diazaborine, benomyl, methotrexate, and other drugs | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Cytoplasmic thioredoxin reductase, key regulatory enzyme that determines the redox state of the thioredoxin system, which acts as a disulfide reductase system and protects cells against both oxidative and reductive stress | Nitrilase, member of the nitrilase branch of the nitrilase superfamily; in closely related species and other S. cerevisiae strain backgrounds YIL164C and adjacent ORF, YIL165C, likely constitute a single ORF encoding a nitrilase gene | Gamma glutamylcysteine synthetase catalyzes the first step in glutathione (GSH) biosynthesis; expression induced by oxidants, cadmium, and mercury | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and cytoplasm; YMR027W is not an essential gene | Transcription factor involved in regulation of arginine-responsive genes; acts with Arg81p and Arg82p | Widely conserved NADPH oxidoreductase containing flavin mononucleotide (FMN), homologous to Oye2p with slight differences in ligand binding and catalytic properties; may be involved in sterol metabolism |
| 245 | 29 | 447 | 0.054 | 0.540 |  |  |  |  |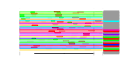  | YIL099W YKL121W YOL047C YBL086C YBR001C YBR059C YBR223C YBR225W YBR241C YBR284W YCR061W YCR062W YDL091C YDL233W YGL036W YHR137W YHR202W YKL036C YLR299W YMR252C YNR034W YOL073C YOL096C YOR028C YOR049C YOR055W YOR273C YPL164C YPR160W | SGA1 NTH2 AKL1 TDP1 UBX3 ARO9 ECM38 SOL1 COQ3 CIN5 RSB1 TPO4 MLH3 GPH1 | Intracellular sporulation-specific glucoamylase involved in glycogen degradation; induced during starvation of a/a diploids late in sporulation, but dispensable for sporulation | Putative protein of unknown function | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | Putative neutral trehalase, required for thermotolerance and may mediate resistance to other cellular stresses | Ser-Thr protein kinase, member (with Ark1p and Prk1p) of the Ark kinase family; involved in endocytosis and actin cytoskeleton organization | Tyrosyl-DNA Phosphodiesterase I, hydrolyzes 3'-phosphotyrosyl bonds to generate 3'-phosphate DNA and tyrosine, involved in the repair of DNA lesions created by topoisomerase I | Putative protein of unknown function; non-essential gene identified in a screen for mutants affected in mannosylphophorylation of cell wall components | Putative transporter, member of the sugar porter family; green fluorescent protein (GFP)-fusion protein localizes to the vacuolar membrane; YBR241C is not an essential gene | Putative protein of unknown function; YBR284W is not an essential gene; null mutant exhibits decreased resistance to rapamycin and wortmannin | UBX (ubiquitin regulatory X) domain-containing protein that interacts with Cdc48p, green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YDL233W is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YGL036W is not an essential gene | Aromatic aminotransferase II, catalyzes the first step of tryptophan, phenylalanine, and tyrosine catabolism | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole, while HA-tagged protein is found in the soluble fraction, suggesting cytoplasmic localization | Gamma-glutamyltranspeptidase, major glutathione-degrading enzyme; involved in detoxification of electrophilic xenobiotics; expression induced mainly by nitrogen starvation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YMR252C is not an essential gene | Protein with a possible role in tRNA export; shows similarity to 6-phosphogluconolactonase non-catalytic domains but does not exhibit this enzymatic activity; homologous to Sol2p, Sol3p, and Sol4p | O-methyltransferase, catalyzes two different O-methylation steps in ubiquinone (Coenzyme Q) biosynthesis; component of a mitochondrial ubiquinone-synthesizing complex | Basic leucine zipper transcriptional factor of the yAP-1 family that mediates pleiotropic drug resistance and salt tolerance; localizes constitutively to the nucleus | Suppressor of sphingoid long chain base (LCB) sensitivity of an LCB-lyase mutation; putative integral membrane transporter or flippase that may transport LCBs from the cytoplasmic side toward the extracytoplasmic side of the membrane | Polyamine transport protein, recognizes spermine, putrescine, and spermidine; localizes to the plasma membrane; member of the major facilitator superfamily | Protein involved in DNA mismatch repair and crossing-over during meiotic recombination; forms a complex with Mlh1p; mammalian homolog is implicated mammalian microsatellite instability | Non-essential glycogen phosphorylase required for the mobilization of glycogen, activity is regulated by cyclic AMP-mediated phosphorylation, expression is regulated by stress-response elements and by the HOG MAP kinase pathway |
| 158 | 29 | 446 | 0.063 | 0.566 | 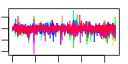 |  |  | 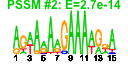 |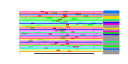  | YBR030W YBR141C YBR155W YDR527W YFL023W YGR093W YGR251W YGR283C YHR069C YHR070W YLR014C YMR185W YMR259C YNL120C YNL299W YNR015W YOL141W YBL018C YDL121C YDR020C YER002W YHR081W YJL125C YJR097W YKL110C YKR025W YLR107W YNL227C YOR091W | CNS1 RBA50 BUD27 DRN1 RRP4 TRM5 PPR1 TRF5 RSP5 PPM2 POP8 DAS2 NOP16 LRP1 GCD14 JJJ3 KTI12 RPC37 REX3 JJJ1 TMA46 | Putative protein of unknown function; predicted protein contains a SET domain (S-adenosyl-L-methionine-binding fold) | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus; YBR141C is not an essential gene | TPR-containing co-chaperone; binds both Hsp82p (Hsp90) and Ssa1p (Hsp70) and stimulates the ATPase activity of SSA1, ts mutants reduce Hsp82p function while over expression suppresses the phenotypes of an HSP82 ts allele and a cpr7 deletion | Protein involved in transcription; interacts with RNA polymerase II subunits Rpb2p, Rpb3, and Rpb11p; has similarity to human RPAP1 | Protein involved in bud-site selection, nutrient signaling, and gene expression controlled by the TOR kinase; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Putative debranching enzyme associated ribonuclease; green fluorescent protein (GFP)-fusion protein localizes to the nucleus | Putative protein of unknown function; deletion mutant has defects in pre-rRNA processing; green fluorescent protein (GFP)-fusion protein localizes to both the nucleus and the nucleolus; YGR251W is an essential gene | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; deletion mutant is resistant to fluconazole; green fluorescent protein (GFP)-fusion protein localizes to the nucleolus | 3'-5' exoribonuclease involved in rRNA processing; component of the exosome 3->5 exonuclease complex with Rrp41p, Rrp42p, Rrp43p and Dis3p | tRNA(m(1)G37)methyltransferase, methylates a tRNA base adjacent to the anticodon that has a role in prevention of frameshifting; highly conserved across Archaea, Bacteria, and Eukarya | Zinc finger transcription factor containing a Zn(2)-Cys(6) binuclear cluster domain, positively regulates transcription of genes involved in uracil biosynthesis; activity may be modulated by interaction with Tup1p | Putative protein of unknown function; essential gene required for viability | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YMR259C is not an essential gene | Poly (A) polymerase involved in nuclear RNA quality control based on: homology with Trf4p, genetic interactions with TRF4 mutants, physical interaction with Mtr4p (TRAMP subunit), and by direct assay; disputed role as a sigma DNA polymerase | Ubiquitin-protein ligase involved in ubiquitin-mediated protein degradation; functions in multivesicular body sorting, heat shock response and ubiquitylation of arrested RNAPII; contains a hect (homologous to E6-AP carboxyl terminus) domain | Putative carboxymethyltransferase, similar to Ppm1p but biochemical activity not yet demonstrated; required for the synthesis of wybutosine, a modified guanosine found at the 3'-position adjacent to the anticodon of phe-tRNA | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic retiuculum; YDL121C is not an essential protein | Predicted protein shares weak similarity with uridine kinases and with phosphoribokinases; null mutant suppresses dst1delta sensitivity for 6-azauracil | Constituent of 66S pre-ribosomal particles, involved in 60S ribosomal subunit biogenesis | Substrate-specific nuclear cofactor for exosome activity in the processing of stable RNAs; required for telomere length maintenance; homolog of mammalian nuclear matrix protein C1D involved in regulation of DNA repair and recombination | Subunit of tRNA (1-methyladenosine) methyltransferase, with Gcd10p, required for the modification of the adenine at position 58 in tRNAs, especially tRNAi-Met; first identified as a negative regulator of GCN4 expression | Protein of unknown function, contains a J-domain, which is a region with homology to the E. coli DnaJ protein | Protein that plays a role, with Elongator complex, in modification of wobble nucleosides in tRNA; involved in sensitivity to G1 arrest induced by zymocin; interacts with chromatin throughout the genome; also interacts with Cdc19p | RNA polymerase III subunit C37 | RNA exonuclease; required for maturation of the RNA component of RNase MRP; functions redundantly with Rnh70p and Rex2p in processing of U5 snRNA and RNase P RNA; member of RNase D family of exonucleases | Co-chaperone that stimulates the ATPase activity of Ssa1p, required for a late step of ribosome biogenesis; associated with the cytosolic large ribosomal subunit; contains a J-domain; mutation causes defects in fluid-phase endocytosis | Protein of unknown function that associates with ribosomes; interacts with GTPase Rbg1p |
| 48 | 33 | 442 | 0.068 | 0.522 | 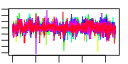 |  |  | 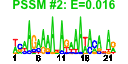 |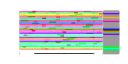  | YBR018C YDR018C YDR043C YDR069C YDR077W YDR203W YDR273W YDR282C YDR330W YDR350C YDR426C YEL020C YEL041W YER038C YER096W YER098W YGL045W YGL208W YGR042W YGR052W YHL044W YIL077C YMR141C YMR159C YMR253C YMR284W YOL100W YOR052C YPL060W YPR005C YDR474C YDR475C YFL044C | GAL7 NRG1 DOA4 SED1 DON1 UBX5 ATP22 YEF1 KRE29 SHC1 UBP9 RIM8 SIP2 FMP48 ATG16 YKU70 PKH2 LPE10 HAL1 JIP4 OTU1 | Galactose-1-phosphate uridyl transferase, synthesizes glucose-1-phosphate and UDP-galactose from UDP-D-glucose and alpha-D-galactose-1-phosphate in the second step of galactose catabolism | Probable membrane protein with three predicted transmembrane domains; homologous to Ybr042cp, similar to C. elegans F55A11.5 and maize 1-acyl-glycerol-3-phosphate acyltransferase; null exhibits no apparent phenotype | Transcriptional repressor that recruits the Cyc8p-Tup1p complex to promoters; mediates glucose repression and negatively regulates a variety of processes including filamentous growth and alkaline pH response | Ubiquitin isopeptidase, required for recycling ubiquitin from proteasome-bound ubiquitinated intermediates, acts at the late endosome/prevacuolar compartment to recover ubiquitin from ubiquitinated membrane proteins en route to the vacuole | Major stress-induced structural GPI-cell wall glycoprotein in stationary-phase cells, associates with translating ribosomes, possible role in mitochondrial genome maintenance; ORF contains two distinct variable minisatellites | Meiosis-specific component of the spindle pole body, part of the leading edge protein (LEP) coat, forms a ring-like structure at the leading edge of the prospore membrane during meiosis II | Putative protein of unknown function | UBX (ubiquitin regulatory X) domain-containing protein that interacts with Cdc48p | Mitochondrial inner membrane protein required for assembly of the F0 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | hypothetical protein with low sequence identity to Pdc1p | ATP-NADH kinase; phosophorylates both NAD and NADH; homooctameric structure consisting of 60-kDa subunits; sequence similarity to Utr1p and Pos5p; overexpression complements certain pos5 phenotypes | Essential subunit of the Mms21-Smc5-Smc6 complex; protein of unknown function; required for growth and DNA repair; heterozygous mutant shows haploinsufficiency in K1 killer toxin resistance | Sporulation-specific activator of Chs3p (chitin synthase III), required for the synthesis of the chitosan layer of ascospores; has similarity to Skt5p, which activates Chs3p during vegetative growth; transcriptionally induced at alkaline pH | Ubiquitin carboxyl-terminal hydrolase, ubiquitin-specific protease that cleaves ubiquitin-protein fusions | Protein of unknown function, involved in the proteolytic activation of Rim101p in response to alkaline pH; has similarity to A. nidulans PalF | One of three beta subunits of the Snf1 serine/threonine protein kinase complex involved in the response to glucose starvation; null mutants exhibit accelerated aging; N-myristoylprotein localized to the cytoplasm and the plasma membrane | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative integral membrane protein, member of DUP240 gene family; green fluorescent protein (GFP)-fusion protein localizes to the plasma membrane in a punctate pattern | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO) | Protein that interacts with the Atg12p-Atg5p conjugate during formation of the pre-autophagosomal structure; essential for autophagy | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern; YMR253C is not an essential gene | Subunit of the telomeric Ku complex (Yku70p-Yku80p), involved in telomere length maintenance, structure and telomere position effect; relocates to sites of double-strand cleavage to promote nonhomologous end joining during DSB repair | Serine/threonine protein kinase involved in sphingolipid-mediated signaling pathway that controls endocytosis; activates Ypk1p and Ykr2p, components of signaling cascade required for maintenance of cell wall integrity; redundant with Pkh1p | Nuclear protein of unknown function; expression induced by nitrogen limitation in a GLN3, GAT1-independent manner and by weak acid | Mitochondrial inner membrane magnesium transporter, involved in maintenance of magnesium concentrations inside mitochondria; indirectly affects splicing of group II introns; functionally and structurally related to Mrs2p | Cytoplasmic protein involved in halotolerance; decreases intracellular Na+ (via Ena1p) and increases intracellular K+ by decreasing efflux; expression repressed by Ssn6p-Tup1p and Sko1p and induced by NaCl, KCl, and sorbitol through Gcn4p | Protein of unknown function; previously annotated as two separate ORFs, YDR474C and YDR475C, which were merged as a result of corrections to the systematic reference sequence | Deubiquitylation enzyme that binds to the chaperone-ATPase Cdc48p; may contribute to regulation of protein degradation by deubiquitylating substrates that have been ubiquitylated by Ufd2p; member of the Ovarian Tumor (OTU) family |
| 108 | 33 | 441 | 0.071 | 0.495 |  |  | 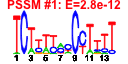 | 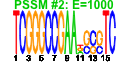 |  | YBR222C YDL238C YDR085C YFR045W YGL180W YGL196W YGR053C YIL055C YIL166C YIR005W YKL200C YKL201C YLL057C YLR030W YLR031W YLR282C YOL089C YOL163W YDR223W YDR273W YEL020C YGL004C YGL124C YGR042W YGR045C YGR213C YHR006W YIL029C YKL095W YLR165C YOL162W YPL222W YPR001W | PCS60 GUD1 AFR1 ATG1 DSD1 IST3 MNN4 JLP1 HAL9 CRF1 DON1 RPN14 MON1 RTA1 STP2 YJU2 PUS5 FMP40 CIT3 | Peroxisomal AMP-binding protein, localizes to both the peroxisomal peripheral membrane and matrix, expression is highly inducible by oleic acid, similar to E. coli long chain acyl-CoA synthetase | Guanine deaminase, a catabolic enzyme of the guanine salvage pathway producing xanthine and ammonia from guanine; activity is low in exponentially-growing cultures but expression is increased in post-diauxic and stationary-phase cultures | Alpha-factor pheromone receptor regulator, negatively regulates pheromone receptor signaling; required for normal mating projection (shmoo) formation; required for Spa2p to recruit Mpk1p to shmoo tip during mating; interacts with Cdc12p | Putative mitochondrial transport protein; null mutant is viable, exhibits decreased levels of chitin and normal resistance to calcofluor white | Protein serine/threonine kinase, required for vesicle formation during autophagy and the cytoplasm-to-vacuole targeting (Cvt) pathway; structurally required for pre-autophagosome formation; forms a complex with Atg13p and Atg17p | D-serine dehydratase (aka D-serine ammonia-lyase); converts D-serine to pyruvate and ammonia; specifc for D-serine, unlike the bacterial enzyme which recognizes both D-serine and L-serine as substrates | Putative protein of unknown function | Putative protein with similarity to the allantoate permease (Dal5p) subfamily of the major facilitator superfamily; mRNA expression is elevated by sulfur limitation; YIL166C is a non-essential gene | Component of the U2 snRNP, required for the first catalytic step of splicing and for spliceosomal assembly; interacts with Rds3p and is required for Mer1p-activated splicing | Putative positive regulator of mannosylphosphate transferase (Mnn6p), involved in mannosylphosphorylation of N-linked oligosaccharides; expression increases in late-logarithmic and stationary growth phases | Fe(II)-dependent sulfonate/alpha-ketoglutarate dioxygenase, involved in sulfonate catabolism for use as a sulfur source, contains sequence that closely resembles a J domain (typified by the E. coli DnaJ protein) | Putative transcription factor containing a zinc finger; overexpression increases salt tolerance through increased expression of the ENA1 (Na+/Li+ extrusion pump) gene while gene disruption decreases both salt tolerance and ENA1 expression | Putative protein of unknown function; member of the Dal5p subfamily of the major facilitator family | Transcriptional corepressor involved in the regulation of ribosomal protein gene transcription via the TOR signaling pathway and protein kinase A, phosphorylated by activated Yak1p which promotes accumulation of Crf1p in the nucleus | Meiosis-specific component of the spindle pole body, part of the leading edge protein (LEP) coat, forms a ring-like structure at the leading edge of the prospore membrane during meiosis II | hypothetical protein with low sequence identity to Pdc1p | Putative non-ATPase subunit of the 19S regulatory particle of the 26S proteasome; localized to the cytoplasm | Protein required for fusion of cvt-vesicles and autophagosomes with the vacuole; associates, as a complex with Ccz1p, with a perivacuolar compartment; potential Cdc28p substrate | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Protein involved in 7-aminocholesterol resistance; has seven potential membrane-spanning regions | Transcription factor, activated by proteolytic processing in response to signals from the SPS sensor system for external amino acids; activates transcription of amino acid permease genes | Putative protein of unknown function; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO) | Essential protein required for pre-mRNA splicing; associates transiently with the spliceosomal NTC ('nineteen complex') and acts after Prp2p to promote the first catalytic reaction of splicing | Pseudouridine synthase, catalyzes only the formation of pseudouridine (Psi)-2819 in mitochondrial 21S rRNA; not essential for viability | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Dual specificity mitochondrial citrate and methylcitrate synthase; catalyzes the condensation of acetyl-CoA and oxaloacetate to form citrate and that of propionyl-CoA and oxaloacetate to form 2-methylcitrate |
| 107 | 27 | 438 | 0.075 | 0.516 |  |  |  |  |  | YBR280C YDR255C YER015W YFL042C YFL043C YGL227W YGR201C YIR014W YIR017C YNL092W YOL117W YPL177C YPR061C YDR043C YDR254W YDR306C YER020W YIL017W YJL142C YKL133C YMR053C YMR085W YMR114C YMR135C YMR141C YMR174C YOR054C | SAF1 RMD5 FAA2 VID30 MET28 RRI2 CUP9 JID1 NRG1 CHL4 GPA2 VID28 STB2 GID8 PAI3 VHS3 | F-Box protein involved in proteasome-dependent degradation of Aah1p during entry of cells into quiescence; interacts with Skp1 | Cytosolic protein required for sporulation; also required for the ubiquitination of the gluconeogenetic enzyme fructose-1,6-bisphosphatase, which is degraded rapidly after the switch from gluconeogenesis to glycolysis | Long chain fatty acyl-CoA synthetase; accepts a wider range of acyl chain lengths than Faa1p, preferring C9:0-C13:0; involved in the activation of endogenous pools of fatty acids | Putative protein of unknown function; YFL042C is not an essential gene | Protein involved in proteasome-dependent catabolite degradation of fructose-1,6-bisphosphatase (FBPase); shifts the balance of nitrogen metabolism toward the production of glutamate; localized to the nucleus and the cytoplasm | Putative protein of unknown function | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; expression directly regulated by the metabolic and meiotic transcriptional regulator Ume6p; YIR014W is a non-essential gene | Transcriptional activator in the Cbf1p-Met4p-Met28p complex, participates in the regulation of sulfur metabolism | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; YNL092W is not an essential gene | Subunit of the COP9 signalosome (CSN) complex that cleaves the ubiquitin-like protein Nedd8 from SCF ubiquitin ligases; plays a role in the mating pheromone response | Homeodomain-containing transcriptional repressor of PTR2, which encodes a major peptide transporter; imported peptides activate ubiquitin-dependent proteolysis, resulting in degradation of Cup9p and de-repression of PTR2 transcription | Probable Hsp40p co-chaperone, has a DnaJ-like domain and appears to be involved in ER-associated degradation of misfolded proteins containing a tightly folded cytoplasmic domain; inhibits replication of Brome mosaic virus in S. cerevisiae | Transcriptional repressor that recruits the Cyc8p-Tup1p complex to promoters; mediates glucose repression and negatively regulates a variety of processes including filamentous growth and alkaline pH response | Outer kinetochore protein required for chromosome stability, interacts with kinetochore proteins Ctf19p, Ctf3p, and Iml3p; exhibits a two-hybrid interaction with Mif2p; association with CEN DNA requires Ctf19p | F-box protein of unknown function; interacts with Sgt1p via a Leucine-Rich Repeat (LRR) domain | Nucleotide binding alpha subunit of the heterotrimeric G protein that interacts with the receptor Gpr1p, has signaling role in response to nutrients; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | Protein involved in proteasome-dependent catabolite degradation of fructose-1,6-bisphosphatase (FBPase); localized to the nucleus and the cytoplasm | Protein that interacts with Sin3p in a two-hybrid assay and is part of a large protein complex with Sin3p and Stb1p | Putative protein of unknown function; YMR085W and adjacent ORF YMR084W are merged in related strains | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and cytoplasm; YMR114C is not an essential gene | Protein of unknown function, involved in proteasome-dependent catabolite inactivation of fructose-1,6-bisphosphatase; contains LisH and CTLH domains, like Vid30p; dosage-dependent regulator of START | Cytoplasmic proteinase A (Pep4p) inhibitor, dependent on Pbs2p and Hog1p protein kinases for osmotic induction; intrinsically unstructured, N-terminal half becomes ordered in the active site of proteinase A upon contact | Functionally redundant (see also SIS2) inhibitory subunit of Ppz1p, a PP1c-related ser/thr protein phosphatase Z isoform; synthetically lethal with sis2; putative phosphopantothenoylcysteine decarboxylase involved in coenzyme A biosynthesis |
| 118 | 35 | 433 | 0.090 | 0.555 |  |  |  |  |  | YAL015C YDR032C YDR294C YER178W YKL035W YKL039W YKL192C YKR066C YLR337W YML028W YML057W YMR261C YMR262W YNL055C YNL071W YNL241C YOL122C YOL143C YPL091W YPL206C YPL260W YCL044C YCR004C YDL086W YEL047C YER004W YJR044C YML110C YML131W YNL045W YNL239W YOR136W YOR142W YPL154C YPL262W | NTG1 PST2 DPL1 PDA1 UGP1 PTM1 ACP1 CCP1 VRP1 TSA1 CMP2 TPS3 POR1 LAT1 ZWF1 SMF1 RIB4 GLR1 MGR1 YCP4 FMP52 VPS55 COQ5 LAP3 IDH2 LSC1 PEP4 FUM1 | DNA N-glycosylase and apurinic/apyrimidinic (AP) lyase involved in base excision repair, localizes to the nucleus and mitochondrion | Protein with similarity to members of a family of flavodoxin-like proteins; induced by oxidative stress in a Yap1p dependent manner; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Dihydrosphingosine phosphate lyase, regulates intracellular levels of sphingolipid long-chain base phosphates (LCBPs), degrades phosphorylated long chain bases, prefers C16 dihydrosphingosine-l-phosphate as a substrate | E1 alpha subunit of the pyruvate dehydrogenase (PDH) complex, catalyzes the direct oxidative decarboxylation of pyruvate to acetyl-CoA; phosphorylated; regulated by glucose | UDP-glucose pyrophosphorylase (UGPase), catalyses the reversible formation of UDP-Glc from glucose 1-phosphate and UTP, involved in a wide variety of metabolic pathways, expression modulated by Pho85p through Pho4p | Protein of unknown function, copurifies with late Golgi vesicles containing the v-SNARE Tlg2p | Mitochondrial matrix acyl carrier protein, involved in biosynthesis of octanoate, which is a precursor to lipoic acid; activated by phosphopantetheinylation catalyzed by Ppt2p | Mitochondrial cytochrome-c peroxidase; degrades reactive oxygen species in mitochondria, involved in the response to oxidative stress | Proline-rich actin-associated protein involved in cytoskeletal organization and cytokinesis; related to mammalian Wiskott-Aldrich syndrome protein (WASP)-interacting protein (WIP) | Ubiquitous housekeeping thioredoxin peroxidase, reduces reactive oxygen, nitrogen and sulfur species using thioredoxin as hydrogen donor; mediates redox regulation of the nuclear localization of Yap1p; deletion results in mutator phenotype | Calcineurin A; one isoform (the other is CNA1) of the catalytic subunit of calcineurin, a Ca++/calmodulin-regulated protein phosphatase which regulates Crz1p (a stress-response transcription factor), the other calcineurin subunit is CNB1 | Regulatory subunit of trehalose-6-phosphate synthase/phosphatase complex, which synthesizes the storage carbohydrate trehalose; expression is induced by stress conditions and repressed by the Ras-cAMP pathway | Protein of unknown function; interacts weakly with Knr4p; YMR262W is not an essential gene | Mitochondrial porin (voltage-dependent anion channel), outer membrane protein required for the maintenance of mitochondrial osmotic stability and mitochondrial membrane permeability; phosphorylated | Dihydrolipoamide acetyltransferase component (E2) of pyruvate dehydrogenase complex, which catalyzes the oxidative decarboxylation of pyruvate to acetyl-CoA | Glucose-6-phosphate dehydrogenase (G6PD), catalyzes the first step of the pentose phosphate pathway; involved in adapting to oxidatve stress; homolog of the human G6PD which is deficient in patients with hemolytic anemia | Divalent metal ion transporter with a broad specificity for di-valent and tri-valent metals; post-translationally regulated by levels of metal ions; member of the Nramp family of metal transport proteins | Lumazine synthase (6,7-dimethyl-8-ribityllumazine synthase, also known as DMRL synthase); catalyzes synthesis of immediate precursor to riboflavin | Cytosolic and mitochondrial glutathione oxidoreductase, converts oxidized glutathione to reduced glutathione | Protein that plays a role in metabolism of glycerophosphocholine to phosphatidylcholine; contains glycerophosphodiester phosphodiesterase motifs | Putative substrate of cAMP-dependent protein kinase (PKA); green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YPL260W is not an essential gene | Subunit, with Yme1p, of the mitochondrial inner membrane i-AAA protease complex, which is responsible for degradation of unfolded or misfolded mitochondrial gene products; required for growth of cells lacking the mitochondrial genome | Protein of unknown function, has sequence and structural similarity to flavodoxins; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YDL086W is not an essential gene | Soluble fumarate reductase, required with isoenzyme Osm1p for anaerobic growth; may interact with ribosomes, based on co-purification experiments; authentic, non-tagged protein is detected in purified mitochondria in high-throughput studies | Protein of unknown function, localized to the mitochondrial outer membrane | Late endosomal protein involved in late endosome to vacuole trafficking; functional homolog of human obesity receptor gene-related protein (OB-RGRP) | 2-hexaprenyl-6-methoxy-1,4-benzoquinone methyltransferase, involved in ubiquinone (Coenzyme Q) biosynthesis; localizes to the matrix face of the mitochondrial inner membrane in a large complex with other ubiquinone biosynthetic enzymes | Putative protein of unknown function with similarity to oxidoreductases; HOG1 and SKO1-dependent mRNA expression is induced after osmotic shock; GFP-fusion protein localizes to the cytoplasm and is induced by the DNA-damaging agent MMS | Leucyl aminopeptidase (leukotriene A4 hydrolase) with epoxide hydrolase activity, metalloenzyme containing one zinc atom; role in vivo is not defined; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Cysteine aminopeptidase with homocysteine-thiolactonase activity; protects cells against homocysteine toxicity; has bleomycin hydrolase activity in vitro; transcription is regulated by galactose via Gal4p; orthologous to human BLMH | Subunit of mitochondrial NAD(+)-dependent isocitrate dehydrogenase, which catalyzes the oxidation of isocitrate to alpha-ketoglutarate in the TCA cycle; phosphorylated | Alpha subunit of succinyl-CoA ligase, which is a mitochondrial enzyme of the TCA cycle that catalyzes the nucleotide-dependent conversion of succinyl-CoA to succinate; phosphorylated | Vacuolar aspartyl protease (proteinase A), required for the posttranslational precursor maturation of vacuolar proteinases; important for protein turnover after oxidative damage; synthesized as a zymogen, self-activates | Fumarase, converts fumaric acid to L-malic acid in the TCA cycle; cytosolic and mitochondrial localization determined by the N-terminal mitochondrial targeting sequence and protein conformation; phosphorylated in mitochondria |
| 126 | 33 | 435 | 0.090 | 0.497 |  |  |  |  |  | YDR242W YGL087C YJR085C YJR149W YMR077C YMR275C YNL157W YNL293W YOL099C YOR003W YOR208W YPR127W YBR077C YGL185C YJL036W YJR036C YJR052W YJR130C YKL189W YKR091W YKR096W YLR031W YLR423C YMR029C YMR097C YMR140W YNL159C YNR007C YOL159C YOR097C YPL003W YPL055C YPL191C | AMD2 MMS2 VPS20 RDS1 IGO1 MSB3 YSP3 PTP2 SLM4 SNX4 HUL4 RAD7 STR2 HYM1 SRL3 ATG17 FAR8 MTG1 SIP5 ASI2 ATG3 YOL159C-A ULA1 LGE1 | Putative amidase | Protein involved in error-free postreplication DNA repair; forms a heteromeric complex with Ubc13p that has a ubiquitin-conjugating activity; cooperates with chromatin-associated RING finger proteins, Rad18p and Rad5p | Putative protein of unknown function; GFP-fusion protein is induced in response to the DNA-damaging agent MMS; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Myristoylated subunit of ESCRTIII, the endosomal sorting complex required for transport of transmembrane proteins into the multivesicular body pathway to the lysosomal/vacuolar lumen; cytoplasmic protein recruited to endosomal membranes | Zinc cluster transcription factor involved in conferring resistance to cycloheximide | hypothetical protein | GTPase-activating protein for Sec4p and several other Rab GTPases, regulates exocytosis via its action on Sec4p, also required for proper actin organization; similar to Msb4p; both Msb3p and Msb4p localize to sites of polarized growth | Putative precursor to the subtilisin-like protease III | Phosphotyrosine-specific protein phosphatase involved in the inactivation of mitogen-activated protein kinase (MAPK) during osmolarity sensing; dephosporylates Hog1p MAPK and regulates its localization; localized to the nucleus | Putative protein of unknown function; expression is activated by transcription factor YRM1/YOR172W; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus | Component of the EGO complex, which is involved in the regulation of microautophagy, and of the GSE complex, which is required for proper sorting of amino acid permease Gap1p; gene exhibits synthetic genetic interaction with MSS4 | Putative protein with sequence similarity to hydroxyacid dehydrogenases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Sorting nexin, involved in the retrieval of late-Golgi SNAREs from the post-Golgi endosome to the trans-Golgi network and in cytoplasm to vacuole transport; contains a PX domain; forms complex with Snx41p and Atg20p | Protein with similarity to hect domain E3 ubiquitin-protein ligases, not essential for viability | Protein that recognizes and binds damaged DNA in an ATP-dependent manner (with Rad16p) during nucleotide excision repair; subunit of Nucleotide Excision Repair Factor 4 (NEF4) and the Elongin-Cullin-Socs (ECS) ligase complex | Cystathionine gamma-synthase, converts cysteine into cystathionine | Component of the RAM signaling network that is involved in regulation of Ace2p activity and cellular morphogenesis, interacts with Kic1p and Sog2p, localizes to sites of polarized growth during budding and during the mating response | Cytoplasmic protein that, when overexpressed, suppresses the lethality of a rad53 null mutation; potential Cdc28p substrate | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and cytoplasm; predicted to contain a PINc (PilT N terminus) domain | Putative protein of unknown function | Scaffold potein responsible for pre-autophagosomal structure organization; interacts with and is required for activation of Apg1p protein kinase; involved in autophagy but not in the Cvt (cytoplasm to vacuole targeting) pathway | Protein involved in G1 cell cycle arrest in response to pheromone, in a pathway different from the Far1p-dependent pathway; interacts with Far3p, Far7p, Far9p, Far10p, and Far11p | Peripheral GTPase of the mitochondrial inner membrane, essential for respiratory competence, likely functions in assembly of the large ribosomal subunit, has homologs in plants and animals | Protein of unknown function; interacts with both the Reg1p/Glc7p phosphatase and the Snf1p kinase | Integral inner nuclear membrane protein that acts with Asi1p and Asi3p to ensure the fidelity of SPS-sensor signalling by maintaining the dormant repressed state of gene expression in the absence of inducing signals | E2-like enzyme involved in autophagy and the cytoplasm-to-vacuole targeting (Cvt) pathway; plays a role in formation of Atg8p-phosphatidylethanolamine conjugates, which are involved in membrane dynamics during autophagy and Cvt | Putative protein of unknown function; identified by sequence comparison with hemiascomycetous yeast species | Putative protein of unknown function; identified as interacting with Hsp82p in a high-throughput two-hybrid screen; YOR097C is not an essential gene | Protein that acts together with Uba3p to activate Rub1p before its conjugation to proteins (neddylation), which may play a role in protein degradation | Protein of unknown function; null mutant forms abnormally large cells | Putative protein of unknown function; diploid deletion strain exhibits high budding index; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm |
| 251 | 44 | 431 | 0.112 | 0.528 | 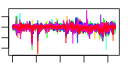 |  | 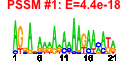 |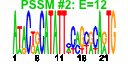  |  | YDR212W YJL186W YKR082W YML046W YMR033W YNR051C YOL076W YOR143C YOR175C YOR355W YBR017C YDL145C YDR303C YDR395W YER125W YGL137W YGL238W YGR218W YGR240C YJR069C YJR075W YKL057C YKL068W YKL210W YLR100W YLR195C YLR347C YML126C YMR308C YNL085W YNL154C YNL287W YOL020W YOL057W YOL098C YOL145C YOR241W YOR254C YOR255W YOR270C YOR326W YPL243W YPL244C YPL254W | TCP1 MNN5 NUP133 PRP39 ARP9 BRE5 MDM20 THI80 ALE1 GDS1 KAP104 COP1 RSC3 SXM1 RSP5 SEC27 CSE1 CRM1 PFK1 HAM1 HOC1 NUP120 YKL068W-A UBA1 ERG27 NMT1 KAP95 ERG13 PSE1 MKT1 YCK2 SEC21 TAT2 CTR9 MET7 SEC63 OSW1 VPH1 MYO2 SRP68 HUT1 HFI1 | Alpha subunit of chaperonin-containing T-complex, which mediates protein folding in the cytosol; involved in maintenance of actin cytoskeleton; homolog to Drosophila melanogaster and mouse tailless complex polypeptide | Alpha-1,2-mannosyltransferase, responsible for addition of the second alpha-1,2-linked mannose of the branches on the mannan backbone of oligosaccharides, localizes to an early Golgi compartment | Subunit of the Nup84p subcomplex of the nuclear pore complex (NPC), localizes to both sides of the NPC, required to establish a normal nucleocytoplasmic concentration gradient of the GTPase Gsp1p | U1 snRNP protein involved in splicing, contains multiple tetriatricopeptide repeats | Component of both the SWI/SNF and RSC chromatin remodeling complexes; actin-related protein involved in transcriptional regulation | Ubiquitin protease cofactor, forms deubiquitination complex with Ubp3p that coregulates anterograde and retrograde transport between the endoplasmic reticulum and Golgi compartments; null is sensitive to brefeldin A | Non-catalytic subunit of the NatB N-terminal acetyltransferase, which catalyzes N-acetylation of proteins with specific N-terminal sequences; involved in mitochondrial inheritance and actin assembly | Thiamine pyrophosphokinase, phosphorylates thiamine to produce the coenzyme thiamine pyrophosphate (thiamine diphosphate) | Lysophospholipid acyltransferase, partially redundant with Slc1p; part of MBOAT family of membrane-bound O-acyltransferases; key component of Lands cycle; may have role in fatty acid exchange at sn-2 position of mature glycerophospholipids | Protein of unknown function, required for growth on glycerol as a carbon source; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Transportin, cytosolic karyopherin beta 2 involved in delivery of heterogeneous nuclear ribonucleoproteins to the nucleoplasm, binds rg-nuclear localization signals on Nab2p and Hrp1p, plays a role in cell-cycle progression | Alpha subunit of COPI vesicle coatomer complex, which surrounds transport vesicles in the early secretory pathway | Component of the RSC chromatin remodeling complex; essential gene required for maintenance of proper ploidy and regulation of ribosomal protein genes and the cell wall/stress response; highly similar to Rsc30p | Nuclear transport factor (karyopherin) involved in protein transport between the cytoplasm and nucleoplasm; similar to Nmd5p, Cse1p, Lph2p, and the human cellular apoptosis susceptibility protein, CAS1 | Ubiquitin-protein ligase involved in ubiquitin-mediated protein degradation; functions in multivesicular body sorting, heat shock response and ubiquitylation of arrested RNAPII; contains a hect (homologous to E6-AP carboxyl terminus) domain | Essential beta'-coat protein of the COPI coatomer, involved in ER-to-Golgi and Golgi-to-ER transport; contains WD40 domains that mediate cargo selective interactions; 45% sequence identity to mammalian beta'-COP | Nuclear envelope protein that mediates the nuclear export of importin alpha (Srp1p), homolog of metazoan CAS protein, required for accurate chromosome segregation | Major karyopherin, involved in export of proteins, RNAs, and ribosomal subunits from the nucleus | Alpha subunit of heterooctameric phosphofructokinase involved in glycolysis, indispensable for anaerobic growth, activated by fructose-2,6-bisphosphate and AMP, mutation inhibits glucose induction of cell cycle-related genes | Conserved protein with deoxyribonucleoside triphosphate pyrophosphohydrolase activity, mediates exclusion of noncanonical purines from deoxyribonucleoside triphosphate pools; mutant is sensitive to the base analog 6-N-hydroxylaminopurine | Alpha-1,6-mannosyltransferase involved in cell wall mannan biosynthesis; subunit of a Golgi-localized complex that also contains Anp1p, Mnn9p, Mnn11p, and Mnn10p; identified as a suppressor of a cell lysis sensitive pkc1-371 allele | Subunit of the Nup84p subcomplex of the nuclear pore complex (NPC), required for even distribution of NPCs around the nuclear envelope, involved in establishment of a normal nucleocytoplasmic concentration gradient of the GTPase Gsp1p | Putative protein of unknown function; identified by homology to Ashbya gossypii | Ubiquitin activating enzyme (E1), involved in ubiquitin-mediated protein degradation and essential for viability | 3-keto sterol reductase, catalyzes the last of three steps required to remove two C-4 methyl groups from an intermediate in ergosterol biosynthesis; mutants are sterol auxotrophs | N-myristoyl transferase, catalyzes the cotranslational, covalent attachment of myristic acid to the N-terminal glycine residue of several proteins involved in cellular growth and signal transduction | Karyopherin beta, forms a dimeric complex with Srp1p (Kap60p) that mediates nuclear import of cargo proteins via a nuclear localization signal (NLS), interacts with nucleoporins to guide transport across the nuclear pore complex | 3-hydroxy-3-methylglutaryl-CoA (HMG-CoA) synthase, catalyzes the formation of HMG-CoA from acetyl-CoA and acetoacetyl-CoA; involved in the second step in mevalonate biosynthesis | Karyopherin/importin that interacts with the nuclear pore complex; acts as the nuclear import receptor for specific proteins, including Pdr1p, Yap1p, Ste12p, and Aft1p | Protein that forms a complex with Pbp1p that may mediate posttranscriptional regulation of HO endonuclease; involved in propagation of M2 dsRNA satellite of L-A virus; contains a DTG signature typical of retroviral proteases | Palmitoylated, plasma membrane-bound casein kinase I isoform; shares redundant functions with Yck1p in morphogenesis, proper septin assembly, endocytic trafficking; provides an essential function overlapping with that of Yck1p | Gamma subunit of coatomer, a heptameric protein complex that together with Arf1p forms the COPI coat; involved in ER to Golgi transport of selective cargo | High affinity tryptophan and tyrosine permease, overexpression confers FK506 and FTY720 resistance | Putative metalloprotease | Component of the Paf1p complex, which is a large complex that binds to and modulates the activity of RNA polymerase II and is required for expression of a subset of genes, including cyclin genes; contains TPR repeats | Folylpolyglutamate synthetase, catalyzes extension of the glutamate chains of the folate coenzymes, required for methionine synthesis and for maintenance of mitochondrial DNA | Essential subunit of Sec63 complex (Sec63p, Sec62p, Sec66p and Sec72p); with Sec61 complex, Kar2p/BiP and Lhs1p forms a channel competent for SRP-dependent and post-translational SRP-independent protein targeting and import into the ER | Protein involved in sporulation; required for the construction of the outer spore wall layers; required for proper localization of Spo14p | Subunit a of vacuolar-ATPase V0 domain, one of two isoforms (Vph1p and Stv1p); Vph1p is located in V-ATPase complexes of the vacuole while Stv1p is located in V-ATPase complexes of the Golgi and endosomes | One of two type V myosin motors (along with MYO4) involved in actin-based transport of cargos; required for the polarized delivery of secretory vesicles, the vacuole, late Golgi elements, peroxisomes, and the mitotic spindle | Core component of the signal recognition particle (SRP) ribonucleoprotein (RNP) complex that functions in targeting nascent secretory proteins to the endoplasmic reticulum (ER) membrane | Protein with a role in UDP-galactose transport to the Golgi lumen, has similarity to human UDP-galactose transporter UGTrel1, exhibits a genetic interaction with S. cerevisiae ERO1 | Adaptor protein required for structural integrity of the SAGA complex, a histone acetyltransferase-coactivator complex that is involved in global regulation of gene expression through acetylation and transcription functions |
| 286 | 36 | 435 | 0.127 | 0.527 | 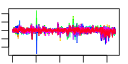 |  | 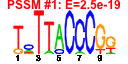 | 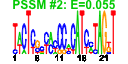 |  | YBL050W YDL115C YGR009C YKL002W YLR025W YAL028W YBL102W YBR290W YDR073W YDR196C YDR197W YDR329C YER063W YGL119W YGL181W YGL219C YGL237C YGR046W YGR232W YHR002W YHR004C YJL053W YJR086W YKL058W YKR062W YLR093C YLR097C YLR151C YLR246W YLR281C YLR283W YMR311C YOL099C YPL224C YPL225W YPR148C | SEC17 IWR1 SEC9 DID4 SNF7 FRT2 SFT2 BSD2 SNF11 CBS2 PEX3 THO1 ABC1 GTS1 MDM34 HAP2 TAM41 NAS6 LEU5 NEM1 PEP8 STE18 TOA2 TFA2 NYV1 HRT3 PCD1 ERF2 GLC8 MMT2 | Peripheral membrane protein required for vesicular transport between ER and Golgi and for the 'priming' step in homotypic vacuole fusion, part of the cis-SNARE complex; has similarity to alpha-SNAP | Protein of unknown function, deletion causes hypersensitivity to the K1 killer toxin | t-SNARE protein important for fusion of secretory vesicles with the plasma membrane; similar to but not functionally redundant with Spo20p; SNAP-25 homolog | Class E Vps protein of the ESCRT-III complex, required for sorting of integral membrane proteins into lumenal vesicles of multivesicular bodies, and for delivery of newly synthesized vacuolar enzymes to the vacuole, involved in endocytosis | One of four subunits of the endosomal sorting complex required for transport III (ESCRT-III); involved in the sorting of transmembrane proteins into the multivesicular body (MVB) pathway; recruited from the cytoplasm to endosomal membranes | Tail-anchored endoplasmic reticulum membrane protein, interacts with homolog Frt1p but is not a substrate of calcineurin (unlike Frt1p), promotes growth in conditions of high Na+, alkaline pH, or cell wall stress; potential Cdc28p substrate | Non-essential tetra-spanning membrane protein found mostly in the late Golgi, can suppress some sed5 alleles; may be part of the transport machinery, but precise function is unknown; similar to mammalian syntaxin 5 | Heavy metal ion homeostasis protein, facilitates trafficking of Smf1p and Smf2p metal transporters to the vacuole where they are degraded, controls metal ion transport, prevents metal hyperaccumulation, functions in copper detoxification | Subunit of the SWI/SNF chromatin remodeling complex involved in transcriptional regulation; interacts with a highly conserved 40-residue sequence of Snf2p | Putative dephospho-CoA kinase (DPCK) that catalyzes the final step in Coenzyme A biosynthesis; essential for viability; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Mitochondrial translational activator of the COB mRNA; interacts with translating ribosomes, acts on the COB mRNA 5'-untranslated leader | Peroxisomal membrane protein (PMP) required required for the proper localization and stability of PMPs; interacts with Pex19p | Conserved nuclear RNA-binding protein; specifically binds to transcribed chromatin in a THO- and RNA-dependent manner, genetically interacts with shuttling hnRNP NAB2; overproduction suppresses transcriptional defect caused by hpr1 mutation | Protein required for ubiquinone (coenzyme Q) biosynthesis and for respiratory growth; exhibits genetic interaction with COQ9, suggesting a common function; similar to prokaryotic proteins involved in early steps of ubiquinone biosynthesis | Protein containing a zinc-finger in the N-terminus and a long Gln-rich region in the C-terminus; regulates ultradian rhythm, cell size, cell cycle, lifespan, sporulation, heat tolerance, and multidrug transport | Mitochondrial outer membrane protein, required for normal mitochondrial morphology and inheritance; localizes to dots on the mitochondrial surface near mtDNA nucleoids; interacts genetically with MDM31 and MDM32 | Subunit of the heme-activated, glucose-repressed Hap2p/3p/4p/5p CCAAT-binding complex, a transcriptional activator and global regulator of respiratory gene expression; contains sequences sufficient for both complex assembly and DNA binding | Mitochondrial protein involved in protein import into the mitochondrial matrix; maintains the functional integrity of the TIM23 protein translocator complex; viability of null mutant is strain-dependent; mRNA is targeted to the bud | Regulatory, non-ATPase subunit of the 26S proteasome; homolog of the human oncoprotein gankyrin, which interacts with the retinoblastoma tumor suppressor (Rb) and cyclin-dependent kinase 4/6 | Mitochondrial carrier protein involved in the accumulation of CoA in the mitochondrial matrix; homolog of human Graves disease protein; does not encode an isozyme of Leu4p, as first hypothesized | Catalytic subunit of Nem1p-Spo7p phosphatase holoenzyme, which regulates nuclear growth by controlling recruitment of Pah1p onto promoters of phospholipid biosynthetic genes; required for normal nuclear envelope morphology and sporulation | Vacuolar protein sorting protein that forms part of the multimeric membrane-associated retromer complex along with Vps35p, Vps29p, Vps17p, and Vps5p; essential for endosome-to-Golgi retrograde protein transport | G protein gamma subunit, forms a dimer with Ste4p to activate the mating signaling pathway, forms a heterotrimer with Gpa1p and Ste4p to dampen signaling; C-terminus is palmitoylated and farnesylated, which are required for normal signaling | TFIIA small subunit; involved in transcriptional activation, acts as antirepressor or as coactivator; homologous to smallest subunit of human and Drosophila TFIIA | TFIIE small subunit, involved in RNA polymerase II transcription initiation | v-SNARE component of the vacuolar SNARE complex involved in vesicle fusion; inhibits ATP-dependent Ca(2+) transport activity of Pmc1p in the vacuolar membrane | Putative SCF-ubiquitin ligase F-box protein, based on both genetic and physical interactions and sequence similarity; identified in association with Cdc53p, Skp1p and Ubi4 in large and small-scale studies | Peroxisomal nudix pyrophosphatase with specificity for coenzyme A and CoA derivatives, may function to remove potentially toxic oxidized CoA disulfide from peroxisomes to maintain the capacity for beta-oxidation of fatty acids | Subunit of a palmitoyltransferase, composed of Erf2p and Shr5p, that adds a palmitoyl lipid moiety to heterolipidated substrates such as Ras1p and Ras2p through a thioester linkage; mutants partially mislocalize Ras2p to the vacuole | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YLR281C is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YLR283W is not an essential gene | Regulatory subunit of protein phosphatase 1 (Glc7p), involved in glycogen metabolism and chromosome segregation; proposed to regulate Glc7p activity via conformational alteration; ortholog of the mammalian protein phosphatase inhibitor 2 | Putative metal transporter involved in mitochondrial iron accumulation; closely related to Mmt1p | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern |
| 25 | 30 | 441 | 0.131 | 0.566 |  |  | 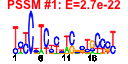 | 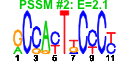 |  | YAL060W YCL044C YCR061W YCR062W YDR474C YDR475C YER088C YER119C YER143W YFL044C YIL155C YJL021C YJR079W YKL085W YKR067W YLL019C YLR142W YLR152C YML120C YML129C YMR031C YMR302C YNL173C YNR014W YOL073C YOR040W YPL134C YPL164C YPR002W YER142C | BDH1 MGR1 JIP4 DOT6 AVT6 DDI1 OTU1 GUT2 BBC1 MDH1 GAT1 KNS1 PUT1 NDI1 COX14 YME2 MDG1 GLO4 ODC1 MLH3 PDH1 MAG1 | NAD-dependent (R,R)-butanediol dehydrogenase, catalyzes oxidation of (R,R)-2,3-butanediol to (3R)-acetoin, oxidation of meso-butanediol to (3S)-acetoin, and reduction of acetoin; enhances use of 2,3-butanediol as an aerobic carbon source | Subunit, with Yme1p, of the mitochondrial inner membrane i-AAA protease complex, which is responsible for degradation of unfolded or misfolded mitochondrial gene products; required for growth of cells lacking the mitochondrial genome | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Protein of unknown function; previously annotated as two separate ORFs, YDR474C and YDR475C, which were merged as a result of corrections to the systematic reference sequence | Protein of unknown function, involved in telomeric gene silencing and filamentation | Vacuolar amino acid transporter, exports aspartate and glutamate from the vacuole; member of a family of seven S. cerevisiae genes (AVT1-7) related to vesicular GABA-glycine transporters | DNA damage-inducible v-SNARE binding protein, contains a ubiquitin-associated (UBA) domain, may act as a negative regulator of constitutive exocytosis, may play a role in S-phase checkpoint control | Deubiquitylation enzyme that binds to the chaperone-ATPase Cdc48p; may contribute to regulation of protein degradation by deubiquitylating substrates that have been ubiquitylated by Ufd2p; member of the Ovarian Tumor (OTU) family | Mitochondrial glycerol-3-phosphate dehydrogenase; expression is repressed by both glucose and cAMP and derepressed by non-fermentable carbon sources in a Snf1p, Rsf1p, Hap2/3/4/5 complex dependent manner | Protein possibly involved in assembly of actin patches; interacts with an actin assembly factor Las17p and with the SH3 domains of Type I myosins Myo3p and Myo5p; localized predominantly to cortical actin patches | Putative protein of unknown function; mutation results in impaired mitochondrial respiration | Mitochondrial malate dehydrogenase, catalyzes interconversion of malate and oxaloacetate; involved in the tricarboxylic acid (TCA) cycle; phosphorylated | Transcriptional activator of genes involved in nitrogen catabolite repression; contains a GATA-1-type zinc finger DNA-binding motif; activity and localization regulated by nitrogen limitation and Ure2p | Nonessential putative protein kinase of unknown cellular role; member of the LAMMER family of protein kinases, which are serine/threonine kinases also capable of phosphorylating tyrosine residues | Proline oxidase, nuclear-encoded mitochondrial protein involved in utilization of proline as sole nitrogen source; PUT1 transcription is induced by Put3p in the presence of proline and the absence of a preferred nitrogen source | Putative protein of unknown function; YLR152C is not an essential gene | NADH:ubiquinone oxidoreductase, transfers electrons from NADH to ubiquinone in the respiratory chain but does not pump protons, in contrast to the higher eukaryotic multisubunit respiratory complex I; phosphorylated; homolog of human AMID | Mitochondrial membrane protein, involved in translational regulation of Cox1p and assembly of cytochrome c oxidase (complex IV); associates with complex IV assembly intermediates and complex III/complex IV supercomplexes | Protein of unknown function with similarity to Ykl050cp and Uso1p; the authentic, non-tagged protein is detected in a phosphorylated state in highly purified mitochondria in high-throughput studies; YMR031C is not an essential gene | Integral inner mitochondrial membrane protein with a role in maintaining mitochondrial nucleoid structure and number; mutants exhibit an increased rate of mitochondrial DNA escape; shows some sequence similarity to exonucleases | Plasma membrane protein involved in G-protein mediated pheromone signaling pathway; overproduction suppresses bem1 mutations | Putative protein of unknown function; expression is cell-cycle regulated, Azf1p-dependent, and heat-inducible | Putative protein of unknown function | Mitochondrial glyoxalase II, catalyzes the hydrolysis of S-D-lactoylglutathione into glutathione and D-lactate | Mitochondrial inner membrane transporter, exports 2-oxoadipate and 2-oxoglutarate from the mitochondrial matrix to the cytosol for lysine and glutamate biosynthesis and lysine catabolism; suppresses, in multicopy, an fmc1 null mutation | Protein involved in DNA mismatch repair and crossing-over during meiotic recombination; forms a complex with Mlh1p; mammalian homolog is implicated mammalian microsatellite instability | Mitochondrial protein that participates in respiration, induced by diauxic shift; homologous to E. coli PrpD, may take part in the conversion of 2-methylcitrate to 2-methylisocitrate | 3-methyl-adenine DNA glycosylase involved in protecting DNA against alkylating agents; initiates base excision repair by removing damaged bases to create abasic sites that are subsequently repaired |
| 33 | 52 | 435 | 0.140 | 0.524 | 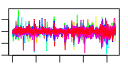 |  | 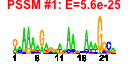 | 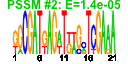 |  | YBL038W YBR146W YBR185C YBR268W YBR282W YCR003W YDR041W YDR071C YDR100W YDR116C YDR494W YEL050C YFR011C YGL068W YGL069C YGR076C YGR084C YGR101W YGR219W YGR220C YHR147C YIL098C YJL046W YJL063C YJL096W YJL180C YJR034W YKL003C YKR006C YKR016W YLL009C YLR069C YLR168C YLR439W YMR024W YMR157C YMR158W YNL122C YNL252C YNL263C YNL315C YNR037C YNR038W YOR158W YOR187W YPL172C YPR100W YPR166C YDR404C YJR079W YKL167C YLR203C | MRPL16 MRPS9 MBA1 MRPL37 MRPL27 MRPL32 RSM10 PAA1 TVP15 MRPL1 RSM28 RML2 AIM13 MNP1 MRPL25 MRP13 PCP1 MRPL9 MRPL6 FMC1 AIM22 MRPL8 MRPL49 ATP12 PET191 MRP17 MRPL13 AIM28 COX17 MEF1 AIM30 MRPL4 MRPL3 AIM36 MRPS8 MRPL17 YIF1 ATP11 RSM19 DBP6 PET123 RAP1 COX10 MRPL51 MRP2 RPB7 MRP49 MSS51 | Mitochondrial ribosomal protein of the large subunit | Mitochondrial ribosomal protein of the small subunit | Protein involved in assembly of mitochondrial respiratory complexes; may act as a receptor for proteins destined for export from the mitochondrial matrix to the inner membrane | Mitochondrial ribosomal protein of the small subunit, has similarity to E. coli S10 ribosomal protein; essential for viability, unlike most other mitoribosomal proteins | Polyamine acetyltransferase; acetylates polyamines (e.g. putrescine, spermidine, spermine) and also aralkylamines (e.g. tryptamine, phenylethylamine); may be involved in transcription and/or DNA replication | Integral membrane protein localized to late Golgi vesicles along with the v-SNARE Tlg2p | Mitochondrial ribosomal protein of the small subunit; genetic interactions suggest a possible role in promoting translation initiation | Mitochondrial ribosomal protein of the large subunit, has similarity to E. coli L2 ribosomal protein; fat21 mutant allele causes inability to utilize oleate and may interfere with activity of the Adr1p transcription factor | Putative protein of unknown function; non-tagged protein is detected in highly purified mitochondria; null mutant displays increased frequency of mitochondrial genome loss (petite formation) and reduced growth rate in minimal glycerol media | Protein associated with the mitochondrial nucleoid; putative mitochondrial ribosomal protein with similarity to E. coli L7/L12 ribosomal protein; required for normal respiratory growth | Mitochondrial serine protease required for the processing of various mitochondrial proteins and maintenance of mitochondrial DNA and morphology; belongs to the rhomboid-GlpG superfamily of intramembrane peptidases | Mitochondrial matrix protein, required for assembly or stability at high temperature of the F1 sector of mitochondrial F1F0 ATP synthase; null mutant temperature sensitive growth on glycerol is suppressed by multicopy expression of Odc1p | Putative lipoate-protein ligase A family member; null mutant displays respiratory growth defect, decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Conserved protein required for assembly of alpha and beta subunits into the F1 sector of mitochondrial F1F0 ATP synthase; mutation of human ATP12 reduces active ATP synthase levels and is associated with the disorder ATPAF2 deficiency | Protein required for assembly of cytochrome c oxidase; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Mitochondrial ribosomal protein of the small subunit; MRP17 exhibits genetic interactions with PET122, encoding a COX3-specific translational activator | Mitochondrial ribosomal protein of the large subunit, not essential for mitochondrial translation | Protein of unknown function; non-tagged protein is detected in purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss and severe growth defect in minimal glycerol media | Copper metallochaperone that transfers copper to Sco1p and Cox11p for eventual delivery to cytochrome c oxidase | Mitochondrial elongation factor involved in translational elongation | Putative protein of unknown function that may be involved in intramitochondrial sorting; similar to Ups1p and to human PRELI; GFP-tagged protein localizes to mitochondria; required for wild-type respiratory growth | Protein of unknown function; non-tagged protein is detected in purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss and reduced growth rate in minimal glycerol media | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YNL122C is not an essential gene | Integral membrane protein required for the fusion of ER-derived COPII transport vesicles with the Golgi; interacts with Yip1p and Yos1p; localizes to the Golgi, the ER, and COPII vesicles | Molecular chaperone, required for the assembly of alpha and beta subunits into the F1 sector of mitochondrial F1F0 ATP synthase | Mitochondrial ribosomal protein of the small subunit, has similarity to E. coli S19 ribosomal protein | Essential protein involved in ribosome biogenesis; putative ATP-dependent RNA helicase of the DEAD-box protein family | Mitochondrial ribosomal protein of the small subunit; PET123 exhibits genetic interactions with PET122, which encodes a COX3 mRNA-specific translational activator | DNA-binding protein involved in either activation or repression of transcription, depending on binding site context; also binds telomere sequences and plays a role in telomeric position effect (silencing) and telomere structure | Heme A:farnesyltransferase, catalyzes the first step in the conversion of protoheme to the heme A prosthetic group required for cytochrome c oxidase activity; human ortholog is associated with mitochondrial disorders | RNA polymerase II subunit B16; forms two subunit dissociable complex with Rpb4p | Putative protein of unknown function; mutation results in impaired mitochondrial respiration | Nuclear encoded protein required for translation of COX1 mRNA; binds to Cox1 protein |
| 74 | 45 | 442 | 0.144 | 0.540 | 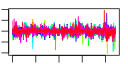 |  | 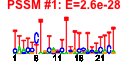 | 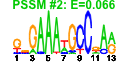 |  | YBR097W YCL068C YDL035C YDR040C YER014W YGL126W YGL127C YIL080W YIL105C YIR015W YKR021W YKR103W YLR131C YLR278C YLR318W YML017W YMR036C YMR069W YMR165C YMR280C YNL269W YNR069C YOR058C YOR148C YOR380W YOR381W YOR388C YPL016W YPL058C YPL073C YPL167C YPR008W YPR095C YPR115W YCR001W YGL114W YHR151C YIL082W YKL134C YML087C YMR101C YMR102C YMR207C YNL242W YPL039W | VPS15 GPR1 ENA1 HEM14 SCS3 SOH1 SLM1 RPR2 ALY1 NFT1 ACE2 EST2 PSP2 MIH1 NAT4 PAH1 CAT8 BSC4 BSC5 ASE1 SPP2 RDR1 FRE3 FDH1 SWI1 PDR12 REV3 HAA1 SYT1 YIL082W-A OCT1 AIM33 SRT1 HFA1 ATG2 | Myristoylated serine/threonine protein kinase involved in vacuolar protein sorting; functions as a membrane-associated complex with Vps34p; active form recruits Vps34p to the Golgi membrane; interacts with the GDP-bound form of Gpa1p | Putative protein of unknown function | Plasma membrane G protein coupled receptor (GPCR) that interacts with the heterotrimeric G protein alpha subunit, Gpa2p, and with Plc1p; sensor that integrates nutritional signals with the modulation of cell fate via PKA and cAMP synthesis | P-type ATPase sodium pump, involved in Na+ and Li+ efflux to allow salt tolerance | Protoporphyrinogen oxidase, a mitochondrial enzyme that catalyzes the seventh step in the heme biosynthetic pathway, converting protoporphyrinogen IX to protoporphyrin IX | Protein required for inositol prototrophy, identified as an ortholog of the FIT family of proteins involved in triglyceride droplet biosynthesis; disputed role in the synthesis of inositol phospholipids from inositol | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; involved in telomere maintenance; conserved with other metazoan MED31 subunits | Phosphoinositide PI4,5P(2) binding protein, forms a complex with Slm2p; acts downstream of Mss4p in a pathway regulating actin cytoskeleton organization in response to stress; subunit of and phosphorylated by the TORC2 complex | Subunit of nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends; not shared between RNase MRP and RNase P, in contrast to all other RNase P protein subunits | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative transporter of the multidrug resistance-associated protein (MRP) subfamily; adjacent ORFs YKR103W and YKR104W are merged in different strain backgrounds. | Transcription factor that activates expression of early G1-specific genes, localizes to daughter cell nuclei after cytokinesis and delays G1 progression in daughters, localization is regulated by phosphorylation; potential Cdc28p substrate | Zinc-cluster protein; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; mutant shows moderate growth defect on caffeine; YLR278C is not an essential gene | Reverse transcriptase subunit of the telomerase holoenzyme, essential for telomerase core catalytic activity, involved in other aspects of telomerase assembly and function; mutations in human homolog are associated with aplastic anemia | Asn rich cytoplasmic protein that contains RGG motifs; high-copy suppressor of group II intron-splicing defects of a mutation in MRS2 and of a conditional mutation in POL1 (DNA polymerase alpha); possible role in mitochondrial mRNA splicing | Protein tyrosine phosphatase involved in cell cycle control; regulates the phosphorylation state of Cdc28p; homolog of S. pombe cdc25 | N alpha-acetyl-transferase, involved in acetylation of the N-terminal residues of histones H4 and H2A | Mg2+-dependent phosphatidate (PA) phosphatase, catalyzes the dephosphorylation of PA to yield diacylglycerol and Pi, responsible for de novo lipid synthesis; homologous to mammalian lipin 1 | Zinc cluster transcriptional activator necessary for derepression of a variety of genes under non-fermentative growth conditions, active after diauxic shift, binds carbon source responsive elements | Protein of unknown function, ORF exhibits genomic organization compatible with a translational readthrough-dependent mode of expression; readthrough is increased upon depletion of Sup35p | Protein of unknown function, ORF exhibits genomic organization compatible with a translational readthrough-dependent mode of expression | Mitotic spindle midzone localized microtubule-associated protein (MAP) family member; required for spindle elongation and stabilization; undergoes cell cycle-regulated degradation by anaphase promoting complex; potential Cdc28p substrate | Essential protein that promotes the first step of splicing and is required for the final stages of spliceosome maturation; interacts with Prp2p, which may release Spp2p from the spliceosome following the first cleavage reaction | Transcriptional repressor involved in the control of multidrug resistance; negatively regulates expression of the PDR5 gene; member of the Gal4p family of zinc cluster proteins | Ferric reductase, reduces siderophore-bound iron prior to uptake by transporters; expression induced by low iron levels | NAD(+)-dependent formate dehydrogenase, may protect cells from exogenous formate | Subunit of the SWI/SNF chromatin remodeling complex, which regulates transcription by remodeling chromosomes; required for transcription of many genes, including ADH1, ADH2, GAL1, HO, INO1 and SUC2 | Plasma membrane ATP-binding cassette (ABC) transporter, weak-acid-inducible multidrug transporter required for weak organic acid resistance; induced by sorbate and benzoate and regulated by War1p; mutants exhibit sorbate hypersensitivity | Catalytic subunit of DNA polymerase zeta, which is involved in DNA repair and translesion synthesis; required for mutagenesis induced by DNA damage | Transcriptional activator involved in the transcription of TPO2, HSP30 and other genes encoding membrane stress proteins; involved in adaptation to weak acid stress | Guanine nucleotide exchange factor (GEF) for Arf proteins; involved in vesicular transport; suppressor of ypt3 mutations; member of the Sec7-domain family | Putative protein of unknown function; predicted member of the oligopeptide transporter (OPT) family of membrane transporters | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Mitochondrial intermediate peptidase, cleaves N-terminal residues of a subset of proteins upon import, after their cleavage by mitochondrial processing peptidase (Mas1p-Mas2p); may contribute to mitochondrial iron homeostasis | Putative protein of unknown function; null mutant displays increased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Cis-prenyltransferase involved in synthesis of long-chain dolichols (19-22 isoprene units; as opposed to Rer2p which synthesizes shorter-chain dolichols); localizes to lipid bodies; transcription is induced during stationary phase | Protein of unknown function; transcription is activated by paralogous transcription factors Yrm1p and Yrr1p along with genes involved in multidrug resistance; mutant shows increased resistance to azoles; YMR102C is not an essential gene | Mitochondrial acetyl-coenzyme A carboxylase, catalyzes the production of malonyl-CoA in mitochondrial fatty acid biosynthesis | Peripheral membrane protein required for the formation of cytosolic sequestering vesicles involved in vacuolar import through both the Cvt pathway and autophagy; interacts with Atg9p and is necessary for its trafficking | Putative protein of unknown function; YPL039W is not an essential gene |
| 213 | 44 | 439 | 0.151 | 0.539 |  |  | 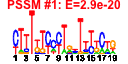 |  |  | YFL061W YJR004C YKL050C YLR371W YLR465C YMR096W YNL146W YNL329C YNL335W YOL011W YOR392W YBR045C YCR014C YDR350C YDR520C YDR543C YER014W YFR026C YIL175W YJR078W YKL055C YKL220C YKR021W YKR105C YLL065W YLR004C YLR306W YLR382C YLR394W YLR424W YML029W YML034W YMR018W YMR036C YMR095C YMR137C YNL337W YNR062C YNR064C YOR140W YOR384W YPL056C YPL277C YPR153W | DDI2 SAG1 ROM2 SNZ1 PEX6 DDI3 PLB3 GIP1 POL4 ATP22 HEM14 BNA2 OAR1 FRE2 ALY1 THI73 UBC12 MSL1 CST9 SPP382 USA1 SRC1 MIH1 SNO1 SNM1 SFL1 FRE5 | Protein whose expression is induced by DNA damage | Alpha-agglutinin of alpha-cells, binds to Aga1p during agglutination, N-terminal half is homologous to the immunoglobulin superfamily and contains binding site for a-agglutinin, C-terminal half is highly glycosylated and contains GPI anchor | Protein of unknown function; the YKL050W protein is a target of the SCFCdc4 ubiquitin ligase complex and YKL050W transcription is regulated by Azf1p | GDP/GTP exchange protein (GEP) for Rho1p and Rho2p; mutations are synthetically lethal with mutations in rom1, which also encodes a GEP | Protein involved in vitamin B6 biosynthesis; member of a stationary phase-induced gene family; coregulated with SNO1; interacts with Sno1p and with Yhr198p, perhaps as a multiprotein complex containing other Snz and Sno proteins | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum; YNL146W is not an essential gene | AAA-peroxin that heterodimerizes with AAA-peroxin Pex1p and participates in the recycling of peroxisomal signal receptor Pex5p from the peroxisomal membrane to the cystosol | hypothetical protein | Phospholipase B (lysophospholipase) involved in phospholipid metabolism; hydrolyzes phosphatidylinositol and phosphatidylserine and displays transacylase activity in vitro | Meiosis-specific regulatory subunit of the Glc7p protein phosphatase, regulates spore wall formation and septin organization, required for expression of some late meiotic genes and for normal localization of Glc7p | DNA polymerase IV, undergoes pair-wise interactions with Dnl4p-Lif1p and Rad27p to mediate repair of DNA double-strand breaks by non-homologous end joining (NHEJ); homologous to mammalian DNA polymerase beta | Mitochondrial inner membrane protein required for assembly of the F0 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis | Putative transcription factor; contains the (Zn(II)2Cys6 motif; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; null mutant is viable and sensitive to caffeine | Protoporphyrinogen oxidase, a mitochondrial enzyme that catalyzes the seventh step in the heme biosynthetic pathway, converting protoporphyrinogen IX to protoporphyrin IX | Putative protein of unknown function | Tryptophan 2,3-dioxygenase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Mitochondrial 3-oxoacyl-[acyl-carrier-protein] reductase, may comprise a type II mitochondrial fatty acid synthase along with Mct1p | Ferric reductase and cupric reductase, reduces siderophore-bound iron and oxidized copper prior to uptake by transporters; expression induced by low iron levels but not by low copper levels | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative transporter of the Major Facilitator Superfamily (MFS) | Putative plasma membrane permease proposed to be involved in carboxylic acid uptake and repressed by thiamine; substrate of Dbf2p/Mob1p kinase; transcription is altered if mitochondrial dysfunction occurs | Enzyme that mediates the conjugation of Rub1p, a ubiquitin-like protein, to other proteins; related to E2 ubiquitin-conjugating enzymes | U2B component of U2 snRNP, involved in splicing, binds the U2 snRNA stem-loop IV in vitro; does not contain the conserved C-terminal RNA binding domain found in other family members | SUMO E3 ligase; required for synaptonemal complex formation; localizes to synapsis initiation sites on meiotic chromosomes; potential Cdc28p substrate | Essential protein that forms a dimer with Ntr2p; also forms a trimer, with Ntr2p and Prp43p, that is involved in spliceosome disassembly; found also in a multisubunit complex with the splicing factor Clf1p; suppressor of prp38-1 mutation | Protein involved in ER-associated protein degradation (ERAD); component of the Hrd1p complex; interacts with the U1 snRNP-specific protein, Snp1p | Inner nuclear membrane (INM) protein with a putative role in sister chromatid segregation, potentially phosphorylated by Cdc28p; contains helix-extension-helix (HEH) motif, nuclear localization signal sequence | Putative protein of unknown function with similarity to human PEX5Rp (peroxin protein 5 related protein); transcription increases during colony development similar to genes involved in peroxisome biogenesis; YMR018W is not an essential gene | Protein tyrosine phosphatase involved in cell cycle control; regulates the phosphorylation state of Cdc28p; homolog of S. pombe cdc25 | Protein of unconfirmed function, involved in pyridoxine metabolism; expression is induced during stationary phase; forms a putative glutamine amidotransferase complex with Snz1p, with Sno1p serving as the glutaminase | Subunit of RNase MRP, which cleaves pre-rRNA and has a role in cell cycle-regulated degradation of daughter cell-specific mRNAs; binds to the NME1 RNA subunit of RNase MRP | Putative membrane protein of unknown function | Epoxide hydrolase, member of the alpha/beta hydrolase fold family; may have a role in detoxification of epoxides | Transcriptional repressor and activator; involved in repression of flocculation-related genes, and activation of stress responsive genes; negatively regulated by cAMP-dependent protein kinase A subunit Tpk2p | Putative ferric reductase with similarity to Fre2p; expression induced by low iron levels; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; deletion mutant is fluconazole resistant | Putative protein of unknown function; localized to the membranes; gene expression regulated by copper levels |
| 168 | 45 | 445 | 0.158 | 0.599 | 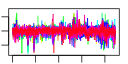 |  | 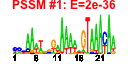 | 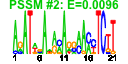 |  | YAR018C YBL010C YBR038W YBR243C YCL012W YCL013W YDR146C YDR147W YGR092W YHR023W YIL038C YIR010W YLR112W YLR353W YML064C YML104C YMR001C YMR032W YOR073W YOR199W YOR239W YPR034W YBL009W YBR092C YCL014W YCL062W YCL063W YDL101C YDL160C YDR325W YHR117W YHR152W YIL131C YJR092W YLR045C YLR131C YML061C YMR198W YNL057W YNL058C YOL123W YOR025W YOR058C YOR195W YPL242C | KIN3 CHS2 ALG7 BUD3 SWI5 EKI1 DBF2 MYO1 NOT3 DSN1 BUD8 TEM1 MDM1 CDC5 HOF1 SGO1 ABP140 ARP7 ALK2 PHO3 VAC17 DUN1 YDL160C-A YCG1 TOM71 SPO12 FKH1 BUD4 STU2 ACE2 PIF1 CIK1 HRP1 HST3 ASE1 SLK19 IQG1 | Nonessential protein kinase with unknown cellular role | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein colocalizes with clathrin-coated vesicles | Chitin synthase II, requires activation from zymogenic form in order to catalyze the transfer of N-acetylglucosamine (GlcNAc) to chitin; required for the synthesis of chitin in the primary septum during cytokinesis | UDP-N-acetyl-glucosamine-1-P transferase, transfers Glc-Nac-P from UDP-GlcNac to Dol-P in the ER in the first step of the dolichol pathway of protein asparagine-linked glycosylation; inhibited by tunicamycin | Protein involved in bud-site selection and required for axial budding pattern; localizes with septins to bud neck in mitosis and may constitute an axial landmark for next round of budding | Transcription factor that activates transcription of genes expressed at the M/G1 phase boundary and in G1 phase; localization to the nucleus occurs during G1 and appears to be regulated by phosphorylation by Cdc28p kinase | Ethanolamine kinase, primarily responsible for phosphatidylethanolamine synthesis via the CDP-ethanolamine pathway; exhibits some choline kinase activity, thus contributing to phosphatidylcholine synthesis via the CDP-choline pathway | Ser/Thr kinase involved in transcription and stress response; functions as part of a network of genes in exit from mitosis; localization is cell cycle regulated; activated by Cdc15p during the exit from mitosis | Type II myosin heavy chain, required for wild-type cytokinesis and cell separation; localizes to the actomyosin ring; binds to myosin light chains Mlc1p and Mlc2p through its IQ1 and IQ2 motifs respectively | Subunit of the CCR4-NOT complex, which is a global transcriptional regulator with roles in transcription initiation and elongation and in mRNA degradation | Essential component of the MIND kinetochore complex (Mtw1p Including Nnf1p-Nsl1p-Dsn1p) which joins kinetochore subunits contacting DNA to those contacting microtubules; important for chromosome segregation | Protein involved in bud-site selection; diploid mutants display a unipolar budding pattern instead of the wild-type bipolar pattern, and bud at the proximal pole | GTP-binding protein of the ras superfamily involved in termination of M-phase; controls actomyosin and septin dynamics during cytokinesis | Intermediate filament protein, required for nuclear and mitochondrial transmission to daughter buds; contains a Phox homology (PX) domain and specifically binds phosphatidylinositol 3-phosphate (PtdIns-3-P) | Polo-like kinase with similarity to Xenopus Plx1 and S. pombe Plo1p; found at bud neck, nucleus and SPBs; has multiple functions in mitosis and cytokinesis through phosphorylation of substrates; may be a Cdc28p substrate | Bud neck-localized, SH3 domain-containing protein required for cytokinesis; regulates actomyosin ring dynamics and septin localization; interacts with the formins, Bni1p and Bnr1p, and with Cyk3p, Vrp1p, and Bni5p | Component of the spindle checkpoint, involved in sensing lack of tension on mitotic chromosomes; protects centromeric Rec8p at meiosis I; required for accurate chromosomal segregation at meiosis II and for mitotic chromosome stability | Nonessential protein that binds actin filaments and localizes to actin patches and cables, has similarity to S-adenosylmethionine (AdoMet)-dependent methyltransferases | Component of both the SWI/SNF and RSC chromatin remodeling complexes; actin-related protein involved in transcriptional regulation | Protein kinase; accumulation and phosphorylation are periodic during the cell cycle; phosphorylated in response to DNA damage; contains characteristic motifs for degradation via the APC pathway; similar to Alk1p and to mammalian haspins | Constitutively expressed acid phosphatase similar to Pho5p; brought to the cell surface by transport vesicles; hydrolyzes thiamin phosphates in the periplasmic space, increasing cellular thiamin uptake; expression is repressed by thiamin | Protein involved in vacuole inheritance; acts as a vacuole-specific receptor for myosin Myo2p | Cell-cycle checkpoint serine-threonine kinase required for DNA damage-induced transcription of certain target genes, phosphorylation of Rad55p and Sml1p, and transient G2/M arrest after DNA damage; also regulates postreplicative DNA repair | Putative protein of unknown function | Non-SMC subunit of the condensin complex (Smc2p-Smc4p-Ycs4p-Brn1p-Ycg1p); required for establishment and maintenance of chromosome condensation, chromosome segregation and for chromatin binding of the condensin complex | Mitochondrial outer membrane protein with similarity to Tom70p; probable minor component of the TOM (translocase of outer membrane) complex responsible for recognition and import of mitochondrially directed proteins | Nucleolar protein of unknown function, positive regulator of exit from mitosis; involved in regulating the release of Cdc14p from the nucleolus in early anaphase; proposed to play similar role in meiosis | Forkhead family transcription factor with a minor role in the expression of G2/M phase genes; negatively regulates transcriptional elongation; positive role in chromatin silencing at HML and HMR; regulates donor preference during switching | Protein involved in bud-site selection and required for axial budding pattern; localizes with septins to bud neck in mitosis and may constitute an axial landmark for next round of budding; potential Cdc28p substrate | Microtubule-associated protein (MAP) of the XMAP215/Dis1 family; regulates microtubule dynamics during spindle orientation and metaphase chromosome alignment; interacts with spindle pole body component Spc72p | Transcription factor that activates expression of early G1-specific genes, localizes to daughter cell nuclei after cytokinesis and delays G1 progression in daughters, localization is regulated by phosphorylation; potential Cdc28p substrate | DNA helicase involved in telomere formation and elongation; acts as a catalytic inhibitor of telomerase; also plays a role in repair and recombination of mitochondrial DNA | Kinesin-associated protein required for both karyogamy and mitotic spindle organization, interacts stably and specifically with Kar3p and may function to target this kinesin to a specific cellular role; has similarity to Vik1p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; YNL058C is not an essential gene | Subunit of cleavage factor I, a five-subunit complex required for the cleavage and polyadenylation of pre-mRNA 3' ends; RRM-containing heteronuclear RNA binding protein and hnRNPA/B family member that binds to poly (A) signal sequences | Member of the Sir2 family of NAD(+)-dependent protein deacetylases; involved along with Hst4p in telomeric silencing, cell cycle progression, radiation resistance, genomic stability and short-chain fatty acid metabolism | Mitotic spindle midzone localized microtubule-associated protein (MAP) family member; required for spindle elongation and stabilization; undergoes cell cycle-regulated degradation by anaphase promoting complex; potential Cdc28p substrate | Kinetochore-associated protein required for normal segregation of chromosomes in meiosis and mitosis; component of the FEAR regulatory network, which promotes Cdc14p release from the nucleolus during anaphase; potential Cdc28p substrate | Essential protein required for determination of budding pattern, promotes localization of axial markers Bud4p and Cdc12p and functionally interacts with Sec3p, localizes to the contractile ring during anaphase, member of the IQGAP family |
| 160 | 33 | 445 | 0.174 | 0.583 | 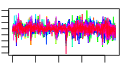 |  | 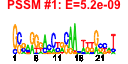 | 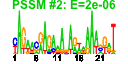 |  | YCR024C YCR036W YDR479C YIL046W YJL060W YJL070C YKL146W YKR089C YLL055W YLR324W YLR326W YML034W YNR019W YOR086C YOR267C YDR030C YDR031W YEL041W YER078C YER163C YFL021W YGR197C YGR241C YHR112C YHR199C YIL077C YIR017C YIR018W YKL037W YLR240W YLR438W YPL206C YPR081C | SLM5 RBK1 PEX29 YIL046W-A BNA3 AVT3 TGL4 YCT1 PEX30 SRC1 AGE1 TCB1 HRK1 RAD28 MIC14 YEF1 GAT1 SNG1 YAP1802 AIM46 MET28 YAP5 AIM26 VPS34 CAR2 GRS2 | Mitochondrial asparaginyl-tRNA synthetase | Putative ribokinase | Peroxisomal integral membrane peroxin, involved in the regulation of peroxisomal size, number and distribution; genetic interactions suggest that Pex28p and Pex29p act at steps upstream of those mediated by Pex30p, Pex31p, and Pex32p | Putative protein of unknown function; identified by expression profiling and mass spectrometry | Arylformamidase, involved in biosynthesis of nicotinic acid from tryptophan via kynurenine pathway; potential Cdc28p substrate | Putative protein of unknown function with similarity to AMP deaminases; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YJL070C is a non-essential gene | Vacuolar transporter, exports large neutral amino acids from the vacuole; member of a family of seven S. cerevisiae genes (AVT1-7) related to vesicular GABA-glycine transporters | Triacylglycerol lipase involved in triacylglycerol mobilization and degradation; found in lipid particles; potential Cdc28p substrate | High-affinity cysteine-specific transporter with similarity to the Dal5p family of transporters; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum; YCT1 is not an essential gene | Peroxisomal integral membrane protein, involved in negative regulation of peroxisome number; partially functionally redundant with Pex31p; genetic interactions suggest action at a step downstream of steps mediated by Pex28p and Pex29p | hypothetical protein | Inner nuclear membrane (INM) protein with a putative role in sister chromatid segregation, potentially phosphorylated by Cdc28p; contains helix-extension-helix (HEH) motif, nuclear localization signal sequence | ADP-ribosylation factor (ARF) GTPase activating protein (GAP) effector, involved in the secretory and endocytic pathways; contains C2C2H2 cysteine/histidine motif | Lipid-binding protein containing three calcium and lipid binding domains; non-tagged protein localizes to mitochondria and GFP-fusion protein localizes to the cell periphery; C-termini of Tcb1p, Tcb2p and Tcb3p interact | Protein kinase implicated in activation of the plasma membrane H(+)-ATPase Pma1p in response to glucose metabolism; plays a role in ion homeostasis | Protein involved in transcription-coupled repair nucleotide excision repair of UV-induced DNA lesions; homolog of human CSA protein | Mitochondrial intermembrane space cysteine motif protein of 14 kDa | ATP-NADH kinase; phosophorylates both NAD and NADH; homooctameric structure consisting of 60-kDa subunits; sequence similarity to Utr1p and Pos5p; overexpression complements certain pos5 phenotypes | Metallopeptidase, localized to the mitochondrial matrix | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Transcriptional activator of genes involved in nitrogen catabolite repression; contains a GATA-1-type zinc finger DNA-binding motif; activity and localization regulated by nitrogen limitation and Ure2p | Protein involved in nitrosoguanidine (MNNG) resistance; expression is regulated by transcription factors involved in multidrug resistance | Protein involved in clathrin cage assembly; binds Pan1p and clathrin; homologous to Yap1801p, member of the AP180 protein family | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO) | Transcriptional activator in the Cbf1p-Met4p-Met28p complex, participates in the regulation of sulfur metabolism | Basic leucine zipper (bZIP) transcription factor | Putative protein of unknown function; null mutant is viable and displays increased frequency of mitochondrial genome loss (petite formation) | Phosphatidylinositol 3-kinase responsible for the synthesis of phosphatidylinositol 3-phosphate; forms membrane-associated signal transduction complex with Vps15p to regulate protein sorting; activated by the GTP-bound form of Gpa1p | L-ornithine transaminase (OTAse), catalyzes the second step of arginine degradation, expression is dually-regulated by allophanate induction and a specific arginine induction process; not nitrogen catabolite repression sensitive | Protein that plays a role in metabolism of glycerophosphocholine to phosphatidylcholine; contains glycerophosphodiester phosphodiesterase motifs | Protein with sequence similarity to Grs1p, which is a glycyl-tRNA synthetase; cannot substitute for Grs1p; possible pseudogene that is expressed at very low levels |
| 249 | 36 | 437 | 0.188 | 0.555 |  |  | 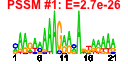 | 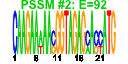 |  | YIL108W YJL165C YPL070W YBR128C YBR151W YCR036W YDL132W YDR251W YDR517W YGL047W YGR167W YIR003W YJR125C YKL007W YKL025C YKR011C YLR132C YLR133W YLR225C YLR250W YLR257W YML057W YMR152W YMR197C YMR210W YMR237W YNL173C YOL013C YOL150C YOL151W YOR018W YOR132W YOR155C YPL099C YPL260W YPR079W | HAL5 MUK1 ATG14 APD1 RBK1 CDC53 PAM1 GRH1 ALG13 CLC1 AIM21 ENT3 CAP1 PAN3 CKI1 SSP120 CMP2 YIM1 VTI1 BCH1 MDG1 HRD1 GRE2 ROD1 VPS17 ISN1 AIM43 MRL1 | Putative metalloprotease | Putative protein kinase; overexpression increases sodium and lithium tolerance, whereas gene disruption increases cation and low pH sensitivity and impairs potassium uptake, suggesting a role in regulation of Trk1p and/or Trk2p transporters | Cytoplasmic protein of unknown function; computational analysis of large-scale protein-protein interaction data suggests a possible role in transcriptional regulation | Subunit of an autophagy-specific phosphatidylinositol 3-kinase complex (with Vps34p, Vps15p, and Vps30p) required for organization of a pre-autophagosomal structure; ATG14 transcription is activated by Gln3p during nitrogen starvation | Protein of unknown function, required for normal localization of actin patches and for normal tolerance of sodium ions and hydrogen peroxide; localizes to both cytoplasm and nucleus | Putative ribokinase | Cullin, structural protein of SCF complexes (which also contain Skp1p, Cdc34p, Hrt1p and an F-box protein) involved in ubiquitination; SCF promotes the G1-S transition by targeting G1 cyclins and the Cln-CDK inhibitor Sic1p for degradation | Essential protein of unknown function; exhibits variable expression during colony morphogenesis; overexpression permits survival without protein phosphatase 2A, inhibits growth, and induces a filamentous phenotype | Acetylated, cis-golgi localized protein involved in ER to Golgi transport; homolog of human GRASP65; forms a complex with the coiled-coil protein Bug1p; mutants are compromised for the fusion of ER-derived vesicles with Golgi membranes | Catalytic component of UDP-GlcNAc transferase, required for the second step of dolichyl-linked oligosaccharide synthesis; anchored to the ER membrane via interaction with Alg14p; similar to bacterial and human glycosyltransferases | Clathrin light chain, subunit of the major coat protein involved in intracellular protein transport and endocytosis; thought to regulate clathrin function, two Clathrin heavy chains (CHC1) form the clathrin triskelion structural component | Protein of unknown function; may interact with ribosomes; GFP-fusion protein colocalizes with Sac1p to the actin cytoskeleton; null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Protein containing an N-terminal epsin-like domain involved in clathrin recruitment and traffic between the Golgi and endosomes; associates with the clathrin adaptor Gga2p | Alpha subunit of the capping protein (CP) heterodimer (Cap1p and Cap2p) which binds to the barbed ends of actin filaments preventing further polymerization; localized predominantly to cortical actin patches | Essential subunit of the Pan2p-Pan3p poly(A)-ribonuclease complex, which acts to control poly(A) tail length and regulate the stoichiometry and activity of postreplication repair complexes | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus | Protein of unknown function; green fluorescent protein (GFP)-tagged protein localizes to mitochondria and nucleus; YLR132C is an essential gene | Choline kinase, catalyzing the first step in phosphatidylcholine synthesis via the CDP-choline (Kennedy pathway); exhibits some ethanolamine kinase activity contributing to phosphatidylethanolamine synthesis via the CDP-ethanolamine pathway | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR225C is not an essential gene | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Putative protein of unknown function | Calcineurin A; one isoform (the other is CNA1) of the catalytic subunit of calcineurin, a Ca++/calmodulin-regulated protein phosphatase which regulates Crz1p (a stress-response transcription factor), the other calcineurin subunit is CNB1 | Protein of unknown function; null mutant displays sensitivity to DNA damaging agents; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein involved in cis-Golgi membrane traffic; v-SNARE that interacts with two t-SNARES, Sed5p and Pep12p; required for multiple vacuolar sorting pathways | Putative acyltransferase with similarity to Eeb1p and Eht1p, has a minor role in medium-chain fatty acid ethyl ester biosynthesis; may be involved in lipid metabolism and detoxification | Member of the ChAPs family (Chs5p-Arf1p-binding proteins: Bch1p, Bch2p, Bud7p, Chs6p), that forms the exomer complex with Chs5p to mediate export of specific cargo proteins from the Golgi to the plasma membrane; may interact with ribosomes | Plasma membrane protein involved in G-protein mediated pheromone signaling pathway; overproduction suppresses bem1 mutations | Ubiquitin-protein ligase required for endoplasmic reticulum-associated degradation (ERAD) of misfolded proteins; genetically linked to the unfolded protein response (UPR); regulated through association with Hrd3p; contains an H2 ring finger | NADPH-dependent methylglyoxal reductase (D-lactaldehyde dehydrogenase); stress induced (osmotic, ionic, oxidative, heat shock and heavy metals); regulated by the HOG pathway | Membrane protein; overexpression confers resistance to the GST substrate o-dinitrobenzene as well as to zinc and calcium; contains 2 PY motifs, which are required for Rod1p interaction with Rsp5p, a hect-type ubiquitin ligase | Subunit of the membrane-associated retromer complex essential for endosome-to-Golgi retrograde protein transport; peripheral membrane protein that assembles onto the membrane with Vps5p to promote vesicle formation | Inosine 5'-monophosphate (IMP)-specific 5'-nucleotidase, catalyzes the breakdown of IMP to inosine, does not show similarity to known 5'-nucleotidases from other organisms | Protein of unknown function;non-tagged protein detected in purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss and severe growth defect in minimal glycerol media | Putative substrate of cAMP-dependent protein kinase (PKA); green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YPL260W is not an essential gene | Membrane protein with similarity to mammalian mannose-6-phosphate receptors, possibly functions as a sorting receptor in the delivery of vacuolar hydrolases |
| 183 | 46 | 443 | 0.213 | 0.536 |  |  | 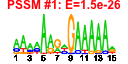 | 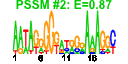 |  | YBR086C YBR127C YCR012W YDL145C YDR303C YGR240C YIL039W YIL109C YJL002C YJL091C YKL080W YLR088W YLR100W YLR195C YLR208W YLR328W YLR378C YMR203W YNL154C YOL020W YOL057W YOR002W YOR067C YOR085W YOR153W YPL145C YPL246C YBR263W YCR084C YGL245W YGL253W YHR098C YHR144C YJL014W YJL133W YJL192C YKL164C YMR130W YMR205C YNL219C YNR021W YOR099W YOR311C YPL245W YPR113W YPR114W | IST2 VMA2 PGK1 COP1 RSC3 PFK1 TED1 SEC24 OST1 GWT1 CBF2 GAA1 ERG27 NMT1 SEC13 NMA1 SEC61 TOM40 YCK2 TAT2 ALG6 ALG8 OST3 NCB2 KES1 RBD2 SHM1 TUP1 GUS1 HXK2 SFB3 DCD1 CCT3 MRS3 SOP4 PIR1 PFK2 ALG9 KTR1 HSD1 PIS1 | Plasma membrane protein that may be involved in osmotolerance, localizes to the mother cell in small-budded cells and to the bud in medium- and large-budded cells; mRNA is transported to the bud tip by an actomyosin-driven process | Subunit B of the eight-subunit V1 peripheral membrane domain of the vacuolar H+-ATPase (V-ATPase), an electrogenic proton pump found throughout the endomembrane system; contains nucleotide binding sites; also detected in the cytoplasm | 3-phosphoglycerate kinase, catalyzes transfer of high-energy phosphoryl groups from the acyl phosphate of 1,3-bisphosphoglycerate to ADP to produce ATP; key enzyme in glycolysis and gluconeogenesis | Alpha subunit of COPI vesicle coatomer complex, which surrounds transport vesicles in the early secretory pathway | Component of the RSC chromatin remodeling complex; essential gene required for maintenance of proper ploidy and regulation of ribosomal protein genes and the cell wall/stress response; highly similar to Rsc30p | Alpha subunit of heterooctameric phosphofructokinase involved in glycolysis, indispensable for anaerobic growth, activated by fructose-2,6-bisphosphate and AMP, mutation inhibits glucose induction of cell cycle-related genes | Conserved phosphoesterase domain-containing protein that acts together with Emp24p/Erv25p in cargo exit from the ER; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO) | Component of the Sec23p-Sec24p heterodimeric complex of the COPII vesicle coat; involved in ER to Golgi transport, cargo selection and autophagy; required for the binding of the Sec13 complex to ER membranes; homologous to Lst1p and Lss1p | Alpha subunit of the oligosaccharyltransferase complex of the ER lumen, which catalyzes asparagine-linked glycosylation of newly synthesized proteins | Protein involved in the inositol acylation of glucosaminyl phosphatidylinositol (GlcN-PI) to form glucosaminyl(acyl)phosphatidylinositol (GlcN(acyl)PI), an intermediate in the biosynthesis of glycosylphosphatidylinositol (GPI) anchors | Essential kinetochore protein, component of the CBF3 multisubunit complex that binds to the CDEIII region of the centromere; Cbf2p also binds to the CDEII region possibly forming a different multimeric complex, ubiquitinated in vivo | Subunit of the GPI (glycosylphosphatidylinositol):protein transamidase complex, removes the GPI-anchoring signal and attaches GPI to proteins in the ER | 3-keto sterol reductase, catalyzes the last of three steps required to remove two C-4 methyl groups from an intermediate in ergosterol biosynthesis; mutants are sterol auxotrophs | N-myristoyl transferase, catalyzes the cotranslational, covalent attachment of myristic acid to the N-terminal glycine residue of several proteins involved in cellular growth and signal transduction | Component of both the Nup84 nuclear pore sub-complex and of the COPII complex (Sar1p, Sec13p, Sec16p, Sec23p, Sec24p, Sec31p, Sfb2p, and Sfb3p) which is important for the formation of ER to Golgi transport vesicles | Nicotinic acid mononucleotide adenylyltransferase, involved in NAD(+) salvage pathway | Essential subunit of Sec61 complex (Sec61p, Sbh1p, and Sss1p); forms a channel for SRP-dependent protein import and retrograde transport of misfolded proteins out of the ER; with Sec63 complex allows SRP-independent protein import into ER | Component of the TOM (translocase of outer membrane) complex responsible for recognition and initial import steps for all mitochondrially directed proteins; constitutes the core element of the protein conducting pore | Palmitoylated, plasma membrane-bound casein kinase I isoform; shares redundant functions with Yck1p in morphogenesis, proper septin assembly, endocytic trafficking; provides an essential function overlapping with that of Yck1p | High affinity tryptophan and tyrosine permease, overexpression confers FK506 and FTY720 resistance | Putative metalloprotease | Alpha 1,3 glucosyltransferase, involved in transfer of oligosaccharides from dolichyl pyrophosphate to asparagine residues of proteins during N-linked protein glycosylation; mutations in human ortholog are associated with disease | Glucosyl transferase, involved in N-linked glycosylation; adds glucose to the dolichol-linked oligosaccharide precursor prior to transfer to protein during lipid-linked oligosaccharide biosynthesis; similar to Alg6p | Gamma subunit of the oligosaccharyltransferase complex of the ER lumen, which catalyzes asparagine-linked glycosylation of newly synthesized proteins; Ost3p is important for N-glycosylation of a subset of proteins | Subunit of a heterodimeric NC2 transcription regulator complex with Bur6p; complex binds to TBP and can repress transcription by preventing preinitiation complex assembly or stimulate activated transcription; homologous to human NC2beta | Member of the oxysterol binding protein family, which includes seven yeast homologs; involved in negative regulation of Sec14p-dependent Golgi complex secretory functions, peripheral membrane protein that localizes to the Golgi complex | Possible rhomboid protease, has similarity to eukaryotic rhomboid proteases including Pcp1p | Mitochondrial serine hydroxymethyltransferase, converts serine to glycine plus 5,10 methylenetetrahydrofolate; involved in generating precursors for purine, pyrimidine, amino acid, and lipid biosynthesis; reverse reaction generates serine | General repressor of transcription, forms complex with Cyc8p, involved in the establishment of repressive chromatin structure through interactions with histones H3 and H4, appears to enhance expression of some genes | Glutamyl-tRNA synthetase (GluRS), forms a complex with methionyl-tRNA synthetase (Mes1p) and Arc1p; complex formation increases the catalytic efficiency of both tRNA synthetases and ensures their correct localization to the cytoplasm | Hexokinase isoenzyme 2 that catalyzes phosphorylation of glucose in the cytosol; predominant hexokinase during growth on glucose; functions in the nucleus to repress expression of HXK1 and GLK1 and to induce expression of its own gene | Member of the Sec24p family; forms a complex, with Sec23p, that is involved in sorting of Pma1p into COPII vesicles; peripheral ER membrane protein; potential Cdc28p substrate | Deoxycytidine monophosphate (dCMP) deaminase required for dCTP and dTTP synthesis; expression is NOT cell cycle regulated | Subunit of the cytosolic chaperonin Cct ring complex, related to Tcp1p, required for the assembly of actin and tubulins in vivo | Mitochondrial iron transporter of the mitochondrial carrier family (MCF), very similar to and functionally redundant with Mrs4p; functions under low-iron conditions; may transport other cations in addition to iron | ER-membrane protein; suppressor of pma1-7, deletion of SOP4 slows down the export of wild-type Pma1p and Pma1-7 from the ER | O-glycosylated protein required for cell wall stability; attached to the cell wall via beta-1,3-glucan; mediates mitochondrial translocation of Apn1p; expression regulated by the cell integrity pathway and by Swi5p during the cell cycle | Putative protein of unknown function; YMR130W is not an essential gene | Beta subunit of heterooctameric phosphofructokinase involved in glycolysis, indispensable for anaerobic growth, activated by fructose-2,6-bisphosphate and AMP, mutation inhibits glucose induction of cell cycle-related genes | Mannosyltransferase, involved in N-linked glycosylation; catalyzes the transfer of mannose from Dol-P-Man to lipid-linked oligosaccharides; mutation of the human ortholog causes type 1 congenital disorders of glycosylation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum; YNR021W is not an essential gene | Alpha-1,2-mannosyltransferase involved in O- and N-linked protein glycosylation; type II membrane protein; member of the KRE2/MNT1 mannosyltransferase family | Endoplasmic reticulum (ER)-resident membrane protein, overproduction induces enlargement of ER-like membrane structures and suppresses a temperature-sensitive sly1 mutation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the nucleus and the cytoplasm | Phosphatidylinositol synthase, required for biosynthesis of phosphatidylinositol, which is a precursor for polyphosphoinositides, sphingolipids, and glycolipid anchors for some of the plasma membrane proteins | Putative protein of unknown function |
| 133 | 43 | 431 | 0.226 | 0.566 | 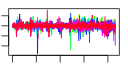 |  | 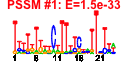 | 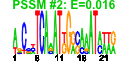 |  | YAL028W YBR201W YCR082W YCR083W YDL159W YDR336W YDR383C YDR425W YDR468C YDR486C YEL057C YER026C YER027C YER045C YER046W YER050C YER065C YER078C YER080W YFR036W YFR042W YGL017W YGR154C YGR247W YHL016C YHL038C YHR004C YHR076W YJL053W YJR086W YJR115W YLR118C YOR258W YDR319C YDR323C YER051W YER087W YFR041C YKL200C YKL201C YLR119W YLR241W YOR003W | FRT2 DER1 AHC2 TRX3 YDL159W-A NKP1 SNX41 TLG1 VPS60 CHO1 GAL83 ACA1 SPO73 RSM18 ICL1 AIM9 CDC26 KEG1 ATE1 GTO1 CPD1 DUR3 CBP2 NEM1 PTC7 PEP8 STE18 HNT3 YFT2 PEP7 JHD1 AIM10 ERJ5 MNN4 SRN2 YSP3 | Tail-anchored endoplasmic reticulum membrane protein, interacts with homolog Frt1p but is not a substrate of calcineurin (unlike Frt1p), promotes growth in conditions of high Na+, alkaline pH, or cell wall stress; potential Cdc28p substrate | Endoplasmic reticulum membrane protein, required for ER-associated protein degradation of misfolded or unassembled proteins; N- and C- termini protrude into the cytoplasm, has similarity to Dfm1p | Protein of unknown function, putative transcriptional regulator; proposed to be a Ada Histone acetyltransferase complex component; GFP tagged protein is localized to the cytoplasm and nucleus | Mitochondrial thioredoxin, highly conserved oxidoreductase required to maintain the redox homeostasis of the cell, forms the mitochondrial thioredoxin system with Trr2p, redox state is maintained by both Trr2p and Glr1p | Putative protein of unknown function; identified by sequence comparison with hemiascomycetous yeast species | Putative protein of unknown function; sumoylated under stress conditions in a genome wide study; YDR336W is not an essential gene | Non-essential kinetochore protein, subunit of the Ctf19 central kinetochore complex (Ctf19p-Mcm21p-Okp1p-Mcm22p-Mcm16p-Ctf3p-Chl4p-Mcm19p- Nkp1p-Nkp2p-Ame1p-Mtw1p) | Sorting nexin, involved in the retrieval of late-Golgi SNAREs from the post-Golgi endosome to the trans-Golgi network; forms a complex with Snx4p and Atg20p | Essential t-SNARE that forms a complex with Tlg2p and Vti1p and mediates fusion of endosome-derived vesicles with the late Golgi; binds the docking complex VFT (Vps fifty-three) through interaction with Vps51p | Cytoplasmic and vacuolar membrane protein involved in late endosome to vacuole transport; required for normal filament maturation during pseudohyphal growth; may function in targeting specific cargo proteins for degradation | Protein of unknown function involved in telomere maintenance; target of UME6 regulation | Phosphatidylserine synthase, functions in phospholipid biosynthesis; catalyzes the reaction CDP-diaclyglycerol + L-serine = CMP + L-1-phosphatidylserine, transcriptionally repressed by myo-inositol and choline | One of three possible beta-subunits of the Snf1 kinase complex, allows nuclear localization of the Snf1 kinase complex in the presence of a nonfermentable carbon source; contains glycogen-binding domain | Basic leucine zipper (bZIP) transcription factor of the ATF/CREB family, may regulate transcription of genes involved in utilization of non-optimal carbon sources | Meiosis-specific protein of unknown function, required for spore wall formation during sporulation; dispensible for both nuclear divisions during meiosis | Mitochondrial ribosomal protein of the small subunit, has similarity to E. coli S18 ribosomal protein | Isocitrate lyase, catalyzes the formation of succinate and glyoxylate from isocitrate, a key reaction of the glyoxylate cycle; expression of ICL1 is induced by growth on ethanol and repressed by growth on glucose | Metallopeptidase, localized to the mitochondrial matrix | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) | Subunit of the Anaphase-Promoting Complex/Cyclosome (APC/C), which is a ubiquitin-protein ligase required for degradation of anaphase inhibitors, including mitotic cyclins, during the metaphase/anaphase transition | Protein of unknown function required for cell viability; localizes to the endoplasmic reticulum membrane | Arginyl-tRNA-protein transferase, catalyzes post-translational conjugation of arginine to the amino termini of acceptor proteins which are then subject to degradation via the N-end rule pathway | Omega-class glutathione transferase; induced under oxidative stress; putative peroxisomal localization | Cyclic nucleotide phosphodiesterase, hydrolyzes ADP-ribose 1'', 2''-cyclic phosphate to ADP-ribose 1''-phosphate; may have a role in tRNA splicing; no detectable phenotype is conferred by null mutation or by overexpression | Plasma membrane transporter for both urea and polyamines, expression is highly sensitive to nitrogen catabolite repression and induced by allophanate, the last intermediate of the allantoin degradative pathway | Mitochondrial protein required for splicing of the group I intron aI5 of the COB pre-mRNA, binds to the RNA to promote splicing; also involved in but not essential for splicing of the COB bI2 intron and the intron in the 21S rRNA gene | Catalytic subunit of Nem1p-Spo7p phosphatase holoenzyme, which regulates nuclear growth by controlling recruitment of Pah1p onto promoters of phospholipid biosynthetic genes; required for normal nuclear envelope morphology and sporulation | Mitochondrially localized type 2C protein phosphatase; expression induced by growth on ethanol and by sustained osmotic stress; possible role in carbon source utilization in low oxygen environments | Vacuolar protein sorting protein that forms part of the multimeric membrane-associated retromer complex along with Vps35p, Vps29p, Vps17p, and Vps5p; essential for endosome-to-Golgi retrograde protein transport | G protein gamma subunit, forms a dimer with Ste4p to activate the mating signaling pathway, forms a heterotrimer with Gpa1p and Ste4p to dampen signaling; C-terminus is palmitoylated and farnesylated, which are required for normal signaling | Putative protein of unknown function | Acyl-protein thioesterase responsible for depalmitoylation of Gpa1p; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and nucleus and is induced in response to the DNA-damaging agent MMS | Member of the third branch of the histidine triad (HIT) superfamily of nucleotide-binding proteins; similar to Aprataxin, a Hint related protein that is mutated in individuals with ataxia with oculomotor apraxia | Putative protein of unknown function, identified as an ortholog of the highly conserved FIT family of proteins involved in triglyceride droplet biosynthesis; interacts with Sst2p and Hsp82p in high-throughput two-hybrid screens | Multivalent adaptor protein that facilitates vesicle-mediated vacuolar protein sorting by ensuring high-fidelity vesicle docking and fusion, which are essential for targeting of vesicles to the endosome; required for vacuole inheritance | JmjC domain family histone demethylase specific for H3-K36, similar to proteins found in human, mouse, drosophila, X. laevis, C. elegans, and S. pombe | Protein with similarity to tRNA synthetases; non-tagged protein is detected in purified mitochondria; null mutant displays decreased frequency of mitochondrial genome loss (petite formation), severe growth defect in minimal glycerol media | Type I membrane protein with a J domain is required to preserve the folding capacity of the endoplasmic reticulum; loss of the non-essential ERJ5 gene leads to a constitutively induced unfolded protein response | Putative positive regulator of mannosylphosphate transferase (Mnn6p), involved in mannosylphosphorylation of N-linked oligosaccharides; expression increases in late-logarithmic and stationary growth phases | Component of the ESCRT-I complex, which is involved in ubiquitin-dependent sorting of proteins into the endosome; suppressor of rna1-1 mutation; may be involved in RNA export from nucleus | Putative protein of unknown function, may be involved in detoxification | Putative precursor to the subtilisin-like protease III |
| 6 | 44 | 438 | 0.227 | 0.530 | 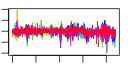 |  | 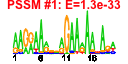 | 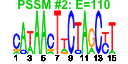 |  | YDR176W YDR320C YGL114W YGL178W YHR151C YIL068C YJR062C YJR110W YKL134C YKR051W YLR072W YLR263W YLR403W YML102W YML122C YMR063W YMR075W YMR102C YMR124W YMR219W YNL105W YNL127W YNL139C YNR008W YOL024W YOL035C YOL037C YOL066C YOL138C YOR062C YOR068C YOR093C YOR156C YOR160W YOR371C YPL039W YPL040C YPL136W YPL148C YPL194W YPR039W YPR177C YPR189W YOL075C | NGG1 SWA2 MTH1 SEC6 NTA1 YMR1 OCT1 RED1 SFP1 CAC2 RIM9 RCO1 ESC1 FAR11 RLR1 LRO1 RIB2 VAM10 NFI1 MTR10 GPB1 ISM1 PPT2 DDC1 SKI3 | Transcriptional regulator involved in glucose repression of Gal4p-regulated genes; component of transcriptional adaptor and histone acetyltransferase complexes, the ADA complex, the SAGA complex, and the SLIK complex | Auxilin-like protein involved in vesicular transport; clathrin-binding protein required for uncoating of clathrin-coated vesicles | Putative protein of unknown function; predicted member of the oligopeptide transporter (OPT) family of membrane transporters | Negative regulator of the glucose-sensing signal transduction pathway, required for repression of transcription by Rgt1p; interacts with Rgt1p and the Snf3p and Rgt2p glucose sensors; phosphorylated by Yck1p, triggering Mth1p degradation | Putative protein of unknown function | Essential 88kDa subunit of the exocyst complex, which mediates polarized targeting of secretory vesicles to active sites of exocytosis; dimeric form of Sec6p interacts with Sec9p in vitro and inhibits t-SNARE assembly | Amidase, removes the amide group from N-terminal asparagine and glutamine residues to generate proteins with N-terminal aspartate and glutamate residues that are targets of ubiquitin-mediated degradation | Phosphatidylinositol 3-phosphate [PI(3)P] phosphatase, regulates the localization and levels of PI(3)P; involved in cytoplasm to vacuole (CVT) transport; has similarity to the conserved myotubularin dual specificity phosphatase family | Mitochondrial intermediate peptidase, cleaves N-terminal residues of a subset of proteins upon import, after their cleavage by mitochondrial processing peptidase (Mas1p-Mas2p); may contribute to mitochondrial iron homeostasis | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern; YLR072W is not an esssential gene | Protein component of the axial elements of the synaptonemal complex, involved in chromosome segregation during the first meiotic division; interacts with Hop1p; required for wild-type levels of Mek1p kinase activity | Transcription factor that controls expression of many ribosome biogenesis genes in response to nutrients and stress, regulates G2/M transitions during mitotic cell cycle and DNA-damage response, involved in cell size modulation | Component of the chromatin assembly complex (with Rlf2p and Msi1p) that assembles newly synthesized histones onto recently replicated DNA, required for building functional kinetochores, conserved from yeast to humans | Protein of unknown function, involved in the proteolytic activation of Rim101p in response to alkaline pH; has similarity to A. nidulans PalI; putative membrane protein | Essential subunit of the histone deacetylase Rpd3S complex; interacts with Eaf3p | Protein of unknown function; transcription is activated by paralogous transcription factors Yrm1p and Yrr1p along with genes involved in multidrug resistance; mutant shows increased resistance to azoles; YMR102C is not an essential gene | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery, cytoplasm, bud, and bud neck; interacts with Crm1p in two-hybrid assay; YMR124W is not an essential gene | Protein localized to the nuclear periphery, involved in telomeric silencing; interacts with PAD4-domain of Sir4p | Protein involved in G1 cell cycle arrest in response to pheromone, in a pathway different from the Far1p-dependent pathway; interacts with Far3p, Far7p, Far8p, Far9p, and Far10p | Subunit of the THO complex, which is required for efficient transcription elongation and involved in transcriptional elongation-associated recombination; required for LacZ RNA expression from certain plasmids | Acyltransferase that catalyzes diacylglycerol esterification; one of several acyltransferases that contribute to triglyceride synthesis; putative homolog of human lecithin cholesterol acyltransferase | Putative protein of unknown function, predicted to have thiol-disulfide oxidoreductase active site | DRAP deaminase, catalyzes the third step of the riboflavin biosynthesis pathway; cytoplasmic tRNA pseudouridine synthase involved in pseudouridylation of cytoplasmic tRNAs at position 32 | Putative protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Protein of unknown function; similar to YKR075Cp and Reg1p; expression regulated by glucose and Rgt1p; GFP-fusion protein is induced in response to the DNA-damaging agent MMS | Protein involved in vacuole morphogenesis; acts at an early step of homotypic vacuole fusion that is required for vacuole tethering | Putative protein of unknown function; deletion causes sensitivity to unfolded protein response-inducing agents | SUMO ligase, catalyzes the covalent attachment of SUMO (Smt3p) to proteins; involved in maintenance of proper telomere length | Nuclear import receptor, mediates the nuclear localization of proteins involved in mRNA-nucleus export; promotes dissociation of mRNAs from the nucleus-cytoplasm mRNA shuttling protein Npl3p | Multistep regulator of cAMP-PKA signaling; inhibits PKA downstream of Gpa2p and Cyr1p, thereby increasing cAMP dependency; inhibits Ras activity through direct interactions with Ira1p/2p; regulated by G-alpha protein Gpa2p; homolog of Gpb2p | Putative protein of unknown function; YPL039W is not an essential gene | Mitochondrial isoleucyl-tRNA synthetase, null mutant is deficient in respiratory growth | Phosphopantetheine:protein transferase (PPTase), activates mitochondrial acyl carrier protein (Acp1p) by phosphopantetheinylation | DNA damage checkpoint protein, part of a PCNA-like complex required for DNA damage response, required for pachytene checkpoint to inhibit cell cycle in response to unrepaired recombination intermediates; potential Cdc28p substrate | Protein involved in exosome mediated 3' to 5' mRNA degradation and translation inhibition of non-poly(A) mRNAs; forms complex with Ski2p and Ski8p; required for repressing propagation of dsRNA viruses | Putative ABC transporter |
| 272 | 38 | 429 | 0.257 | 0.547 |  |  | 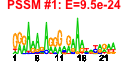 | 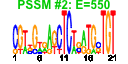 |  | YCL033C YHL032C YIR036C YIR037W YJL151C YJL152W YOR161C YAL008W YBR016W YBR024W YBR111C YCL017C YCR069W YCR076C YDL057W YDL078C YER083C YGL127C YGL165C YGL198W YGL199C YGR086C YGR137W YHL031C YHR057C YIL023C YIL108W YJL153C YJR074W YJR103W YJR117W YKR048C YLR121C YLR350W YNL322C YOR089C YOR230W YOR347C | GUT1 IRC24 HYR1 SNA3 PNS1 FUN14 SCO2 YSA1 NFS1 CPR4 MDH3 GET2 SOH1 YIP4 PIL1 GOS1 CPR2 YKE4 INO1 MOG1 URA8 STE24 NAP1 YPS3 ORM2 KRE1 VPS21 WTM1 PYK2 | Putative protein-methionine-R-oxide reductase; involved in response to oxidative stress; similar to mouse Sepx1p and fly SelRp; YCL033C is not an essential gene | Glycerol kinase, converts glycerol to glycerol-3-phosphate; glucose repression of expression is mediated by Adr1p and Ino2p-Ino4p; derepression of expression on non-fermentable carbon sources is mediated by Opi1p and Rsf1p | Putative benzil reductase;(GFP)-fusion protein localizes to the cytoplasm and is induced by the DNA-damaging agent MMS; sequence similarity with short-chain dehydrogenase/reductases; null mutant has increased spontaneous Rad52 foci | Thiol peroxidase that functions as a hydroperoxide receptor to sense intracellular hydroperoxide levels and transduce a redox signal to the Yap1p transcription factor | Integral membrane protein localized to vacuolar intralumenal vesicles, computational analysis of large-scale protein-protein interaction data suggests a possible role in either cell wall synthesis or protein-vacuolar targeting | Protein of unknown function; has similarity to Torpedo californica tCTL1p, which is postulated to be a choline transporter, neither null mutation nor overexpression affects choline transport | Mitochondrial protein of unknown function | Plasma membrane protein of unknown function; has similarity to hydrophilins, which are hydrophilic, glycine-rich proteins involved in the adaptive response to hyperosmotic conditions | Protein anchored to the mitochondrial inner membrane, similar to Sco1p and may have a redundant function with Sco1p in delivery of copper to cytochrome c oxidase; interacts with Cox2p | Nudix hydrolase family member with ADP-ribose pyrophosphatase activity | Cysteine desulfurase involved in iron-sulfur cluster (Fe/S) biogenesis; required for the post-transcriptional thio-modification of mitochondrial and cytoplasmic tRNAs; essential protein located predominantly in mitochondria | Peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; has a potential role in the secretory pathway | Putative protein of unknown function; YCR076C is not an essential gene | Putative protein of unknown function; YDL057W is not an essential gene | Peroxisomal malate dehydrogenase, catalyzes interconversion of malate and oxaloacetate; involved in the glyoxylate cycle | Subunit of the GET complex; required for meiotic nuclear division and for the retrieval of HDEL proteins from the Golgi to the ER in an ERD2 dependent fashion; may be involved in cell wall function | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; involved in telomere maintenance; conserved with other metazoan MED31 subunits | Protein that interacts with Rab GTPases, localized to late Golgi vesicles; computational analysis of large-scale protein-protein interaction data suggests a possible role in vesicle-mediated transport | Primary component of eisosomes, which are large immobile cell cortex structures associated with endocytosis; null mutants show activation of Pkc1p/Ypk1p stress resistance pathways; detected in phosphorylated state in mitochondria | v-SNARE protein involved in Golgi transport, homolog of the mammalian protein GOS-28/GS28 | Zinc transporter; localizes to the ER; null mutant is sensitive to calcofluor white, leads to zinc accumulation in cytosol; ortholog of the mouse KE4 and member of the ZIP (ZRT, IRT-like Protein) family | Putative metalloprotease | Inositol 1-phosphate synthase, involved in synthesis of inositol phosphates and inositol-containing phospholipids; transcription is coregulated with other phospholipid biosynthetic genes by Ino2p and Ino4p, which bind the UASINO DNA element | Conserved nuclear protein that interacts with GTP-Gsp1p, which is a Ran homolog of the Ras GTPase family, and stimulates nucleotide release, involved in nuclear protein import, nucleotide release is inhibited by Yrb1p | Minor CTP synthase isozyme (see also URA7), catalyzes the ATP-dependent transfer of the amide nitrogen from glutamine to UTP, forming CTP, the final step in de novo biosynthesis of pyrimidines; involved in phospholipid biosynthesis | Highly conserved zinc metalloprotease that functions in two steps of a-factor maturation, C-terminal CAAX proteolysis and the first step of N-terminal proteolytic processing; contains multiple transmembrane spans | Protein that interacts with mitotic cyclin Clb2p; required for the regulation of microtubule dynamics during mitosis; controls bud morphogenesis; involved in the transport of H2A and H2B histones to the nucleus | Aspartic protease, attached to the plasma membrane via a glycosylphosphatidylinositol (GPI) anchor | Evolutionarily conserved protein with similarity to Orm1p, required for resistance to agents that induce the unfolded protein response; human ortholog is located in the endoplasmic reticulum | Cell wall glycoprotein involved in beta-glucan assembly; serves as a K1 killer toxin membrane receptor | GTPase required for transport during endocytosis and for correct sorting of vacuolar hydrolases; localized in endocytic intermediates; detected in mitochondria; geranylgeranylation required for membrane association; mammalian Rab5 homolog | Transcriptional repressor involved in regulation of meiosis and silencing, required for nuclear localization of the ribonucleotide reductase small subunit Rnr2p and Rnr4p; contains WD repeats | Pyruvate kinase that appears to be modulated by phosphorylation; PYK2 transcription is repressed by glucose, and Pyk2p may be active under low glycolytic flux |
| 241 | 31 | 449 | 0.263 | 0.558 |  |  | 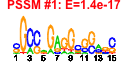 | 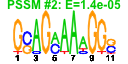 |  | YAL049C YDL009C YDL010W YDL057W YGL165C YPL135W YBL095W YBR005W YDR380W YGL166W YIL051C YJL185C YJR059W YKL023W YLR155C YLR157C YLR158C YLR160C YLR205C YMR184W YMR261C YMR262W YNL036W YNL055C YNL157W YOL110W YOL111C YPL015C YPL031C YPR023C YPR075C | AIM2 GRX6 ISU1 RCR1 ARO10 CUP2 MMF1 PTK2 ASP3-1 HMX1 ADD37 TPS3 NCE103 POR1 IGO1 SHR5 MDY2 HST2 PHO85 EAF3 OPY2 | Cytoplasmic protein of unknown function, potential Hsp82p interactor; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) | Monothiol glutaredoxin that binds an iron-sulfur cluster; more similar in activity to dithiol than other monothiol glutaredoxins; forms homodimers; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Putative protein of unknown function; YDL057W is not an essential gene | Conserved protein of the mitochondrial matrix, performs a scaffolding function during assembly of iron-sulfur clusters, interacts physically and functionally with yeast frataxin (Yfh1p); isu1 isu2 double mutant is inviable | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein of the ER membrane involved in cell wall chitin deposition; may function in the endosomal-vacuolar trafficking pathway, helping determine whether plasma membrane proteins are degraded or routed to the plasma membrane | Phenylpyruvate decarboxylase, catalyzes decarboxylation of phenylpyruvate to phenylacetaldehyde, which is the first specific step in the Ehrlich pathway | Copper-binding transcription factor; activates transcription of the metallothionein genes CUP1-1 and CUP1-2 in response to elevated copper concentrations | Mitochondrial protein involved in maintenance of the mitochondrial genome | Putative protein of unknown function; mRNA is weakly cell cycle regulated, peaking in G2 phase; YJL185C is a non-essential gene | Putative serine/threonine protein kinase involved in regulation of ion transport across plasma membrane; enhances spermine uptake | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Cell-wall L-asparaginase II, involved in asparagine catabolism; expression is induced during nitrogen starvation; four copies of ASP3 are present in the genome reference strain S288C | ER localized, heme-binding peroxidase involved in the degradation of heme; does not exhibit heme oxygenase activity despite similarity to heme oxygenases; expression regulated by AFT1 | Protein of unknown function involved in ER-associated protein degradation; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and is induced in response to the DNA-damaging agent MMS; YMR184W is not an essential gene | Regulatory subunit of trehalose-6-phosphate synthase/phosphatase complex, which synthesizes the storage carbohydrate trehalose; expression is induced by stress conditions and repressed by the Ras-cAMP pathway | Protein of unknown function; interacts weakly with Knr4p; YMR262W is not an essential gene | Carbonic anhydrase; poorly transcribed under aerobic conditions and at an undetectable level under anaerobic conditions; involved in non-classical protein export pathway | Mitochondrial porin (voltage-dependent anion channel), outer membrane protein required for the maintenance of mitochondrial osmotic stability and mitochondrial membrane permeability; phosphorylated | hypothetical protein | Subunit of a palmitoyltransferase, composed of Shr5p and Erf2p, that adds a palmitoyl lipid moiety to heterolipidated substrates such as Ras1p and Ras2p through a thioester linkage; palmitoylation is required for Ras2p membrane localization | Protein required for efficient mating; involved in shmoo formation and nuclear migration in the pre-zygote; associates with ribosomes and interacts with YOR164C; contains a ubiquitin-like (UBL) domain | Cytoplasmic member of the silencing information regulator 2 (Sir2) family of NAD(+)-dependent protein deacetylases; modulates nucleolar (rDNA) and telomeric silencing; possesses NAD(+)-dependent histone deacetylase activity in vitro | Cyclin-dependent kinase, with ten cyclin partners; involved in environmental stress response; in phosphate-rich conditions, Pho85p-Pho80p complex phosphorylates Pho4p which in turn represses PHO5 | Esa1p-associated factor, nonessential component of the NuA4 acetyltransferase complex, homologous to Drosophila dosage compensation protein MSL3 | Integral membrane protein that functions in the signaling branch of the high-osmolarity glycerol (HOG) pathway; interacts with Ste50p; overproduction blocks cell cycle arrest in the presence of mating pheromone |
| 230 | 51 | 450 | 0.291 | 0.565 | 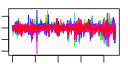 |  | 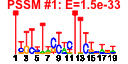 | 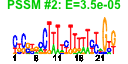 |  | YGR099W YHR120W YHR205W YIR033W YLR442C YNL198C YNR065C YOR237W YPL189W YAL033W YCR047C YCR101C YCRX20C YDL041W YDL042C YDL240W YEL019C YER176W YGL094C YGR166W YGR198W YGR286C YHR118C YHR207C YIL129C YIL147C YIL149C YIL159W YIR004W YIR030C YLL004W YLL038C YLR018C YLR135W YLR139C YLR141W YLR223C YLR341W YLR343W YLR364W YNL210W YNL251C YNR029C YNR048W YNR066C YOL091W YOR129C YOR333C YPL007C YPL157W YPR089W | TEL2 MSH1 SCH9 MGA2 SIR3 HES1 GUP2 POP5 BUD23 SIR2 LRG1 MMS21 ECM32 PAN2 KRE11 YPP1 BIO2 ORC6 SET5 TAO3 SLN1 MLP2 BNR1 DJP1 DCG1 ORC3 ENT4 POM34 SLX4 SLS1 RRN5 IFH1 SPO77 GAS2 MER1 NRD1 CRF1 SPO21 TFC8 TGS1 | Essential DNA-binding protein specific to single-stranded yeast telomeric DNA repeats, required for telomere length regulation and telomere position effect | DNA-binding protein of the mitochondria involved in repair of mitochondrial DNA, has ATPase activity and binds to DNA mismatches; has homology to E. coli MutS; transcription is induced during meiosis | Protein kinase involved in transcriptional activation of osmostress-responsive genes; regulates G1 progression, cAPK activity, nitrogen activation of the FGM pathway; involved in life span regulation; homologous to mammalian Akt/PKB | ER membrane protein involved in regulation of OLE1 transcription, acts with homolog Spt23p; inactive ER form dimerizes and one subunit is then activated by ubiquitin/proteasome-dependent processing followed by nuclear targeting | Silencing protein that interacts with Sir2p and Sir4p, and histone H3 and H4 tails, to establish a transcriptionally silent chromatin state; required for spreading of silenced chromatin; recruited to chromatin through interaction with Rap1p | Protein of unknown function; YNR065C is not an essential gene | Protein implicated in the regulation of ergosterol biosynthesis; one of a seven member gene family with a common essential function and non-essential unique functions; similar to human oxysterol binding protein (OSBP) | Probable membrane protein with a possible role in proton symport of glycerol; member of the MBOAT family of putative membrane-bound O-acyltransferases; Gup1p homolog | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends | Protein involved in bud-site selection; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Putative protein of unknown function; localizes to the membrane fraction; YCR101C is not an essential gene | Conserved NAD+ dependent histone deacetylase of the Sirtuin family involved in regulation of lifespan; plays roles in silencing at HML, HMR, telomeres, and the rDNA locus; negatively regulates initiation of DNA replication | Putative GTPase-activating protein (GAP) involved in the Pkc1p-mediated signaling pathway that controls cell wall integrity; appears to specifically regulate 1,3-beta-glucan synthesis | SUMO ligase involved in chromosomal organization and DNA repair; essential subunit of the Mms21-Smc5-Smc6 complex; mutants are sensitive to methyl methanesulfonate and show increased spontaneous mutation and mitotic recombination | DNA dependent ATPase/DNA helicase belonging to the Dna2p- and Nam7p-like family of helicases that is involved in modulating translation termination; interacts with the translation termination factors, localized to polysomes | Essential subunit of the Pan2p-Pan3p poly(A)-ribonuclease complex, which acts to control poly(A) tail length and regulate the stoichiometry and activity of postreplication repair complexes | Protein involved in biosynthesis of cell wall beta-glucans; subunit of the TRAPP (transport protein particle) complex, which is involved in the late steps of endoplasmic reticulum to Golgi transport | Cargo-transport protein involved in endocytosis; may interact with ribosomes, based on co-purification experiments; GFP-fusion protein localizes to the cytoplasm; YGR198W is an essential gene | Biotin synthase, catalyzes the conversion of dethiobiotin to biotin, which is the last step of the biotin biosynthesis pathway; complements E. coli bioB mutant | Subunit of the origin recognition complex, which directs DNA replication by binding to replication origins and is also involved in transcriptional silencing; phosphorylated by Cdc28p | Zinc-finger protein of unknown function, contains one canonical and two unusual fingers in unusual arrangements; deletion enhances replication of positive-strand RNA virus | Protein involved in cell morphogenesis and proliferation, associated with protein kinase Cbk1p; mutants activate OCH1 transcription | Histidine kinase osmosensor that regulates a MAP kinase cascade; transmembrane protein with an intracellular kinase domain that signals to Ypd1p and Ssk1p, thereby forming a phosphorelay system similar to bacterial two-component regulators | Myosin-like protein associated with the nuclear envelope, connects the nuclear pore complex with the nuclear interior; involved in the Tel1p pathway that controls telomere length | Formin, nucleates the formation of linear actin filaments, involved in cell processes such as budding and mitotic spindle orientation which require the formation of polarized actin cables, functionally redundant with BNI1 | Cytosolic J-domain-containing protein, required for peroxisomal protein import and involved in peroxisome assembly, homologous to E. coli DnaJ | Protein of unknown function, expression is sensitive to nitrogen catabolite repression and regulated by Dal80p; contains transmembrane domain | Subunit of the origin recognition complex, which directs DNA replication by binding to replication origins and is also involved in transcriptional silencing | Protein of unknown function, contains an N-terminal epsin-like domain | Integral membrane protein of the nuclear pore; has an important role in maintaining the architecture of the pore complex | Subunit of a complex, with Slx1p, that hydrolyzes 5' branches from duplex DNA in response to stalled or converging replication forks; function overlaps with that of Sgs1p-Top3p | Mitochondrial membrane protein that coordinates expression of mitochondrially-encoded genes; may facilitate delivery of mRNA to membrane-bound translation machinery | Protein involved in transcription of rDNA by RNA polymerase I; transcription factor, member of UAF (upstream activation factor) family along with Rrn9p and Rrn10p | Essential protein with a highly acidic N-terminal domain; IFH1 exhibits genetic interactions with FHL1, overexpression interferes with silencing at telomeres and HM loci; potential Cdc28p substrate | Meiosis-specific protein of unknown function, required for spore wall formation during sporulation; dispensable for both nuclear divisions during meiosis | 1,3-beta-glucanosyltransferase, involved with Gas4p in spore wall assembly; has similarity to Gas1p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR364W is not an essential gene | Protein with RNA-binding motifs required for meiosis-specific mRNA splicing; required for chromosome pairing and meiotic recombination | RNA-binding protein that interacts with the C-terminal domain of the RNA polymerase II large subunit (Rpo21p), required for transcription termination and 3' end maturation of nonpolyadenylated RNAs | Putative protein of unknown function, deletion confers reduced fitness in saline | Transcriptional corepressor involved in the regulation of ribosomal protein gene transcription via the TOR signaling pathway and protein kinase A, phosphorylated by activated Yak1p which promotes accumulation of Crf1p in the nucleus | Putative membrane-localized protein of unknown function | Component of the meiotic outer plaque of the spindle pole body, involved in modifying the meiotic outer plaque that is required prior to prospore membrane formation | Putative component of the outer plaque of the spindle pole body; may be involved in cation homeostasis or multidrug resistance | One of six subunits of RNA polymerase III transcription initiation factor complex (TFIIIC); part of TFIIIC TauB domain that binds BoxB promoter sites of tRNA and other genes; linker between TauB and TauA domains; human homolog is TFIIIC-90 | Trimethyl guanosine synthase, conserved nucleolar methyl transferase responsible for conversion of the m(7)G cap structure of snRNAs and snoRNAs to m(2,2,7)G; also required for ribosome synthesis and nucleolar morphology | Protein of unknown function; exhibits genetic interaction with ERG11 and protein-protein interaction with Hsp82p |
| 297 | 34 | 441 | 0.294 | 0.506 | 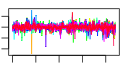 |  |  |  |  | YDL200C YDR124W YGL134W YHR180W YIL055C YJL037W YJL045W YJL100W YJL116C YJR106W YKL162C YLR122C YLR125W YLR173W YLR284C YLR411W YML118W YMR068W YMR084W YMR103C YMR201C YNL012W YNL203C YOR022C YOR032C YOR040W YOR190W YOR392W YPL006W YPL114W YPR005C YPR093C YPR192W YPR193C | MGT1 PCL10 IRC18 LSB6 NCA3 ECM27 ECI1 CTR3 NGL3 AVO2 RAD14 SPO1 HMS1 GLO4 SPR1 NCR1 HAL1 ASR1 AQY1 HPA2 | DNA repair methyltransferase (6-O-methylguanine-DNA methylase) involved in protection against DNA alkylation damage | Putative protein of unknown function; non-essential gene; expression is strongly induced by alpha factor | Pho85p cyclin; recruits, activates, and targets Pho85p cyclin-dependent protein kinase to its substrate | Putative protein of unknown function | Putative protein of unknown function; expression induced in respiratory-deficient cells and in carbon-limited chemostat cultures; similar to adjacent ORF, YJL038C; null mutant displays increased levels of spontaneous Rad52 foci | Minor succinate dehydrogenase isozyme; homologous to Sdh1p, the major isozyme reponsible for the oxidation of succinate and transfer of electrons to ubiquinone; induced during the diauxic shift in a Cat8p-dependent manner | Phosphatidylinositol 4-kinase that binds Las17p, which is a homolog of human Wiskott-Aldrich Syndrome protein involved in actin patch assembly and actin polymerization | Protein that functions with Nca2p to regulate mitochondrial expression of subunits 6 (Atp6p) and 8 (Atp8p ) of the Fo-F1 ATP synthase; member of the SUN family | Non-essential protein of unknown function | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the mitochondrion | Putative protein of unknown function; mutant has decreased Ty3 transposition; YLR125W is not an essential gene | Peroxisomal delta3,delta2-enoyl-CoA isomerase, hexameric protein that converts 3-hexenoyl-CoA to trans-2-hexenoyl-CoA, essential for the beta-oxidation of unsaturated fatty acids, oleate-induced | High-affinity copper transporter of the plasma membrane, acts as a trimer; gene is disrupted by a Ty2 transposon insertion in many laboratory strains of S. cerevisiae | Putative endonuclease, has a domain similar to a magnesium-dependent endonuclease motif in mRNA deadenylase Ccr4p; similar to Ngl1p and Ngl2p | Component of a complex containing the Tor2p kinase and other proteins, which may have a role in regulation of cell growth | Putative protein of unknown function; YMR084W and adjacent ORF YMR085W are merged in related strains | Protein that recognizes and binds damaged DNA during nucleotide excision repair; subunit of Nucleotide Excision Repair Factor 1 (NEF1); contains zinc finger motif; homolog of human XPA protein | Meiosis-specific protein with similarity to phospholipase B, required for meiotic spindle pole body duplication and separation; required for spore formation | Protein with similarity to bovine phospholipase A1; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Basic helix-loop-helix (bHLH) protein with similarity to myc-family transcription factors; overexpression confers hyperfilamentous growth and suppresses the pseudohyphal filamentation defect of a diploid mep1 mep2 homozygous null mutant | Mitochondrial glyoxalase II, catalyzes the hydrolysis of S-D-lactoylglutathione into glutathione and D-lactate | Sporulation-specific exo-1,3-beta-glucanase; contributes to ascospore thermoresistance | Vacuolar membrane protein that transits through the biosynthetic vacuolar protein sorting pathway, involved in sphingolipid metabolism; glycoprotein and functional orthologue of human Niemann Pick C1 (NPC1) protein | Cytoplasmic protein involved in halotolerance; decreases intracellular Na+ (via Ena1p) and increases intracellular K+ by decreasing efflux; expression repressed by Ssn6p-Tup1p and Sko1p and induced by NaCl, KCl, and sorbitol through Gcn4p | Protein involved in a putative alcohol-responsive signaling pathway; accumulates in the nucleus under alcohol stress; contains a Ring/PHD finger domain | Spore-specific water channel that mediates the transport of water across cell membranes, developmentally controlled; may play a role in spore maturation, probably by allowing water outflow, may be involved in freeze tolerance | Tetrameric histone acetyltransferase with similarity to Gcn5p, Hat1p, Elp3p, and Hpa3p; acetylates histones H3 and H4 in vitro and exhibits autoacetylation activity |
| 196 | 48 | 440 | 0.308 | 0.575 |  |  |  |  |  | YBL098W YBR115C YCL018W YDL198C YDR035W YER052C YER069W YER090W YGL202W YGR124W YHR018C YHR162W YIL116W YJR025C YJR109C YJR111C YKL211C YLR027C YMR062C YNL129W YOL058W YOL064C YOL140W YOR130C YBR104W YBR248C YCL009C YCL030C YDL131W YDL182W YDR158W YDR510W YER055C YER073W YGL009C YGL026C YHR063C YJL071W YKL120W YMR108W YNL104C YOR108W YOR184W YOR221C YOR222W YPL273W YPR058W YPR145W | BNA4 LYS2 LEU2 GGC1 ARO3 HOM3 ARG5,6 TRP2 ARO8 ASN2 ARG4 HIS5 BNA1 CPA2 TRP3 AAT2 ECM40 KIC1 ARG1 MET22 ARG8 ORT1 YMC2 HIS7 ILV6 HIS4 LYS21 LYS20 HOM2 SMT3 HIS1 ALD5 LEU1 TRP5 PAN5 ARG2 OAC1 ILV2 LEU4 LEU9 SER1 MCT1 ODC2 SAM4 YMC1 ASN1 | Kynurenine 3-mono oxygenase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Alpha aminoadipate reductase, catalyzes the reduction of alpha-aminoadipate to alpha-aminoadipate 6-semialdehyde, which is the fifth step in biosynthesis of lysine; activation requires posttranslational phosphopantetheinylation by Lys5p | Beta-isopropylmalate dehydrogenase (IMDH), catalyzes the third step in the leucine biosynthesis pathway | Mitochondrial GTP/GDP transporter, essential for mitochondrial genome maintenance; has a role in mitochondrial iron transport; member of the mitochondrial carrier family | 3-deoxy-D-arabino-heptulosonate-7-phosphate (DAHP) synthase, catalyzes the first step in aromatic amino acid biosynthesis and is feedback-inhibited by phenylalanine or high concentration of tyrosine or tryptophan | Aspartate kinase (L-aspartate 4-P-transferase); cytoplasmic enzyme that catalyzes the first step in the common pathway for methionine and threonine biosynthesis; expression regulated by Gcn4p and the general control of amino acid synthesis | Protein that is processed in the mitochondrion to yield acetylglutamate kinase and N-acetyl-gamma-glutamyl-phosphate reductase, which catalyze the 2nd and 3rd steps in arginine biosynthesis; enzymes form a complex with Arg2p | Anthranilate synthase, catalyzes the initial step of tryptophan biosynthesis, forms multifunctional hetero-oligomeric anthranilate synthase:indole-3-glycerol phosphate synthase enzyme complex with Trp3p | Aromatic aminotransferase I, expression is regulated by general control of amino acid biosynthesis | Asparagine synthetase, isozyme of Asn1p; catalyzes the synthesis of L-asparagine from L-aspartate in the asparagine biosynthetic pathway | Argininosuccinate lyase, catalyzes the final step in the arginine biosynthesis pathway | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the mitochondrion | Histidinol-phosphate aminotransferase, catalyzes the seventh step in histidine biosynthesis; responsive to general control of amino acid biosynthesis; mutations cause histidine auxotrophy and sensitivity to Cu, Co, and Ni salts | 3-hydroxyanthranilic acid dioxygenase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Large subunit of carbamoyl phosphate synthetase, which catalyzes a step in the synthesis of citrulline, an arginine precursor | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the mitochondria | Bifunctional enzyme exhibiting both indole-3-glycerol-phosphate synthase and anthranilate synthase activities, forms multifunctional hetero-oligomeric anthranilate synthase:indole-3-glycerol phosphate synthase enzyme complex with Trp2p | Cytosolic aspartate aminotransferase, involved in nitrogen metabolism; localizes to peroxisomes in oleate-grown cells | Mitochondrial ornithine acetyltransferase, catalyzes the fifth step in arginine biosynthesis; also possesses acetylglutamate synthase activity, regenerates acetylglutamate while forming ornithine | Protein kinase of the PAK/Ste20 kinase family, required for cell integrity possibly through regulating 1,6-beta-glucan levels in the wall; physically interacts with Cdc31p (centrin), which is a component of the spindle pole body | Arginosuccinate synthetase, catalyzes the formation of L-argininosuccinate from citrulline and L-aspartate in the arginine biosynthesis pathway; potential Cdc28p substrate | Bisphosphate-3'-nucleotidase, involved in salt tolerance and methionine biogenesis; dephosphorylates 3'-phosphoadenosine-5'-phosphate and 3'-phosphoadenosine-5'-phosphosulfate, intermediates of the sulfate assimilation pathway | Acetylornithine aminotransferase, catalyzes the fourth step in the biosynthesis of the arginine precursor ornithine | Ornithine transporter of the mitochondrial inner membrane, exports ornithine from mitochondria as part of arginine biosynthesis; human ortholog is associated with hyperammonaemia-hyperornithinaemia-homocitrullinuria (HHH) syndrome | Mitochondrial protein, putative inner membrane transporter with a role in oleate metabolism and glutamate biosynthesis; member of the mitochondrial carrier (MCF) family; has similarity with Ymc1p | Imidazole glycerol phosphate synthase (glutamine amidotransferase:cyclase), catalyzes the fifth and sixth steps of histidine biosynthesis and also produces 5-aminoimidazole-4-carboxamide ribotide (AICAR), a purine precursor | Regulatory subunit of acetolactate synthase, which catalyzes the first step of branched-chain amino acid biosynthesis; enhances activity of the Ilv2p catalytic subunit, localizes to mitochondria | Multifunctional enzyme containing phosphoribosyl-ATP pyrophosphatase, phosphoribosyl-AMP cyclohydrolase, and histidinol dehydrogenase activities; catalyzes the second, third, ninth and tenth steps in histidine biosynthesis | Homocitrate synthase isozyme, catalyzes the condensation of acetyl-CoA and alpha-ketoglutarate to form homocitrate, which is the first step in the lysine biosynthesis pathway; highly similar to the other isozyme, Lys20p | Homocitrate synthase isozyme, catalyzes the condensation of acetyl-CoA and alpha-ketoglutarate to form homocitrate, which is the first step in the lysine biosynthesis pathway; highly similar to the other isozyme, Lys21p | Aspartic beta semi-aldehyde dehydrogenase, catalyzes the second step in the common pathway for methionine and threonine biosynthesis; expression regulated by Gcn4p and the general control of amino acid synthesis | Ubiquitin-like protein of the SUMO family, conjugated to lysine residues of target proteins; regulates chromatid cohesion, chromosome segregation, APC-mediated proteolysis, DNA replication and septin ring dynamics | ATP phosphoribosyltransferase, a hexameric enzyme, catalyzes the first step in histidine biosynthesis; mutations cause histidine auxotrophy and sensitivity to Cu, Co, and Ni salts; transcription is regulated by general amino acid control | Mitochondrial aldehyde dehydrogenase, involved in regulation or biosynthesis of electron transport chain components and acetate formation; activated by K+; utilizes NADP+ as the preferred coenzyme; constitutively expressed | Isopropylmalate isomerase, catalyzes the second step in the leucine biosynthesis pathway | Tryptophan synthase involved in tryptophan biosynthesis, regulated by the general control system of amino acid biosynthesis | 2-dehydropantoate 2-reductase, part of the pantothenic acid pathway, structurally homologous to E. coli panE | Acetylglutamate synthase (glutamate N-acetyltransferase), mitochondrial enzyme that catalyzes the first step in the biosynthesis of the arginine precursor ornithine; forms a complex with Arg5,6p | Mitochondrial inner membrane transporter, transports oxaloacetate, sulfate, and thiosulfate; member of the mitochondrial carrier family | Acetolactate synthase, catalyses the first common step in isoleucine and valine biosynthesis and is the target of several classes of inhibitors, localizes to the mitochondria; expression of the gene is under general amino acid control | Alpha-isopropylmalate synthase (2-isopropylmalate synthase); the main isozyme responsible for the first step in the leucine biosynthesis pathway | Alpha-isopropylmalate synthase II (2-isopropylmalate synthase), catalyzes the first step in the leucine biosynthesis pathway; the minor isozyme, responsible for the residual alpha-IPMS activity detected in a leu4 null mutant | 3-phosphoserine aminotransferase, catalyzes the formation of phosphoserine from 3-phosphohydroxypyruvate, required for serine and glycine biosynthesis; regulated by the general control of amino acid biosynthesis mediated by Gcn4p | Predicted malonyl-CoA:ACP transferase, putative component of a type-II mitochondrial fatty acid synthase that produces intermediates for phospholipid remodeling | Mitochondrial inner membrane transporter, exports 2-oxoadipate and 2-oxoglutarate from the mitochondrial matrix to the cytosol for use in lysine and glutamate biosynthesis and in lysine catabolism | S-adenosylmethionine-homocysteine methyltransferase, functions along with Mht1p in the conversion of S-adenosylmethionine (AdoMet) to methionine to control the methionine/AdoMet ratio | Mitochondrial protein, putative inner membrane transporter with a role in oleate metabolism and glutamate biosynthesis; member of the mitochondrial carrier (MCF) family; has similarity with Ymc2p | Asparagine synthetase, isozyme of Asn2p; catalyzes the synthesis of L-asparagine from L-aspartate in the asparagine biosynthetic pathway |
| 237 | 56 | 431 | 0.316 | 0.538 |  |  |  |  |  | YDR079W YHR011W YHR057C YJR113C YPL013C YBL038W YBL107C YBR120C YBR185C YBR262C YBR268W YBR282W YCR003W YDL107W YDR405W YDR462W YEL050C YFR011C YGL068W YGL069C YGL236C YGR021W YGR076C YGR084C YGR207C YGR208W YGR219W YGR220C YHR059W YHR147C YIL070C YIL157C YJL063C YJL180C YJL206C YJR034W YKL003C YKL087C YKL138C YKR085C YLL009C YLR069C YMR157C YMR158W YMR188C YMR193W YMR267W YNL122C YNL185C YNR037C YOL033W YOR187W YPL097W YPL132W YPL173W YPR100W | PET100 DIA4 CPR2 RSM7 MRPS16 MRPL16 CBP6 MBA1 AIM5 MRPL37 MRPL27 MRPL32 MSS2 MRP20 MRPL28 RML2 AIM13 MNP1 MTO1 MRPL25 MRP13 SER2 MRPL9 FYV4 MRPL6 MAM33 COA1 MRPL8 ATP12 PET191 MRP17 CYT2 HSK3 MRPL20 COX17 MEF1 AIM36 MRPS8 MRPS17 MRPL24 PPA2 MRPL19 RSM19 MSE1 RAP1 MSY1 COX11 MRPL40 MRPL51 | Chaperone that specifically facilitates the assembly of cytochrome c oxidase, integral to the mitochondrial inner membrane; interacts with a subcomplex of subunits VII, VIIa, and VIII (Cox7p, Cox9p, and Cox8p) but not with the holoenzyme | Probable mitochondrial seryl-tRNA synthetase, mutant displays increased invasive and pseudohyphal growth | Peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; has a potential role in the secretory pathway | Mitochondrial ribosomal protein of the small subunit, has similarity to E. coli S7 ribosomal protein | Mitochondrial ribosomal protein of the small subunit | Mitochondrial ribosomal protein of the large subunit | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YBL107C is not an essential gene | Mitochondrial translational activator of the COB mRNA; phosphorylated | Protein involved in assembly of mitochondrial respiratory complexes; may act as a receptor for proteins destined for export from the mitochondrial matrix to the inner membrane | Protein of unknown function; non-tagged protein is detected in purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss and reduced growth rate in minimal glycerol media | Peripherally bound inner membrane protein of the mitochondrial matrix involved in membrane insertion of C-terminus of Cox2p, interacts genetically and physically with Cox18p | Mitochondrial ribosomal protein of the large subunit, has similarity to E. coli L2 ribosomal protein; fat21 mutant allele causes inability to utilize oleate and may interfere with activity of the Adr1p transcription factor | Putative protein of unknown function; non-tagged protein is detected in highly purified mitochondria; null mutant displays increased frequency of mitochondrial genome loss (petite formation) and reduced growth rate in minimal glycerol media | Protein associated with the mitochondrial nucleoid; putative mitochondrial ribosomal protein with similarity to E. coli L7/L12 ribosomal protein; required for normal respiratory growth | Mitochondrial protein, forms a heterodimer complex with Mss1p that performs the 5-carboxymethylaminomethyl modification of the wobble uridine base in mitochondrial tRNAs; required for respiration in paromomycin-resistant 15S rRNA mutants | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein that interacts with frataxin (Yfh1p); putative homolog of mammalian electron transfer flavoprotein complex subunit ETF-beta; authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Phosphoserine phosphatase of the phosphoglycerate pathway, involved in serine and glycine biosynthesis, expression is regulated by the available nitrogen source | Protein of unknown function, required for survival upon exposure to K1 killer toxin | Acidic protein of the mitochondrial matrix involved in oxidative phosphorylation; related to the human complement receptor gC1q-R | Mitochondrial inner membrane protein required for assembly of the cytochrome c oxidase complex (complex IV); interacts with complex IV assembly factor Shy1p during the early stages of assembly | Conserved protein required for assembly of alpha and beta subunits into the F1 sector of mitochondrial F1F0 ATP synthase; mutation of human ATP12 reduces active ATP synthase levels and is associated with the disorder ATPAF2 deficiency | Putative protein of unknown function; similar to transcriptional regulators from the Zn[2]-Cys[6] binuclear cluster protein family; mRNA is weakly cell cycle regulated, peaking in S phase; induced rapidly upon MMS treatment | Protein required for assembly of cytochrome c oxidase; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Mitochondrial ribosomal protein of the small subunit; MRP17 exhibits genetic interactions with PET122, encoding a COX3-specific translational activator | Cytochrome c1 heme lyase, involved in maturation of cytochrome c1, which is a subunit of the mitochondrial ubiquinol-cytochrome-c reductase; links heme covalently to apocytochrome c1 | Essential subunit of the Dam1 complex (aka DASH complex), couples kinetochores to the force produced by MT depolymerization thereby aiding in chromosome segregation; is transferred to the kinetochore prior to mitosis | Copper metallochaperone that transfers copper to Sco1p and Cox11p for eventual delivery to cytochrome c oxidase | Mitochondrial elongation factor involved in translational elongation | Protein of unknown function; non-tagged protein is detected in purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss and reduced growth rate in minimal glycerol media | Mitochondrial inorganic pyrophosphatase, required for mitochondrial function and possibly involved in energy generation from inorganic pyrophosphate | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YNL122C is not an essential gene | Mitochondrial ribosomal protein of the small subunit, has similarity to E. coli S19 ribosomal protein | Mitochondrial glutamyl-tRNA synthetase, encoded by a nuclear gene | DNA-binding protein involved in either activation or repression of transcription, depending on binding site context; also binds telomere sequences and plays a role in telomeric position effect (silencing) and telomere structure | Mitochondrial tyrosyl-tRNA synthetase | Mitochondrial inner membrane protein required for delivery of copper to the Cox1p subunit of cytochrome c oxidase; association with mitochondrial ribosomes suggests that copper delivery may occur during translation of Cox1p |
| 222 | 50 | 441 | 0.325 | 0.538 |  |  |  |  |  | YBL014C YJL197W YJR140C YLR223C YLR272C YML010W YML076C YML098W YMR277W YNL233W YNR011C YOL006C YOL034W YOR127W YPR097W YCL045C YCR042C YDL161W YDR243C YIR002C YJL061W YJL095W YJR089W YKL086W YKL205W YKR078W YLR039C YLR095C YLR096W YLR114C YLR143W YLR148W YLR410W YML096W YML099C YMR126C YMR231W YMR287C YMR288W YNL068C YNL272C YOL004W YOR048C YOR093C YOR109W YOR160W YPL009C YPL115C YPL195W YPR055W | RRN6 UBP12 HIR3 IFH1 YCS4 SPT5 WAR1 TAF13 FCP1 BNI4 PRP2 TOP1 SMC5 RGA1 TAF2 ENT1 PRP28 MPH1 NUP82 BCK1 BIR1 SRX1 LOS1 RIC1 IOC2 KIN2 AVL9 PEP3 VIP1 ARG81 DLT1 PEP5 SEM1 HSH155 FKH2 SEC2 SIN3 TAT1 SRO77 MTR10 BEM3 APL5 SEC8 | Protein involved in the transcription of 35S rRNA genes by RNA polymerase I; component of the core factor (CF) complex also composed of Rrn11p, Rrn7p and TATA-binding protein | Ubiquitin carboxyl-terminal hydrolase, ubiquitin-specific protease present in the nucleus and cytoplasm that cleaves ubiquitin from ubiquitinated proteins | Transcriptional corepressor involved in the cell cycle-regulated transcription of histone genes HTA1, HTB1, HHT1, and HHT2; involved in position-dependent gene silencing and nucleosome reassembly | Essential protein with a highly acidic N-terminal domain; IFH1 exhibits genetic interactions with FHL1, overexpression interferes with silencing at telomeres and HM loci; potential Cdc28p substrate | Non-SMC subunit of the condensin complex (Smc2p-Smc4p-Ycs4p-Brn1p-Ycg1p); required for establishment and maintenance of chromosome condensation, chromosome segregation, chromatin binding of condensin and silencing at the mating type locus | Protein that forms a complex with Spt4p and mediates both activation and inhibition of transcription elongation; Spt4p-Spt5p complex also plays a role in pre-mRNA processing | Homodimeric Zn2Cys6 zinc finger transcription factor; binds to a weak acid response element to induce transcription of PDR12 and FUN34, encoding an acid transporter and a putative ammonia transporter, respectively | TFIID subunit (19 kDa), involved in RNA polymerase II transcription initiation, similar to histone H4 with atypical histone fold motif of Spt3-like transcription factors | Carboxy-terminal domain (CTD) phosphatase, essential for dephosphorylation of the repeated C-terminal domain of the RNA polymerase II large subunit (Rpo21p) | Targeting subunit for Glc7p protein phosphatase, localized to the bud neck, required for localization of chitin synthase III to the bud neck via interaction with the chitin synthase III regulatory subunit Skt5p | RNA-dependent ATPase in the DEAH-box family, required for activation of the spliceosome before the first transesterification step in RNA splicing | Topoisomerase I, nuclear enzyme that relieves torsional strain in DNA by cleaving and re-sealing the phosphodiester backbone; relaxes both positively and negatively supercoiled DNA; functions in replication, transcription, and recombination | Structural maintenance of chromosomes (SMC) protein; essential subunit of the Mms21-Smc5-Smc6 complex; required for growth and DNA repair; S. pombe homolog forms a heterodimer with S. pombe Rad18p that is involved in DNA repair | GTPase-activating protein for the polarity-establishment protein Cdc42p; implicated in control of septin organization, pheromone response, and haploid invasive growth | Protein that contains a Phox homology (PX) domain and binds phosphoinositides; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the ER; interacts with Gal80p; YCL045C is not an essential gene | TFIID subunit (150 kDa), involved in RNA polymerase II transcription initiation | Epsin-like protein involved in endocytosis and actin patch assembly and functionally redundant with Ent2p; binds clathrin via a clathrin-binding domain motif at C-terminus | RNA helicase in the DEAD-box family, involved in RNA isomerization at the 5' splice site | Member of the DEAH family of helicases, functions in an error-free DNA damage bypass pathway that involves homologous recombination, mutations confer a mutator phenotype | Nucleoporin, subunit of the nuclear pore complex (NPC); forms a subcomplex with Nup159p and Nsp1p, interacts with Nup116p and is required for proper localization of Nup116p in the NPC | Mitogen-activated protein (MAP) kinase kinase kinase acting in the protein kinase C signaling pathway, which controls cell integrity; upon activation by Pkc1p phosphorylates downstream kinases Mkk1p and Mkk2p | Essential chromosomal passenger protein involved in coordinating cell cycle events for proper chromosome segregation; C-terminal region binds Sli15p, and the middle region, upon phosphorylation, localizes Cbf2p to the spindle at anaphase | Sulfiredoxin, contributes to oxidative stress resistance by reducing cysteine-sulfinic acid groups in the peroxiredoxins Tsa1p and Ahp1p that are formed upon exposure to oxidants; conserved in higher eukaryotes | Nuclear pore protein involved in nuclear export of pre-tRNA | Cytoplasmic protein of unknown function, has similarity to Vps5p; potential Cdc28p substrate; contains a Phox homology (PX) domain and specifically binds phosphatidylinositol 3-phosphate (PtdIns-3-P) | Protein involved in retrograde transport to the cis-Golgi network; forms heterodimer with Rgp1p that acts as a GTP exchange factor for Ypt6p; involved in transcription of rRNA and ribosomal protein genes | Member of a complex (Isw1b) with Isw1p and Ioc4p that exhibits nucleosome-stimulated ATPase activity and acts within coding regions to coordinate transcription elongation with termination and processing, contains a PHD finger motif | Serine/threonine protein kinase involved in regulation of exocytosis; localizes to the cytoplasmic face of the plasma membrane; closely related to Kin1p | Conserved protein involved in exocytic transport from the Golgi; mutation is synthetically lethal with apl2 vps1 double mutation; member of a protein superfamily with orthologs in diverse organisms | Putative protein of unknown function; green fluorescent protein (GFP)-tagged protein localizes to the cytoplasm; YLR143W is not an essential gene | Component of CORVET tethering complex; vacuolar peripheral membrane protein that promotes vesicular docking/fusion reactions in conjunction with SNARE proteins, required for vacuolar biogenesis | Inositol hexakisphosphate (IP6) and inositol heptakisphosphate (IP7) kinase; generation of IP7 by Vip1p is important for phosphate signaling; likely involved in cortical actin cytoskeleton function, by analogy with S. pombe ortholog asp1 | Putative protein of unknown function with similarity to asparagine synthetases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YML096W is not an essential gene and partially overlaps the verified gene RAD10 | Zinc-finger transcription factor of the Zn(2)-Cys(6) binuclear cluster domain type, involved in the regulation of arginine-responsive genes; acts with Arg80p and Arg82p | Protein of unknown function, mutant sensitive to 6-azauracil (6AU) and mycophenolic acid (MPA) | Component of CORVET tethering complex; peripheral vacuolar membrane protein required for protein trafficking and vacuole biogenesis; interacts with Pep7p | Component of the lid subcomplex of the regulatory subunit of the 26S proteasome; ortholog of human DSS1 | U2-snRNP associated splicing factor that forms extensive associations with the branch site-3' splice site-3' exon region upon prespliceosome formation; similarity to the mammalian U2 snRNP-associated splicing factor SAP155 | Forkhead family transcription factor with a major role in the expression of G2/M phase genes; positively regulates transcriptional elongation; negative role in chromatin silencing at HML and HMR; substrate of the Cdc28p/Clb5p kinase | Guanyl-nucleotide exchange factor for the small G-protein Sec4p, located on cytoplasmic vesicles; essential for post-Golgi vesicle transport | Component of the Sin3p-Rpd3p histone deacetylase complex, involved in transcriptional repression and activation of diverse processes, including mating-type switching and meiosis; involved in the maintenance of chromosomal integrity | Amino acid transport protein for valine, leucine, isoleucine, and tyrosine, low-affinity tryptophan and histidine transporter; overexpression confers FK506 and FTY720 resistance | Putative protein of unknown function; deletion causes sensitivity to unfolded protein response-inducing agents | Protein with roles in exocytosis and cation homeostasis; functions in docking and fusion of post-Golgi vesicles with plasma membrane; homolog of Sro7p and Drosophila lethal giant larvae tumor suppressor; interacts with SNARE protein Sec9p | Nuclear import receptor, mediates the nuclear localization of proteins involved in mRNA-nucleus export; promotes dissociation of mRNAs from the nucleus-cytoplasm mRNA shuttling protein Npl3p | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YPL009C is not an essential gene | Rho GTPase activating protein (RhoGAP) involved in control of the cytoskeleton organization; targets the essential Rho-GTPase Cdc42p, which controls establishment and maintenance of cell polarity, including bud-site assembly | Delta adaptin-like subunit of the clathrin associated protein complex (AP-3); functions in transport of alkaline phosphatase to the vacuole via the alternate pathway, suppressor of loss of casein kinase 1 function | Essential 121kDa subunit of the exocyst complex (Sec3p, Sec5p, Sec6p, Sec8p, Sec10p, Sec15p, Exo70p, and Exo84p), which has the essential function of mediating polarized targeting of secretory vesicles to active sites of exocytosis |
| 244 | 45 | 444 | 0.326 | 0.547 |  |  |  |  |  | YDR419W YGR031W YJL047C YJL058C YLR125W YNL019C YNL033W YBL094C YBL098W YBR033W YBR294W YCL006C YCR099C YDL048C YDL139C YDL159W YDR437W YDR521W YEL067C YER184C YGR030C YGR096W YGR126W YHR211W YIL028W YIL143C YIL165C YIL167W YIR042C YJR119C YJR153W YLL016W YLR123C YLR162W YLR233C YLR234W YLR376C YLR457C YOL132W YOL164W YOR030W YPR092W YPR135W YPR194C YPR196W | RAD30 YJL047C-A BIT61 BNA4 EDS1 SUL1 STP4 SCM3 YDL159W-A GPI19 POP6 TPC1 FLO5 SSL2 JHD2 PGU1 EST1 TOP3 PSY3 NBP1 GAS4 YOL164W-A MET30 CTF4 OPT2 | DNA polymerase eta, involved in the predominantly error-free bypass replication of DNA lesions, catalyzes the efficient and accurate synthesis of DNA opposite cyclobutane pyrimidine dimers; homolog of human POLH and bacterial DinB proteins | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function | Subunit of TORC2 (Tor2p-Lst8p-Avo1-Avo2-Tsc11p-Bit61p-Slm1p-Slm2p), a membrane-associated complex that regulates cell cycle-dependent actin cytoskeletal dynamics during polarized growth and cell wall integrity | Putative protein of unknown function; mutant has decreased Ty3 transposition; YLR125W is not an essential gene | Kynurenine 3-mono oxygenase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Putative zinc cluster protein; YBR033W is not an essential gene | High affinity sulfate permease; sulfate uptake is mediated by specific sulfate transporters Sul1p and Sul2p, which control the concentration of endogenous activated sulfate intermediates | Protein containing a Kruppel-type zinc-finger domain; has similarity to Stp1p, Stp2p, and Stp3p | Nonhistone component of centromeric chromatin that binds stoichiometrically to CenH3-H4 histones, required for kinetochore assembly; contains nuclear export signal (NES); required for G2/M progression and localization of Cse4p | Putative protein of unknown function; identified by sequence comparison with hemiascomycetous yeast species | Subunit of GPI-GlcNAc transferase involved in synthesis of N-acetylglucosaminyl phosphatidylinositol (GlcNAc-PI), which is the first intermediate in glycosylphosphatidylinositol (GPI) anchor synthesis, shares similarity with mammalian PIG-P | Putative zinc cluster protein; deletion confers sensitivity to Calcufluor white, and prevents growth on glycerol or lactate as sole carbon source | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends | Mitochondrial membrane transporter that mediates uptake of the essential cofactor thiamine pyrophosphate (ThPP) into mitochondria; expression appears to be regulated by carbon source; member of the mitochondrial carrier family | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus and is induced in response to the DNA-damaging agent MMS | Lectin-like cell wall protein (flocculin) involved in flocculation, binds to mannose chains on the surface of other cells, confers floc-forming ability that is chymotrypsin resistant but heat labile; similar to Flo1p | Component of the holoenzyme form of RNA polymerase transcription factor TFIIH, has DNA-dependent ATPase/helicase activity and is required, with Rad3p, for unwinding promoter DNA; involved in DNA repair; homolog of human ERCC3 | Putative protein of unknown function; in closely related species and other S. cerevisiae strain backgrounds YIL165C and adjacent ORF, YIL164C, likely constitute a single ORF encoding a nitrilase gene | Putative protein of unknown function; YIR042C is a non-essential gene | JmjC domain family histone demethylase specific for H3-K4 (lysine at position 4 of the histone H3 protein); removes methyl groups specifically added by Set1p methyltransferase | Endo-polygalacturonase, pectolytic enzyme that hydrolyzes the alpha-1,4-glycosidic bonds in the rhamnogalacturonan chains in pectins | Putative protein of unknown function; overexpression confers resistance to the antimicrobial peptide MiAMP1 | TLC1 RNA-associated factor involved in telomere length regulation as the recruitment subunit of the telomerase holoenzyme, has a possible role in activating Est2p-TLC1-RNA bound to the telomere | DNA Topoisomerase III, conserved protein that functions in a complex with Sgs1p and Rmi1p to relax single-stranded negatively-supercoiled DNA preferentially, involved in telomere stability and regulation of mitotic recombination | Protein of unknown function; deletion results in a mutator phenotype suggesting a role for this protein as a mutational suppressor; deletion increases sensitivity to anticancer drugs oxaliplatin and cisplatin but not mitomycin C | Spindle pole body (SPB) component, required for the insertion of the duplication plaque into the nuclear membrane during SPB duplication; essential for bipolar spindle formation; component of the Mps2p-Bbp1p complex | 1,3-beta-glucanosyltransferase, involved with Gas2p in spore wall assembly; has similarity to Gas1p; localizes to the cell wall | Putative protein of unknown function; identified by fungal homology and RT-PCR | F-box protein containing five copies of the WD40 motif, controls cell cycle function, sulfur metabolism, and methionine biosynthesis as part of the ubiquitin ligase complex; interacts with and regulates Met4p, localizes within the nucleus | Chromatin-associated protein, required for sister chromatid cohesion; interacts with DNA polymerase alpha (Pol1p) and may link DNA synthesis to sister chromatid cohesion | Oligopeptide transporter; member of the OPT family, with potential orthologs in S. pombe and C. albicans | Putative maltose activator |
| 26 | 51 | 447 | 0.331 | 0.613 |  |  | 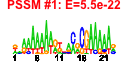 | 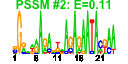 |  | YAR007C YAR008W YBL009W YBR070C YBR089W YBR156C YCR065W YDL003W YDL102W YDL127W YDL205C YDL227C YDR113C YDR261C YDR309C YDR500C YDR507C YDR509W YDR524C YDR526C YDR528W YGR014W YGR109C YGR151C YGR152C YHR149C YHR153C YHR154W YHR173C YIL015W YJL097W YKL112W YKR077W YKR083C YLR212C YLR342W YLR452C YML027W YMR144W YMR199W YNL231C YNL300W YOL007C YOR026W YOR033C YOR195W YPL241C YDR451C YFR028C YHR084W YMR012W | RFA1 SEN34 ALK2 ALG14 SLI15 HCM1 MCD1 POL3 PCL2 HEM3 HO PDS1 EXG2 GIC2 RPL37B GIN4 AGE1 HLR1 MSB2 CLB6 RSR1 SKG6 SPO16 RTT107 BAR1 PHS1 ABF1 MSA2 DAD2 TUB4 FKS1 SST2 YOX1 CLN1 PDR16 CSI2 BUB3 EXO1 SLK19 CIN2 YHP1 CDC14 STE12 CLU1 | Subunit of heterotrimeric Replication Protein A (RPA), which is a highly conserved single-stranded DNA binding protein involved in DNA replication, repair, and recombination | Subunit of the tRNA splicing endonuclease, which is composed of Sen2p, Sen15p, Sen34p, and Sen54p; Sen34p contains the active site for tRNA 3' splice site cleavage and has similarity to Sen2p and to Archaeal tRNA splicing endonuclease | Protein kinase; accumulation and phosphorylation are periodic during the cell cycle; phosphorylated in response to DNA damage; contains characteristic motifs for degradation via the APC pathway; similar to Alk1p and to mammalian haspins | Component of UDP-GlcNAc transferase required for the second step of dolichyl-linked oligosaccharide synthesis; anchors the catalytic subunit Alg13p to the ER membrane; similar to bacterial and human glycosyltransferases | Subunit of the Ipl1p-Sli15p-Bir1p complex that regulates kinetochore-microtubule attachments, activation of the spindle tension checkpoint, and mitotic spindle disassembly; regulates the activity and localization of the Ipl1p aurora kinase | Forkhead transcription factor that drives S-phase specific expression of genes involved in chromosome segregation, spindle dynamics, and budding; suppressor of calmodulin mutants with specific SPB assembly defects; telomere maintenance role | Essential protein required for sister chromatid cohesion in mitosis and meiosis; subunit of the cohesin complex; expression is cell cycle regulated and peaks in S phase | Catalytic subunit of DNA polymerase delta; required for chromosomal DNA replication during mitosis and meiosis, intragenic recombination, repair of double strand DNA breaks, and DNA replication during nucleotide excision repair (NER) | G1 cyclin, associates with Pho85p cyclin-dependent kinase (Cdk) to contribute to entry into the mitotic cell cycle, essential for cell morphogenesis; localizes to sites of polarized cell growth | Phorphobilinogen deaminase, catalyzes the conversion of 4-porphobilinogen to hydroxymethylbilane, the third step in the heme biosynthetic pathway; localizes to both the cytoplasm and nucleus; expression is regulated by Hap2p-Hap3p | Site-specific endonuclease required for gene conversion at the MAT locus (homothallic switching) through the generation of a ds DNA break; expression restricted to mother cells in late G1 as controlled by Swi4p-Swi6p, Swi5p and Ash1p | Securin that inhibits anaphase by binding separin Esp1p, also blocks cyclin destruction and mitotic exit, essential for cell cycle arrest in mitosis in the presence of DNA damage or aberrant mitotic spindles; also present in meiotic nuclei | Exo-1,3-beta-glucanase, involved in cell wall beta-glucan assembly; may be anchored to the plasma membrane via a glycosylphosphatidylinositol (GPI) anchor | Protein of unknown function involved in initiation of budding and cellular polarization, interacts with Cdc42p via the Cdc42/Rac-interactive binding (CRIB) domain | Protein component of the large (60S) ribosomal subunit, has similarity to Rpl37Ap and to rat L37 ribosomal protein | Protein kinase involved in bud growth and assembly of the septin ring, proposed to have kinase-dependent and kinase-independent activities; undergoes autophosphorylation; similar to Kcc4p and Hsl1p | ADP-ribosylation factor (ARF) GTPase activating protein (GAP) effector, involved in the secretory and endocytic pathways; contains C2C2H2 cysteine/histidine motif | Protein involved in regulation of cell wall composition and integrity and response to osmotic stress; overproduction suppresses a lysis sensitive PKC mutation; similar to Lre1p, which functions antagonistically to protein kinase A | Mucin family member involved in the Cdc42p- and MAP kinase-dependent filamentous growth signaling pathway; also functions as an osmosensor in parallel to the Sho1p-mediated pathway; potential Cdc28p substrate | B-type cyclin involved in DNA replication during S phase; activates Cdc28p to promote initiation of DNA synthesis; functions in formation of mitotic spindles along with Clb3p and Clb4p; most abundant during late G1 | GTP-binding protein of the ras superfamily required for bud site selection, morphological changes in response to mating pheromone, and efficient cell fusion; localized to the plasma membrane; significantly similar to mammalian Rap GTPases | Integral membrane protein that localizes primarily to growing sites such as the bud tip or the cell periphery; potential Cdc28p substrate; Skg6p interacts with Zds1p and Zds2p | Protein of unknown function, required for spore formation | Protein implicated in Mms22-dependent DNA repair during S phase, DNA damage induces phosphorylation by Mec1p at one or more SQ/TQ motifs; interacts with Mms22p and Slx4p; has four BRCT domains; has a role in regulation of Ty1 transposition | Aspartyl protease secreted into the periplasmic space of mating type a cells, helps cells find mating partners, cleaves and inactivates alpha factor allowing cells to recover from alpha-factor-induced cell cycle arrest | Protein of unknown function; homolog of mammalian PTPLA; involved in sphingolipid biosynthesis, protein trafficking; required for cell viability | DNA binding protein with possible chromatin-reorganizing activity involved in transcriptional activation, gene silencing, and DNA replication and repair | Putative transcriptional activator, that interacts with G1-specific transcription factor, MBF and G1-specific promoters; ortholog of Msa2p, an MBF and SBF activator that regulates G1-specific transcription and cell cycle initiation | Essential subunit of the Dam1 complex (aka DASH complex), couples kinetochores to the force produced by MT depolymerization thereby aiding in chromosome segregation; is transferred to the kinetochore prior to mitosis | Gamma-tubulin, involved in nucleating microtubules from both the cytoplasmic and nuclear faces of the spindle pole body | Catalytic subunit of 1,3-beta-D-glucan synthase, functionally redundant with alternate catalytic subunit Gsc2p; binds to regulatory subunit Rho1p; involved in cell wall synthesis and maintenance; localizes to sites of cell wall remodeling | GTPase-activating protein for Gpa1p, regulates desensitization to alpha factor pheromone; also required to prevent receptor-independent signaling of the mating pathway; member of the RGS (regulator of G-protein signaling) family | Homeodomain-containing transcriptional repressor, binds to Mcm1p and to early cell cycle boxes (ECBs) in the promoters of cell cycle-regulated genes expressed in M/G1 phase; expression is cell cycle-regulated; potential Cdc28p substrate | Putative protein of unknown function; localized to the nucleus; YMR144W is not an essential gene | G1 cyclin involved in regulation of the cell cycle; activates Cdc28p kinase to promote the G1 to S phase transition; late G1 specific expression depends on transcription factor complexes, MBF (Swi6p-Mbp1p) and SBF (Swi6p-Swi4p) | Phosphatidylinositol transfer protein (PITP) controlled by the multiple drug resistance regulator Pdr1p, localizes to lipid particles and microsomes, controls levels of various lipids, may regulate lipid synthesis, homologous to Pdr17p | Glycosylphosphatidylinositol-dependent cell wall protein, expression is periodic and decreases in respone to ergosterol perturbation or upon entry into stationary phase; depletion increases resistance to lactic acid | Protein of unknown function; green fluorescent protein (GFP)- fusion protein localizes to the mother side of the bud neck and the vacuole; YOL007C is not an essential gene | Kinetochore checkpoint WD40 repeat protein that localizes to kinetochores during prophase and metaphase, delays anaphase in the presence of unattached kinetochores; forms complexes with Mad1p-Bub1p and with Cdc20p, binds Mad2p and Mad3p | 5'-3' exonuclease and flap-endonuclease involved in recombination, double-strand break repair and DNA mismatch repair; member of the Rad2p nuclease family, with conserved N and I nuclease domains | Kinetochore-associated protein required for normal segregation of chromosomes in meiosis and mitosis; component of the FEAR regulatory network, which promotes Cdc14p release from the nucleolus during anaphase; potential Cdc28p substrate | Tubulin folding factor C (putative) involved in beta-tubulin (Tub2p) folding; isolated as mutant with increased chromosome loss and sensitivity to benomyl | One of two homeobox transcriptional repressors (see also Yox1p), that bind to Mcm1p and to early cell cycle box (ECB) elements of cell cycle regulated genes, thereby restricting ECB-mediated transcription to the M/G1 interval | Protein phosphatase required for mitotic exit; located in the nucleolus until liberated by the FEAR and Mitotic Exit Network in anaphase, enabling it to act on key substrates to effect a decrease in CDK/B-cyclin activity and mitotic exit | Transcription factor that is activated by a MAP kinase signaling cascade, activates genes involved in mating or pseudohyphal/invasive growth pathways; cooperates with Tec1p transcription factor to regulate genes specific for invasive growth | eIF3 component of unknown function; deletion causes defects in mitochondrial organization but not in growth or translation initiation, can rescue cytokinesis and mitochondrial organization defects of the Dictyostelium cluA- mutant |
| 217 | 39 | 455 | 0.334 | 0.568 |  |  | 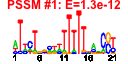 |  |  | YBR290W YDR196C YDR197W YDR248C YGL096W YHR001W YIR024C YJL112W YJR066W YLR097C YML002W YOL053W YDR168W YDR532C YER050C YFR016C YGL087C YGL152C YGL153W YGR044C YGR186W YJL102W YJL131C YJR107W YKL061W YKR016W YLR102C YLR193C YLR211C YLR254C YLR289W YLR326W YML030W YNL100W YOL108C YOR064C YOR147W YPL257W YPR027C | BSD2 CBS2 TOS8 OSH7 MDV1 TOR1 HRT3 AIM39 CDC37 RSM18 MMS2 PEX14 RME1 TFG1 MEF2 AIM23 AIM28 APC9 UPS1 NDL1 GUF1 AIM31 AIM37 INO4 YNG1 MDM32 YPL257W-B | Heavy metal ion homeostasis protein, facilitates trafficking of Smf1p and Smf2p metal transporters to the vacuole where they are degraded, controls metal ion transport, prevents metal hyperaccumulation, functions in copper detoxification | Putative dephospho-CoA kinase (DPCK) that catalyzes the final step in Coenzyme A biosynthesis; essential for viability; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Mitochondrial translational activator of the COB mRNA; interacts with translating ribosomes, acts on the COB mRNA 5'-untranslated leader | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Homeodomain-containing transcription factor; SBF regulated target gene that in turn regulates expression of genes involved in G1/S phase events such as bud site selection, bud emergence and cell cycle progression; similarity to Cup9p | Member of an oxysterol-binding protein family with seven members in S. cerevisiae; family members have overlapping, redundant functions in sterol metabolism and collectively perform a function essential for viability | Protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; interacts with Arh1p, a mitochondrial oxidoreductase; deletion mutant has a respiratory growth defect | Peripheral protein of the cytosolic face of the mitochondrial outer membrane, required for mitochondrial fission; interacts with Fis1p and with the dynamin-related GTPase Dnm1p; contains WD repeats | PIK-related protein kinase and rapamycin target; subunit of TORC1, a complex that controls growth in response to nutrients by regulating translation, transcription, ribosome biogenesis, nutrient transport and autophagy; involved in meiosis | Putative SCF-ubiquitin ligase F-box protein, based on both genetic and physical interactions and sequence similarity; identified in association with Cdc53p, Skp1p and Ubi4 in large and small-scale studies | Putative protein of unknown function; expression induced by heat and by calcium shortage | Putative protein of unknown function; YOL053W is not an essential gene; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) | Essential Hsp90p co-chaperone; necessary for passage through the START phase of the cell cycle; stabilizes protein kinase nascent chains and participates along with Hsp90p in their folding | Protein of unknown function that localizes to the nuclear side of the spindle pole body and along short spindles; deletion mutants have short telomeres; forms a complex with Spc105p | Mitochondrial ribosomal protein of the small subunit, has similarity to E. coli S18 ribosomal protein | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and bud; interacts with Spa2p; YFL016C is not an essential gene | Protein involved in error-free postreplication DNA repair; forms a heteromeric complex with Ubc13p that has a ubiquitin-conjugating activity; cooperates with chromatin-associated RING finger proteins, Rad18p and Rad5p | Peroxisomal membrane peroxin that is a central component of the peroxisomal protein import machinery; interacts with both PTS1 (Pex5p) and PTS2 (Pex7p), peroxisomal matrix protein signal recognition factors and membrane receptor Pex13p | Zinc finger protein involved in control of meiosis; prevents meiosis by repressing IME1 expression and promotes mitosis by activating CLN2 expression; directly repressed by a1-a2 regulator; mediates cell type control of sporulation | TFIIF (Transcription Factor II) largest subunit; involved in both transcription initiation and elongation of RNA polymerase II; homologous to human RAP74 | Mitochondrial elongation factor involved in translational elongation | Putative protein of unknown function; the authentic non-tagged protein is detected in highly purified mitochondria; null mutant is viable, displays increased frequency of mitochondrial genome loss (petite formation) | Putative protein of unknown function; has sequence or structural similarity to lipases | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endosome | Protein of unknown function; non-tagged protein is detected in purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss and severe growth defect in minimal glycerol media | Subunit of the Anaphase-Promoting Complex/Cyclosome (APC/C), which is a ubiquitin-protein ligase required for degradation of anaphase inhibitors, including mitotic cyclins, during the metaphase/anaphase transition | Mitochondrial intermembrane space protein that regulates alternative processing and sorting of Mgm1p and other proteins; required for normal mitochondrial morphology; ortholog of human PRELI | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR211C is not an essential gene; ORF contains an intron | Homolog of nuclear distribution factor NudE, NUDEL; interacts with Pac1p and regulates dynein targeting to microtubule plus ends | Mitochondrial GTPase of unknown function, similar to E. coli elongation factor-type GTP-binding protein LepA and to LK1236.1 from Caenorhabditis elegans | hypothetical protein | Putative protein of unknown function; GFP-fusion protein localizes to mitochondria; null mutant is viable and displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth rate in minimal glycerol media | Putative protein of unknown function; non-tagged protein is detected in purified mitochondria; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Transcription factor required for derepression of inositol-choline-regulated genes involved in phospholipid synthesis; forms a complex, with Ino2p, that binds the inositol-choline-responsive element through a basic helix-loop-helix domain | Subunit of the NuA3 histone acetyltransferase complex that acetylates histone H3; contains PHD finger domain that interacts with methylated histone H3, has similarity to the human tumor suppressor ING1 | Mitochondrial inner membrane protein with similarity to Mdm31p, required for normal mitochondrial morphology and inheritance; interacts genetically with MMM1, MDM10, MDM12, and MDM34 | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Putative protein of unknown function |
| 1 | 36 | 444 | 0.353 | 0.549 |  |  |  |  |  | YBL033C YBR046C YBR047W YBR256C YBR293W YBR297W YDR423C YER175C YGL117W YGL134W YGL218W YGR112W YHL036W YHR017W YHR029C YHR071W YHR161C YIL164C YJL088W YJR130C YKL189W YKR096W YLR189C YLR193C YLR240W YML117W YMR097C YMR197C YNL160W YNL183C YOL055C YOL119C YOR111W YPL224C YPL250C YPR158W | RIB1 ZTA1 FMP23 RIB5 VBA2 MAL33 CAD1 TMT1 PCL10 SHY1 MUP3 YSC83 YHI9 PCL5 YAP1801 NIT1 ARG3 STR2 HYM1 ATG26 UPS1 VPS34 NAB6 MTG1 VTI1 YGP1 NPR1 THI20 MCH4 MMT2 ICY2 | GTP cyclohydrolase II; catalyzes the first step of the riboflavin biosynthesis pathway | Zeta-crystallin homolog, found in the cytoplasm and nucleus; has similarity to E. coli quinone oxidoreductase and to human zeta-crystallin, which has quinone oxidoreductase activity | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Riboflavin synthase; catalyzes the last step of the riboflavin biosynthesis pathway | Permease of basic amino acids in the vacuolar membrane | MAL-activator protein, part of complex locus MAL3; nonfunctional in genomic reference strain S288C | AP-1-like bZIP transcriptional activator involved in stress responses, iron metabolism, and pleiotropic drug resistance; controls a set of genes involved in stabilizing proteins; binds consensus sequence TTACTAA; 5' UTR contains uORFs | Trans-aconitate methyltransferase, cytosolic enzyme that catalyzes the methyl esterification of 3-isopropylmalate, an intermediate of the leucine biosynthetic pathway, and trans-aconitate, which inhibits the citric acid cycle | Putative protein of unknown function | Pho85p cyclin; recruits, activates, and targets Pho85p cyclin-dependent protein kinase to its substrate | Mitochondrial inner membrane protein required for assembly of cytochrome c oxidase (complex IV); associates with complex IV assembly intermediates and complex III/complex IV supercomplexes; similar to human SURF1 involved in Leigh Syndrome | Low affinity methionine permease, similar to Mup1p | Non-essential mitochondrial protein of unknown function; mRNA induced during meiosis, peaking between mid to late prophase of meiosis I; similar to S. douglasii YSD83 | Protein of unknown function; null mutant is defective in unfolded protein response; possibly involved in a membrane regulation metabolic pathway; member of the PhzF superfamily, though most likely not involved in phenazine production | Cyclin, interacts with Pho85p cyclin-dependent kinase (Cdk), induced by Gcn4p at level of transcription, specifically required for Gcn4p degradation, may be sensor of cellular protein biosynthetic capacity | Protein involved in clathrin cage assembly; binds Pan1p and clathrin; homologous to Yap1802p, member of the AP180 protein family | Nitrilase, member of the nitrilase branch of the nitrilase superfamily; in closely related species and other S. cerevisiae strain backgrounds YIL164C and adjacent ORF, YIL165C, likely constitute a single ORF encoding a nitrilase gene | Ornithine carbamoyltransferase (carbamoylphosphate:L-ornithine carbamoyltransferase), catalyzes the sixth step in the biosynthesis of the arginine precursor ornithine | Cystathionine gamma-synthase, converts cysteine into cystathionine | Component of the RAM signaling network that is involved in regulation of Ace2p activity and cellular morphogenesis, interacts with Kic1p and Sog2p, localizes to sites of polarized growth during budding and during the mating response | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and cytoplasm; predicted to contain a PINc (PilT N terminus) domain | UDP-glucose:sterol glucosyltransferase, conserved enzyme involved in synthesis of sterol glucoside membrane lipids, involved in autophagy | Mitochondrial intermembrane space protein that regulates alternative processing and sorting of Mgm1p and other proteins; required for normal mitochondrial morphology; ortholog of human PRELI | Phosphatidylinositol 3-kinase responsible for the synthesis of phosphatidylinositol 3-phosphate; forms membrane-associated signal transduction complex with Vps15p to regulate protein sorting; activated by the GTP-bound form of Gpa1p | Putative RNA-binding protein, based on computational analysis of large-scale protein-protein interaction data | Peripheral GTPase of the mitochondrial inner membrane, essential for respiratory competence, likely functions in assembly of the large ribosomal subunit, has homologs in plants and animals | Protein involved in cis-Golgi membrane traffic; v-SNARE that interacts with two t-SNARES, Sed5p and Pep12p; required for multiple vacuolar sorting pathways | Cell wall-related secretory glycoprotein; induced by nutrient deprivation-associated growth arrest and upon entry into stationary phase; may be involved in adaptation prior to stationary phase entry; has similarity to Sps100p | Protein kinase that stabilizes several plasma membrane amino acid transporters by antagonizing their ubiquitin-mediated degradation | Multifunctional protein with both hydroxymethylpyrimidine kinase and thiaminase activities; involved in thiamine biosynthesis and also in thiamine degradation; member of a gene family with THI21 and THI22; functionally redundant with Thi21p | Protein with similarity to mammalian monocarboxylate permeases, which are involved in transport of monocarboxylic acids across the plasma membrane; mutant is not deficient in monocarboxylate transport | Putative metal transporter involved in mitochondrial iron accumulation; closely related to Mmt1p | Protein of unknown function; mobilized into polysomes upon a shift from a fermentable to nonfermentable carbon source; potential Cdc28p substrate |
| 250 | 44 | 441 | 0.357 | 0.589 | 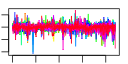 |  | 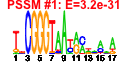 |  |  | YDR257C YLL022C YNL207W YNL232W YNL254C YOL005C YOL145C YOR265W YOR337W YOR359W YPL193W YCR051W YDL208W YDL209C YEL053C YFR037C YGL151W YGL211W YGR251W YHR167W YIL020C YJR003C YLR275W YLR323C YML031W YML082W YMR013C YMR127C YMR214W YMR221C YNL023C YNL140C YNL164C YOL034W YOL142W YOL144W YOR154W YOR205C YOR229W YOR233W YOR269W YOR301W YOR336W YPL101W | SET7 HIF1 RIO2 CSL4 RPB11 CTR9 RBL2 TEA1 VTS1 RSA1 NHP2 CWC2 MAK10 RSC8 NUT1 NCS6 THP2 HIS6 SMD2 CWC24 NDC1 SEC59 SAS2 SCJ1 FMP42 FAP1 IBD2 SMC5 RRP40 NOP8 VPS33 LRC5 WTM2 KIN3 PAC1 RAX1 KRE5 HAP1 | Nuclear protein that contains a SET-domain, which have been shown to mediate methyltransferase activity in other proteins | Non-essential component of the HAT-B histone acetyltransferase complex (Hat1p-Hat2p-Hif1p), localized to the nucleus; has a role in telomeric silencing | Essential serine kinase involved in the processing of the 20S pre-rRNA into mature 18S rRNA; has similarity to Rio1p | Subunit of the exosome, which is an essential complex present in both nucleus and cytoplasm that mediates RNA processing and degradation | hypothetical protein | RNA polymerase II subunit B12.5; part of central core; similar to Rpc19p and bacterial alpha subunit | Component of the Paf1p complex, which is a large complex that binds to and modulates the activity of RNA polymerase II and is required for expression of a subset of genes, including cyclin genes; contains TPR repeats | Protein involved in microtubule morphogenesis, required for protection from excess free beta-tubulin; proposed to be involved the folding of beta-tubulin; similar to mouse beta-tubulin cofactor A | Ty1 enhancer activator required for full levels of Ty enhancer-mediated transcription; C6 zinc cluster DNA-binding protein | Post-transcriptional gene regulator, RNA-binding protein containing a SAM domain; shows genetic interactions with Vti1p, which is a v-SNARE involved in cis-Golgi membrane traffic | Protein involved in the assembly of 60S ribosomal subunits; functionally interacts with Dbp6p; functions in a late nucleoplasmic step of the assembly | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; contains ankyrin (Ank) repeats; YCR051W is not an essential gene | Nuclear protein related to mammalian high mobility group (HMG) proteins, essential for function of H/ACA-type snoRNPs, which are involved in 18S rRNA processing | Protein involved in pre-mRNA splicing, component of a complex containing Cef1p; interacts with Prp19p; contains an RNA recognition motif; has similarity to S. pombe Cwf2p | Non-catalytic subunit of N-terminal acetyltransferase of the NatC type, required for replication of dsRNA virus; expression is glucose-repressible | Component of the RSC chromatin remodeling complex; essential for viability and mitotic growth; homolog of SWI/SNF subunit Swi3p, but unlike Swi3p, does not activate transcription of reporters | Component of the RNA polymerase II mediator complex, which is required for transcriptional activation and also has a role in basal transcription | Protein required for thiolation of the uridine at the wobble position of Gln, Lys, and Glu tRNAs; has a role in urmylation and in invasive and pseudohyphal growth; inhibits replication of Brome mosaic virus in S. cerevisiae | Putative protein of unknown function; deletion mutant has defects in pre-rRNA processing; green fluorescent protein (GFP)-fusion protein localizes to both the nucleus and the nucleolus; YGR251W is an essential gene | Subunit of the THO complex, which connects transcription elongation and mitotic recombination, and of the TREX complex, which is recruited to activated genes and couples transcription to mRNA export; involved in telomere maintenance | Phosphoribosyl-5-amino-1-phosphoribosyl-4- imidazolecarboxiamide isomerase, catalyzes the fourth step in histidine biosynthesis; mutations cause histidine auxotrophy and sensitivity to Cu, Co, and Ni salts | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Core Sm protein Sm D2; part of heteroheptameric complex (with Smb1p, Smd1p, Smd3p, Sme1p, Smx3p, and Smx2p) that is part of the spliceosomal U1, U2, U4, and U5 snRNPs; homolog of human Sm D2 | Essential protein, component of a complex containing Cef1p; has similarity to S. pombe Cwf24p | Nuclear envelope protein with multiple putative transmembrane domains, required for nuclear pore complex assembly and spindle pole body duplication; required for meiosis II | Putative protein predicted to have carbon-sulfur lyase activity; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and the cytoplasm; YNML082W is not an essential gene | Dolichol kinase, catalyzes the terminal step in dolichyl monophosphate (Dol-P) biosynthesis; required for viability and for normal rates of lipid intermediate synthesis and protein N-glycosylation | Histone acetyltransferase (HAT) catalytic subunit of the SAS complex (Sas2p-Sas4p-Sas5p), which acetylates free histones and nucleosomes and regulates transcriptional silencing; member of the MYSTacetyltransferase family | One of several homologs of bacterial chaperone DnaJ, located in the ER lumen where it cooperates with Kar2p to mediate maturation of proteins | Protein that binds to Fpr1p (FKBP12), conferring rapamycin resistance by competing with rapamycin for Fpr1p binding; has similarity to putative transcription factors, including D. melanogaster shuttle craft and human NFX1 | Component of the BUB2-dependent spindle checkpoint pathway, interacts with Bfa1p and functions upstream of Bub2p and Bfa1p | Structural maintenance of chromosomes (SMC) protein; essential subunit of the Mms21-Smc5-Smc6 complex; required for growth and DNA repair; S. pombe homolog forms a heterodimer with S. pombe Rad18p that is involved in DNA repair | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex | Nucleolar protein required for 60S ribosomal subunit biogenesis | ATP-binding protein that is a subunit of the HOPS complex and the CORVET tethering complex; essential for membrane docking and fusion at both the Golgi-to-endosome and endosome-to-vacuole stages of protein transport | Protein of unknown function; non-tagged protein is detected in purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss and severe growth defect in minimal glycerol media | Transcriptional repressor involved in regulation of meiosis and silencing; contains WD repeats | Nonessential protein kinase with unknown cellular role | Protein involved in nuclear migration, part of the dynein/dynactin pathway; targets dynein to microtubule tips, which is necessary for sliding of microtubules along bud cortex; synthetic lethal with bni1; homolog of human LIS1 | Protein involved in bud site selection during bipolar budding; localization requires Rax2p; has similarity to members of the insulin-related peptide superfamily | Protein required for beta-1,6 glucan biosynthesis; mutations result in aberrant morphology and severe growth defects | Zinc finger transcription factor involved in the complex regulation of gene expression in response to levels of heme and oxygen; the S288C sequence differs from other strain backgrounds due to a Ty1 insertion in the carboxy terminus |
| 116 | 41 | 437 | 0.370 | 0.566 | 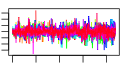 |  | 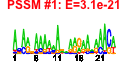 | 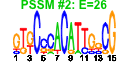 |  | YBL099W YBL100C YBR003W YER141W YFL018C YGR121C YHR198C YIL125W YIL162W YJL171C YKR039W YLR089C YLR246W YLR304C YLR438W YML091C YOL155C YOR136W YOR142W YPL111W YPL221W YPL262W YPL265W YBL030C YCL025C YCR005C YDR247W YDR342C YDR343C YEL052W YHR092C YLL001W YMR145C YMR272C YMR291W YNL037C YNR001C YOR135C YPL057C YPR002W YPR154W | ATP1 COQ1 COX15 LPD1 MEP1 AIM18 KGD1 SUC2 GAP1 ALT1 ERF2 ACO1 CAR2 RPM2 HPF1 IDH2 LSC1 CAR1 FLC1 FUM1 DIP5 PET9 AGP1 CIT2 VHS1 HXT7 HXT6 AFG1 HXT4 DNM1 NDE1 SCS7 IDH1 CIT1 SUR1 PDH1 PIN3 | Alpha subunit of the F1 sector of mitochondrial F1F0 ATP synthase, which is a large, evolutionarily conserved enzyme complex required for ATP synthesis; phosphorylated | Hexaprenyl pyrophosphate synthetase, catalyzes the first step in ubiquinone (coenzyme Q) biosynthesis | Protein required for the hydroxylation of heme O to form heme A, which is an essential prosthetic group for cytochrome c oxidase | Dihydrolipoamide dehydrogenase, the lipoamide dehydrogenase component (E3) of the pyruvate dehydrogenase and 2-oxoglutarate dehydrogenase multi-enzyme complexes | Ammonium permease; belongs to a ubiquitous family of cytoplasmic membrane proteins that transport only ammonium (NH4+); expression is under the nitrogen catabolite repression regulation | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Component of the mitochondrial alpha-ketoglutarate dehydrogenase complex, which catalyzes a key step in the tricarboxylic acid (TCA) cycle, the oxidative decarboxylation of alpha-ketoglutarate to form succinyl-CoA | Invertase, sucrose hydrolyzing enzyme; a secreted, glycosylated form is regulated by glucose repression, and an intracellular, nonglycosylated enzyme is produced constitutively | GPI-anchored cell wall protein of unknown function; induced in response to cell wall damaging agents and by mutations in genes involved in cell wall biogenesis; sequence similarity to YBR162C/TOS1, a covalently bound cell wall protein | General amino acid permease; localization to the plasma membrane is regulated by nitrogen source | Putative alanine transaminase (glutamic pyruvic transaminase); the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Subunit of a palmitoyltransferase, composed of Erf2p and Shr5p, that adds a palmitoyl lipid moiety to heterolipidated substrates such as Ras1p and Ras2p through a thioester linkage; mutants partially mislocalize Ras2p to the vacuole | Aconitase, required for the tricarboxylic acid (TCA) cycle and also independently required for mitochondrial genome maintenance; phosphorylated; component of the mitochondrial nucleoid; mutation leads to glutamate auxotrophy | L-ornithine transaminase (OTAse), catalyzes the second step of arginine degradation, expression is dually-regulated by allophanate induction and a specific arginine induction process; not nitrogen catabolite repression sensitive | Protein subunit of mitochondrial RNase P, has roles in nuclear transcription, cytoplasmic and mitochondrial RNA processing, and mitochondrial translation; distributed to mitochondria, cytoplasmic processing bodies, and the nucleus | Haze-protective mannoprotein that reduces the particle size of aggregated proteins in white wines | Subunit of mitochondrial NAD(+)-dependent isocitrate dehydrogenase, which catalyzes the oxidation of isocitrate to alpha-ketoglutarate in the TCA cycle; phosphorylated | Alpha subunit of succinyl-CoA ligase, which is a mitochondrial enzyme of the TCA cycle that catalyzes the nucleotide-dependent conversion of succinyl-CoA to succinate; phosphorylated | Arginase, responsible for arginine degradation, expression responds to both induction by arginine and nitrogen catabolite repression; disruption enhances freeze tolerance | Putative FAD transporter; required for uptake of FAD into endoplasmic reticulum; involved in cell wall maintenance | Fumarase, converts fumaric acid to L-malic acid in the TCA cycle; cytosolic and mitochondrial localization determined by the N-terminal mitochondrial targeting sequence and protein conformation; phosphorylated in mitochondria | Dicarboxylic amino acid permease, mediates high-affinity and high-capacity transport of L-glutamate and L-aspartate; also a transporter for Gln, Asn, Ser, Ala, and Gly | Major ADP/ATP carrier of the mitochondrial inner membrane, exchanges cytosolic ADP for mitochondrially synthesized ATP; phosphorylated; required for viability in many common lab strains carrying a mutation in the polymorphic SAL1 gene | Low-affinity amino acid permease with broad substrate range, involved in uptake of asparagine, glutamine, and other amino acids; expression is regulated by the SPS plasma membrane amino acid sensor system (Ssy1p-Ptr3p-Ssy5p) | Citrate synthase, catalyzes the condensation of acetyl coenzyme A and oxaloacetate to form citrate, peroxisomal isozyme involved in glyoxylate cycle; expression is controlled by Rtg1p and Rtg2p transcription factors | Cytoplasmic serine/threonine protein kinase; identified as a high-copy suppressor of the synthetic lethality of a sis2 sit4 double mutant, suggesting a role in G1/S phase progression; homolog of Sks1p | High-affinity glucose transporter of the major facilitator superfamily, nearly identical to Hxt6p, expressed at high basal levels relative to other HXTs, expression repressed by high glucose levels | High-affinity glucose transporter of the major facilitator superfamily, nearly identical to Hxt7p, expressed at high basal levels relative to other HXTs, repression of expression by high glucose requires SNF3 | Conserved protein that may act as a chaperone in the degradation of misfolded or unassembled cytochrome c oxidase subunits; localized to matrix face of the mitochondrial inner membrane; member of the AAA family but lacks a protease domain | High-affinity glucose transporter of the major facilitator superfamily, expression is induced by low levels of glucose and repressed by high levels of glucose | Dynamin-related GTPase required for mitochondrial fission and the maintenance of mitochondrial morphology, assembles on the cytoplasmic face of mitochondrial tubules at sites at which division will occur; also participates in endocytosis | Mitochondrial external NADH dehydrogenase, a type II NAD(P)H:quinone oxidoreductase that catalyzes the oxidation of cytosolic NADH; Nde1p and Nde2p provide cytosolic NADH to the mitochondrial respiratory chain | Sphingolipid alpha-hydroxylase, functions in the alpha-hydroxylation of sphingolipid-associated very long chain fatty acids, has both cytochrome b5-like and hydroxylase/desaturase domains, not essential for growth | Putative kinase of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YMR291W is not an essential gene | Subunit of mitochondrial NAD(+)-dependent isocitrate dehydrogenase, which catalyzes the oxidation of isocitrate to alpha-ketoglutarate in the TCA cycle | Citrate synthase, catalyzes the condensation of acetyl coenzyme A and oxaloacetate to form citrate; the rate-limiting enzyme of the TCA cycle; nuclear encoded mitochondrial protein | Probable catalytic subunit of a mannosylinositol phosphorylceramide (MIPC) synthase, forms a complex with probable regulatory subunit Csg2p; function in sphingolipid biosynthesis is overlapping with that of Csh1p | Mitochondrial protein that participates in respiration, induced by diauxic shift; homologous to E. coli PrpD, may take part in the conversion of 2-methylcitrate to 2-methylisocitrate | Protein that induces appearance of [PIN+] prion when overproduced |
| 200 | 50 | 438 | 0.376 | 0.544 |  |  | 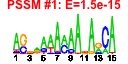 | 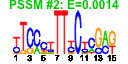 |  | YBR196C YCL045C YFL037W YGL022W YGR054W YJR075W YKL104C YLR354C YLR379W YMR221C YNL044W YNL283C YNL291C YPR159W YBR127C YBR159W YDL015C YDL095W YDR233C YGL001C YGL200C YGL213C YGR279C YHR107C YHR108W YIL021W YIL039W YIL109C YJL002C YJL091C YJL167W YJR001W YJR139C YKL077W YKL080W YKL127W YLR043C YLR378C YMR203W YNL090W YNL263C YOR067C YOR085W YOR204W YOR321W YPL028W YPL053C YPL145C YPR036W YPR181C | PGI1 TUB2 STT3 HOC1 GFA1 TAL1 FMP42 YIP3 WSC2 MID1 KRE6 VMA2 IFA38 TSC13 PMT1 RTN1 ERG26 EMP24 SKI8 SCW4 CDC12 GGA2 RPB3 TED1 SEC24 OST1 GWT1 ERG20 AVT1 HOM6 CBF2 PGM1 TRX2 SEC61 TOM40 RHO2 YIF1 ALG8 OST3 DED1 PMT3 ERG10 KTR6 KES1 VMA13 SEC23 | Glycolytic enzyme phosphoglucose isomerase, catalyzes the interconversion of glucose-6-phosphate and fructose-6-phosphate; required for cell cycle progression and completion of the gluconeogenic events of sporulation | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the ER; interacts with Gal80p; YCL045C is not an essential gene | Beta-tubulin; associates with alpha-tubulin (Tub1p and Tub3p) to form tubulin dimer, which polymerizes to form microtubules | Subunit of the oligosaccharyltransferase complex of the ER lumen, which catalyzes asparagine-linked glycosylation of newly synthesized proteins; forms a subcomplex with Ost3p and Ost4p and is directly involved in catalysis | Eukaryotic initiation factor (eIF) 2A; associates specifically with both 40S subunits and 80 S ribosomes, and interacts genetically with both eIF5b and eIF4E; homologous to mammalian eIF2A | Alpha-1,6-mannosyltransferase involved in cell wall mannan biosynthesis; subunit of a Golgi-localized complex that also contains Anp1p, Mnn9p, Mnn11p, and Mnn10p; identified as a suppressor of a cell lysis sensitive pkc1-371 allele | Glutamine-fructose-6-phosphate amidotransferase, catalyzes the formation of glucosamine-6-P and glutamate from fructose-6-P and glutamine in the first step of chitin biosynthesis | Transaldolase, enzyme in the non-oxidative pentose phosphate pathway; converts sedoheptulose 7-phosphate and glyceraldehyde 3-phosphate to erythrose 4-phosphate and fructose 6-phosphate | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein localized to COPII vesicles, proposed to be involved in ER to Golgi transport; interacts with members of the Rab GTPase family and Yip1p; also interacts with Rtn1p | Partially redundant sensor-transducer of the stress-activated PKC1-MPK1 signaling pathway involved in maintenance of cell wall integrity and recovery from heat shock; secretory pathway Wsc2p is required for the arrest of secretion response | N-glycosylated integral membrane protein of the ER membrane and plasma membrane, functions as a stretch-activated Ca2+-permeable cation channel required for Ca2+ influx stimulated by pheromone; interacts with Cch1p; forms an oligomer | Protein required for beta-1,6 glucan biosynthesis; putative beta-glucan synthase; appears functionally redundant with Skn1p | Subunit B of the eight-subunit V1 peripheral membrane domain of the vacuolar H+-ATPase (V-ATPase), an electrogenic proton pump found throughout the endomembrane system; contains nucleotide binding sites; also detected in the cytoplasm | Microsomal beta-keto-reductase; contains oleate response element (ORE) sequence in the promoter region; mutants exhibit reduced VLCFA synthesis, accumulate high levels of dihydrosphingosine, phytosphingosine and medium-chain ceramides | Enoyl reductase that catalyzes the last step in each cycle of very long chain fatty acid elongation, localizes to the ER, highly enriched in a structure marking nuclear-vacuolar junctions, coimmunoprecipitates with elongases Fen1p and Sur4p | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; acts in a complex with Pmt2p, can instead interact with Pmt3p in some conditions; target for new antifungals | ER membrane protein that interacts with exocyst subunit Sec6p and with Yip3p; also interacts with Sbh1p; null mutant has an altered (mostly cisternal) ER morphology; member of the RTNLA (reticulon-like A) subfamily | C-3 sterol dehydrogenase, catalyzes the second of three steps required to remove two C-4 methyl groups from an intermediate in ergosterol biosynthesis | Integral membrane component of endoplasmic reticulum-derived COPII-coated vesicles, which function in ER to Golgi transport | Protein involved in exosome mediated 3' to 5' mRNA degradation and translation inhibition of non-poly(A) mRNAs as well as double-strand break formation during meiotic recombination; required for repressing propagation of dsRNA viruses | Cell wall protein with similarity to glucanases; scw4 scw10 double mutants exhibit defects in mating | Component of the septin ring of the mother-bud neck that is required for cytokinesis; septins recruit proteins to the neck and can act as a barrier to diffusion at the membrane, and they comprise the 10nm filaments seen with EM | Golgi-localized protein with homology to gamma-adaptin, interacts with and regulates Arf1p and Arf2p in a GTP-dependent manner in order to facilitate traffic through the late Golgi | RNA polymerase II third largest subunit B44, part of central core; similar to prokaryotic alpha subunit | Conserved phosphoesterase domain-containing protein that acts together with Emp24p/Erv25p in cargo exit from the ER; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO) | Component of the Sec23p-Sec24p heterodimeric complex of the COPII vesicle coat; involved in ER to Golgi transport, cargo selection and autophagy; required for the binding of the Sec13 complex to ER membranes; homologous to Lst1p and Lss1p | Alpha subunit of the oligosaccharyltransferase complex of the ER lumen, which catalyzes asparagine-linked glycosylation of newly synthesized proteins | Protein involved in the inositol acylation of glucosaminyl phosphatidylinositol (GlcN-PI) to form glucosaminyl(acyl)phosphatidylinositol (GlcN(acyl)PI), an intermediate in the biosynthesis of glycosylphosphatidylinositol (GPI) anchors | Farnesyl pyrophosphate synthetase, has both dimethylallyltranstransferase and geranyltranstransferase activities; catalyzes the formation of C15 farnesyl pyrophosphate units for isoprenoid and sterol biosynthesis | Vacuolar transporter, imports large neutral amino acids into the vacuole; member of a family of seven S. cerevisiae genes (AVT1-7) related to vesicular GABA-glycine transporters | Homoserine dehydrogenase (L-homoserine:NADP oxidoreductase), dimeric enzyme that catalyzes the third step in the common pathway for methionine and threonine biosynthesis; enzyme has nucleotide-binding, dimerization and catalytic regions | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Essential kinetochore protein, component of the CBF3 multisubunit complex that binds to the CDEIII region of the centromere; Cbf2p also binds to the CDEII region possibly forming a different multimeric complex, ubiquitinated in vivo | Phosphoglucomutase, minor isoform; catalyzes the conversion from glucose-1-phosphate to glucose-6-phosphate, which is a key step in hexose metabolism | Cytoplasmic thioredoxin isoenzyme of the thioredoxin system which protects cells against oxidative and reductive stress, forms LMA1 complex with Pbi2p, acts as a cofactor for Tsa1p, required for ER-Golgi transport and vacuole inheritance | Essential subunit of Sec61 complex (Sec61p, Sbh1p, and Sss1p); forms a channel for SRP-dependent protein import and retrograde transport of misfolded proteins out of the ER; with Sec63 complex allows SRP-independent protein import into ER | Component of the TOM (translocase of outer membrane) complex responsible for recognition and initial import steps for all mitochondrially directed proteins; constitutes the core element of the protein conducting pore | Non-essential small GTPase of the Rho/Rac subfamily of Ras-like proteins, involved in the establishment of cell polarity and in microtubule assembly | Integral membrane protein required for the fusion of ER-derived COPII transport vesicles with the Golgi; interacts with Yip1p and Yos1p; localizes to the Golgi, the ER, and COPII vesicles | Glucosyl transferase, involved in N-linked glycosylation; adds glucose to the dolichol-linked oligosaccharide precursor prior to transfer to protein during lipid-linked oligosaccharide biosynthesis; similar to Alg6p | Gamma subunit of the oligosaccharyltransferase complex of the ER lumen, which catalyzes asparagine-linked glycosylation of newly synthesized proteins; Ost3p is important for N-glycosylation of a subset of proteins | ATP-dependent DEAD (Asp-Glu-Ala-Asp)-box RNA helicase, required for translation initiation of all yeast mRNAs; mutations in human DEAD-box DBY are a frequent cause of male infertility | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; acts in a complex with Pmt5p, can instead interact with Pmt1p in some conditions; target for new antifungals | Acetyl-CoA C-acetyltransferase (acetoacetyl-CoA thiolase), cytosolic enzyme that transfers an acetyl group from one acetyl-CoA molecule to another, forming acetoacetyl-CoA; involved in the first step in mevalonate biosynthesis | Probable mannosylphosphate transferase involved in the synthesis of core oligosaccharides in protein glycosylation pathway; member of the KRE2/MNT1 mannosyltransferase family | Member of the oxysterol binding protein family, which includes seven yeast homologs; involved in negative regulation of Sec14p-dependent Golgi complex secretory functions, peripheral membrane protein that localizes to the Golgi complex | Subunit H of the eight-subunit V1 peripheral membrane domain of the vacuolar H+-ATPase (V-ATPase), an electrogenic proton pump found throughout the endomembrane system; serves as an activator or a structural stabilizer of the V-ATPase | GTPase-activating protein; component of the Sec23p-Sec24p heterodimeric complex of the COPII vesicle coat, involved in ER to Golgi transport and autophagy; stimulates the GDP-bound form of Sar1p |
| 59 | 44 | 447 | 0.379 | 0.541 | 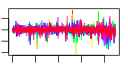 |  | 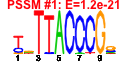 | 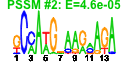 |  | YCL017C YCL065W YDR073W YDR378C YER176W YGL003C YGL119W YGL151W YGL237C YHL020C YHL030W YIL089W YIR002C YKL010C YKL052C YKR048C YLL058W YLR011W YLR091W YLR114C YLR331C YLR417W YLR427W YML114C YMR154C YMR188C YMR279C YNL020C YOR221C YOR299W YOR336W YOR353C YPL011C YPL103C YPL107W YPL125W YPL155C YPL215W YPR040W YPR055W YPR076W YBR007C YDR421W YNL139C | NFS1 SNF11 LSM6 ECM32 CDH1 ABC1 NUT1 HAP2 OPI1 ECM29 MPH1 UFD4 ASK1 NAP1 LOT6 AVL9 VPS36 MAG2 TAF8 RIM13 MRPS17 ARK1 MCT1 BUD7 KRE5 SOG2 TAF3 FMP30 KAP120 KIP2 CBP3 TIP41 SEC8 DSF2 ARO80 RLR1 | Cysteine desulfurase involved in iron-sulfur cluster (Fe/S) biogenesis; required for the post-transcriptional thio-modification of mitochondrial and cytoplasmic tRNAs; essential protein located predominantly in mitochondria | Subunit of the SWI/SNF chromatin remodeling complex involved in transcriptional regulation; interacts with a highly conserved 40-residue sequence of Snf2p | Lsm (Like Sm) protein; part of heteroheptameric complexes (Lsm2p-7p and either Lsm1p or 8p): cytoplasmic Lsm1p complex involved in mRNA decay; nuclear Lsm8p complex part of U6 snRNP and possibly involved in processing tRNA, snoRNA, and rRNA | DNA dependent ATPase/DNA helicase belonging to the Dna2p- and Nam7p-like family of helicases that is involved in modulating translation termination; interacts with the translation termination factors, localized to polysomes | Cell-cycle regulated activator of the anaphase-promoting complex/cyclosome (APC/C), which directs ubiquitination of cyclins resulting in mitotic exit; targets the APC/C to specific substrates including CDC20, ASE1, CIN8 and FIN1 | Protein required for ubiquinone (coenzyme Q) biosynthesis and for respiratory growth; exhibits genetic interaction with COQ9, suggesting a common function; similar to prokaryotic proteins involved in early steps of ubiquinone biosynthesis | Component of the RNA polymerase II mediator complex, which is required for transcriptional activation and also has a role in basal transcription | Subunit of the heme-activated, glucose-repressed Hap2p/3p/4p/5p CCAAT-binding complex, a transcriptional activator and global regulator of respiratory gene expression; contains sequences sufficient for both complex assembly and DNA binding | Transcriptional regulator of a variety of genes; phosphorylation by protein kinase A stimulates Opi1p function in negative regulation of phospholipid biosynthetic genes; involved in telomere maintenance | Major component of the proteasome; tethers the proteasome core particle to the regulatory particle, and enhances the stability of the proteasome | Putative protein of unknown function | Member of the DEAH family of helicases, functions in an error-free DNA damage bypass pathway that involves homologous recombination, mutations confer a mutator phenotype | Ubiquitin-protein ligase (E3) that interacts with Rpt4p and Rpt6p, two subunits of the 19S particle of the 26S proteasome; cytoplasmic E3 involved in the degradation of ubiquitin fusion proteins | Essential subunit of the Dam1 complex (aka DASH complex), couples kinetochores to the force produced by MT depolymerization thereby aiding in chromosome segregation; phosphorylated during the cell cycle by cyclin-dependent kinases | Protein that interacts with mitotic cyclin Clb2p; required for the regulation of microtubule dynamics during mitosis; controls bud morphogenesis; involved in the transport of H2A and H2B histones to the nucleus | Putative protein of unknown function with similarity to Str2p, which is a cystathionine gamma-synthase important in sulfur metabolism; YLL058W is not an essential gene | FMN-dependent NAD(P)H:quinone reductase that may be involved in quinone detoxification; gene expression increases in cultures shifted to a lower temperature | Protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YLR091W is not an esssential gene | Conserved protein involved in exocytic transport from the Golgi; mutation is synthetically lethal with apl2 vps1 double mutation; member of a protein superfamily with orthologs in diverse organisms | Component of the ESCRT-II complex; contains the GLUE (GRAM Like Ubiquitin binding in EAP45) domain which is involved in interactions with ESCRT-I and ubiquitin-dependent sorting of proteins into the endosome | Cytoplasmic protein of unknown function predicted to encode a DNA-3-methyladenine glycosidase II that catalyzes the hydrolysis of alkylated DNA | TFIID subunit (65 kDa), involved in RNA polymerase II transcription initiation | Calpain-like cysteine protease involved in proteolytic activation of Rim101p in response to alkaline pH; has similarity to A. nidulans palB | Mitochondrial ribosomal protein of the small subunit | Putative protein of unknown function; identified as a heat-induced gene in a high-throughout screen; YMR279C is not an essential gene | Serine/threonine protein kinase involved in regulation of the cortical actin cytoskeleton; involved in control of endocytosis | Predicted malonyl-CoA:ACP transferase, putative component of a type-II mitochondrial fatty acid synthase that produces intermediates for phospholipid remodeling | Member of the ChAPs family of proteins (Chs5p-Arf1p-binding proteins: Bch1p, Bch2p, Bud7p, Chs6p), that forms the exomer complex with Chs5p to mediate export of specific cargo proteins, including Chs3p, from the Golgi to the plasma membrane | Protein required for beta-1,6 glucan biosynthesis; mutations result in aberrant morphology and severe growth defects | Key component of the RAM signaling network, required for proper cell morphogenesis and cell separation after mitosis | TFIID subunit (47 kDa), involved in promoter binding and RNA polymerase II transcription initiation | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria; YPL107W is not an essential gene | Karyopherin with a role in the assembly or export of 60S ribosomal subunits | Kinesin-related motor protein involved in mitotic spindle positioning, stabilizes microtubules by targeting Bik1p to the plus end; Kip2p levels are controlled during the cell cycle | Mitochondrial protein required for assembly of ubiquinol cytochrome-c reductase complex (cytochrome bc1 complex); interacts with Cbp4p and function is partially redundant with that of Cbp4p | Protein that interacts physically and genetically with Tap42p, which regulates protein phosphatase 2A; component of the TOR (target of rapamycin) signaling pathway | Essential 121kDa subunit of the exocyst complex (Sec3p, Sec5p, Sec6p, Sec8p, Sec10p, Sec15p, Exo70p, and Exo84p), which has the essential function of mediating polarized targeting of secretory vesicles to active sites of exocytosis | Deletion suppressor of mpt5 mutation | Zinc finger transcriptional activator of the Zn2Cys6 family; activates transcription of aromatic amino acid catabolic genes in the presence of aromatic amino acids | Subunit of the THO complex, which is required for efficient transcription elongation and involved in transcriptional elongation-associated recombination; required for LacZ RNA expression from certain plasmids |
| 18 | 34 | 450 | 0.380 | 0.575 | 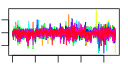 |  | 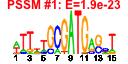 | 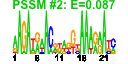 |  | YBL004W YBL018C YCR058C YDL152W YDR020C YDR021W YDR075W YEL055C YER002W YER082C YGR083C YHR081W YHR085W YIL090W YIL091C YIL104C YJL025W YJL098W YKL083W YKR079C YKR099W YLR074C YNL186W YNL227C YOL021C YOL079W YOL144W YOR048C YOR146W YOR154W YOR233W YPL044C YPL068C YPL157W | UTP20 POP8 PWP2 DAS2 FAL1 PPH3 POL5 NOP16 UTP7 GCD2 LRP1 IPI1 ICE2 SHQ1 RRN7 SAP185 TRZ1 BAS1 BUD20 UBP10 JJJ1 DIS3 NOP8 TAT1 VPS33 KIN3 TGS1 | Component of the small-subunit (SSU) processome, which is involved in the biogenesis of the 18S rRNA | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends | Conserved 90S pre-ribosomal component essential for proper endonucleolytic cleavage of the 35 S rRNA precursor at A0, A1, and A2 sites; contains eight WD-repeats; PWP2 deletion leads to defects in cell cycle and bud morphogenesis | Predicted protein shares weak similarity with uridine kinases and with phosphoribokinases; null mutant suppresses dst1delta sensitivity for 6-azauracil | Nucleolar protein required for maturation of 18S rRNA, member of the eIF4A subfamily of DEAD-box ATP-dependent RNA helicases | Catalytic subunit of an evolutionarily conserved protein phosphatase complex containing Psy2p and the regulatory subunit Psy4p; required for cisplatin resistance; involved in activation of Gln3p | DNA Polymerase phi; has sequence similarity to the human MybBP1A and weak sequence similarity to B-type DNA polymerases, not required for chromosomal DNA replication; required for the synthesis of rRNA | Constituent of 66S pre-ribosomal particles, involved in 60S ribosomal subunit biogenesis | Nucleolar protein, component of the small subunit (SSU) processome containing the U3 snoRNA that is involved in processing of pre-18S rRNA | Delta subunit of the translation initiation factor eIF2B, the guanine-nucleotide exchange factor for eIF2; activity subsequently regulated by phosphorylated eIF2; first identified as a negative regulator of GCN4 expression | Substrate-specific nuclear cofactor for exosome activity in the processing of stable RNAs; required for telomere length maintenance; homolog of mammalian nuclear matrix protein C1D involved in regulation of DNA repair and recombination | Essential component of the Rix1 complex (with Rix1p and Ipi3p) that is required for processing of ITS2 sequences from 35S pre-rRNA; Rix1 complex associates with Mdn1p in pre-60S ribosomal particles | Integral ER membrane protein with type-III transmembrane domains; mutations cause defects in cortical ER morphology in both the mother and daughter cells | Protein required for cell viability | Essential nuclear protein, required for accumulation of box H/ACA snoRNAs and for rRNA processing; interacts with Naf1p | Protein involved in the transcription of 35S rRNA genes by RNA polymerase I; component of the core factor (CF) complex also composed of Rrn11p, Rrn6p and TATA-binding protein | Protein that forms a complex with the Sit4p protein phosphatase and is required for its function; member of a family of similar proteins including Sap4p, Sap155p, and Sap190p | tRNase Z, involved in RNA processing, has two putative nucleotide triphosphate-binding motifs (P-loop) and a conserved histidine motif, homolog of the human candidate prostate cancer susceptibility gene ELAC2 | Myb-related transcription factor involved in regulating basal and induced expression of genes of the purine and histidine biosynthesis pathways; also involved in regulation of meiotic recombination at specific genes | Protein involved in bud-site selection; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Ubiquitin-specific protease that deubiquitinates ubiquitin-protein moieties; may regulate silencing by acting on Sir4p; involved in posttranscriptionally regulating Gap1p and possibly other transporters; primarily located in the nucleus | Co-chaperone that stimulates the ATPase activity of Ssa1p, required for a late step of ribosome biogenesis; associated with the cytosolic large ribosomal subunit; contains a J-domain; mutation causes defects in fluid-phase endocytosis | Catalytic component of the exosome, involved in RNA processing and degradation; binds Gsp1p/Ran and enhances the GEF activity of Srm1p; implicated in mitotic control; homologous to the E. coli RNase R of the RNase II family | Nucleolar protein required for 60S ribosomal subunit biogenesis | Amino acid transport protein for valine, leucine, isoleucine, and tyrosine, low-affinity tryptophan and histidine transporter; overexpression confers FK506 and FTY720 resistance | ATP-binding protein that is a subunit of the HOPS complex and the CORVET tethering complex; essential for membrane docking and fusion at both the Golgi-to-endosome and endosome-to-vacuole stages of protein transport | Nonessential protein kinase with unknown cellular role | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus and is induced in response to the DNA-damaging agent MMS | Trimethyl guanosine synthase, conserved nucleolar methyl transferase responsible for conversion of the m(7)G cap structure of snRNAs and snoRNAs to m(2,2,7)G; also required for ribosome synthesis and nucleolar morphology |
| 182 | 38 | 433 | 0.393 | 0.579 |  |  | 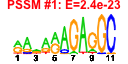 |  |  | YBR005W YBR105C YDR251W YDR293C YDR380W YHR030C YHR137W YIL117C YJL213W YJR103W YLR266C YOR385W YAR033W YBR068C YBR287W YDL234C YDR055W YGL062W YHR159W YIL046W YIL121W YJL171C YJL172W YJL217W YJR046W YKL213C YKL218C YLR090W YMR034C YMR035W YMR195W YNL073W YNL074C YNL193W YNR059W YOR219C YOR317W YPL265W | RCR1 VID24 PAM1 SSD1 ARO10 SYN8 ARO9 PRM5 URA8 PDR8 MST28 BAP2 GYP7 PST1 PYC1 YIL046W-A QDR2 CPS1 TAH11 DOA1 SRY1 XDJ1 IMP2' ICY1 MSK1 MLF3 MNT4 STE13 FAA1 DIP5 | Protein of the ER membrane involved in cell wall chitin deposition; may function in the endosomal-vacuolar trafficking pathway, helping determine whether plasma membrane proteins are degraded or routed to the plasma membrane | Peripheral membrane protein located at Vid (vacuole import and degradation) vesicles; regulates fructose-1,6-bisphosphatase (FBPase) targeting to the vacuole; involved in proteasome-dependent catabolite degradation of FBPase | Essential protein of unknown function; exhibits variable expression during colony morphogenesis; overexpression permits survival without protein phosphatase 2A, inhibits growth, and induces a filamentous phenotype | Protein with a role in maintenance of cellular integrity, interacts with components of the TOR pathway; ssd1 mutant of a clinical S. cerevisiae strain displays elevated virulence | Phenylpyruvate decarboxylase, catalyzes decarboxylation of phenylpyruvate to phenylacetaldehyde, which is the first specific step in the Ehrlich pathway | Endosomal SNARE related to mammalian syntaxin 8 | Aromatic aminotransferase II, catalyzes the first step of tryptophan, phenylalanine, and tyrosine catabolism | Pheromone-regulated protein, predicted to have 1 transmembrane segment; induced during cell integrity signaling | Protein of unknown function that may interact with ribosomes; periodically expressed during the yeast metabolic cycle; phosphorylated in vitro by the mitotic exit network (MEN) kinase complex, Dbf2p/Mob1p | Minor CTP synthase isozyme (see also URA7), catalyzes the ATP-dependent transfer of the amide nitrogen from glutamine to UTP, forming CTP, the final step in de novo biosynthesis of pyrimidines; involved in phospholipid biosynthesis | Transcription factor; targets include ATP-binding cassette (ABC) transporters, major facilitator superfamily transporters, and other genes involved in the pleiotropic drug resistance (PDR) phenomenon | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YOR385W is not an essential gene | Putative integral membrane protein, involved in vesicle formation; forms complex with Mst27p; member of DUP240 gene family; binds COPI and COPII vesicles | High-affinity leucine permease, functions as a branched-chain amino acid permease involved in the uptake of leucine, isoleucine and valine; contains 12 predicted transmembrane domains | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the ER; YBR287W is not an essential gene | GTPase-activating protein for yeast Rab family members including: Ypt7p (most effective), Ypt1p, Ypt31p, and Ypt32p (in vitro); involved in vesicle mediated protein trafficking | Cell wall protein that contains a putative GPI-attachment site; secreted by regenerating protoplasts; up-regulated by activation of the cell integrity pathway, as mediated by Rlm1p; upregulated by cell wall damage via disruption of FKS1 | Pyruvate carboxylase isoform, cytoplasmic enzyme that converts pyruvate to oxaloacetate; highly similar to isoform Pyc2p but differentially regulated; mutations in the human homolog are associated with lactic acidosis | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; potential Cdc28p substrate | Putative protein of unknown function; identified by expression profiling and mass spectrometry | Multidrug transporter of the major facilitator superfamily, required for resistance to quinidine, barban, cisplatin, and bleomycin; may have a role in potassium uptake | GPI-anchored cell wall protein of unknown function; induced in response to cell wall damaging agents and by mutations in genes involved in cell wall biogenesis; sequence similarity to YBR162C/TOS1, a covalently bound cell wall protein | Vacuolar carboxypeptidase yscS; expression is induced under low-nitrogen conditions | Cytoplasmic protein of unknown function; expression induced by calcium shortage and via the copper sensing transciption factor Mac1p during conditons of copper deficiency; mRNA is cell cycle regulated, peaking in G1 phase | DNA replication licensing factor, required for pre-replication complex assembly | WD repeat protein required for ubiquitin-mediated protein degradation, forms complex with Cdc48p, plays a role in controlling cellular ubiquitin concentration; also promotes efficient NHEJ in postdiauxic/stationary phase | 3-hydroxyaspartate dehydratase, deaminates L-threo-3-hydroxyaspartate to form oxaloacetate and ammonia; required for survival in the presence of hydroxyaspartate | Putative chaperone, homolog of E. coli DnaJ, closely related to Ydj1p; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative transporter, member of the SLC10 carrier family; identified in a transposon mutagenesis screen as a gene involved in azole resistance; YMR034C is not an essential gene | Transcriptional activator involved in maintenance of ion homeostasis and protection against DNA damage caused by bleomycin and other oxidants, contains a C-terminal leucine-rich repeat | Protein of unknown function, required for viability in rich media of cells lacking mitochondrial DNA; mutants have an invasive growth defect with elongated morphology; induced by amino acid starvation | Mitochondrial lysine-tRNA synthetase, required for import of both aminoacylated and deacylated forms of tRNA(Lys) into mitochondria | Serine-rich protein of unknown function; overproduction suppresses the growth inhibition caused by exposure to the immunosuppressant leflunomide | Putative protein of unknown function; exhibits a two-hybrid interaction with Yhr151cp in a large-scale analysis | Putative alpha-1,3-mannosyltransferase, not required for protein O-glycosylation | Dipeptidyl aminopeptidase, Golgi integral membrane protein that cleaves on the carboxyl side of repeating -X-Ala- sequences, required for maturation of alpha factor, transcription is induced by a-factor | Long chain fatty acyl-CoA synthetase with a preference for C12:0-C16:0 fatty acids; involved in the activation of imported fatty acids; localized to both lipid particles and mitochondrial outer membrane; essential for stationary phase | Dicarboxylic amino acid permease, mediates high-affinity and high-capacity transport of L-glutamate and L-aspartate; also a transporter for Gln, Asn, Ser, Ala, and Gly |
| 181 | 50 | 433 | 0.406 | 0.596 |  |  |  |  |  | YBR014C YCL043C YDL078C YDL103C YER048C YFR041C YGL157W YGR192C YHR043C YJL020C YKL152C YKR003W YLR301W YMR004W YMR015C YMR092C YNL079C YPL049C YBL056W YBR082C YCR088W YDL029W YDL052C YDL070W YDR388W YER027C YER062C YER177W YFR010W YFR042W YGL221C YGL242C YGR284C YIL010W YIL137C YIL153W YIL154C YJL218W YKL206C YLR028C YLR192C YMR099C YMR226C YNL098C YNL281W YOL031C YOR176W YOR267C YPL106C YPL208W | GRX7 PDI1 MDH3 QRI1 CAJ1 ERJ5 TDH3 DOG2 BBC1 GPM1 OSH6 MVP1 ERG5 AIP1 TPM1 DIG1 PTC3 UBC4 ABP1 ARP2 SLC1 BDF2 RVS167 GAL83 HOR2 BMH1 UBP6 KEG1 NIF3 ERV29 DOT5 TMA108 RPP2B IMP2' ADD66 ADE16 HCR1 TMA29 RAS2 HCH1 SLS1 HEM15 HRK1 SSE1 RKM1 | Monothiol glutaredoxin; more similar in activity to dithiol than other monothiol glutaredoxins; membrane localized; forms homodimers; does not bind metal ions | Protein disulfide isomerase, multifunctional protein resident in the endoplasmic reticulum lumen, essential for the formation of disulfide bonds in secretory and cell-surface proteins, unscrambles non-native disulfide bonds | Peroxisomal malate dehydrogenase, catalyzes interconversion of malate and oxaloacetate; involved in the glyoxylate cycle | UDP-N-acetylglucosamine pyrophosphorylase, catalyzes the formation of UDP-N-acetylglucosamine (UDP-GlcNAc), which is important in cell wall biosynthesis, protein N-glycosylation, and GPI anchor biosynthesis | Nuclear type II J heat shock protein of the E. coli dnaJ family, contains a leucine zipper-like motif, binds to non-native substrates for presentation to Ssa3p, may function during protein translocation, assembly and disassembly | Type I membrane protein with a J domain is required to preserve the folding capacity of the endoplasmic reticulum; loss of the non-essential ERJ5 gene leads to a constitutively induced unfolded protein response | Oxidoreductase, catalyzes NADPH-dependent reduction of the bicyclic diketone bicyclo[2.2.2]octane-2,6-dione (BCO2,6D) to the chiral ketoalcohol (1R,4S,6S)-6-hydroxybicyclo[2.2.2]octane-2-one (BCO2one6ol) | Glyceraldehyde-3-phosphate dehydrogenase, isozyme 3, involved in glycolysis and gluconeogenesis; tetramer that catalyzes the reaction of glyceraldehyde-3-phosphate to 1,3 bis-phosphoglycerate; detected in the cytoplasm and cell-wall | 2-deoxyglucose-6-phosphate phosphatase, member of a family of low molecular weight phosphatases, similar to Dog1p, induced by oxidative and osmotic stress, confers 2-deoxyglucose resistance when overexpressed | Protein possibly involved in assembly of actin patches; interacts with an actin assembly factor Las17p and with the SH3 domains of Type I myosins Myo3p and Myo5p; localized predominantly to cortical actin patches | Tetrameric phosphoglycerate mutase, mediates the conversion of 3-phosphoglycerate to 2-phosphoglycerate during glycolysis and the reverse reaction during gluconeogenesis | Member of an oxysterol-binding protein family with overlapping, redundant functions in sterol metabolism and which collectively perform a function essential for viability; GFP-fusion protein localizes to the cell periphery | Protein of unknown function that interacts with Sec72p | Protein required for sorting proteins to the vacuole; overproduction of Mvp1p suppresses several dominant VPS1 mutations; Mvp1p and Vps1p act in concert to promote membrane traffic to the vacuole | C-22 sterol desaturase, a cytochrome P450 enzyme that catalyzes the formation of the C-22(23) double bond in the sterol side chain in ergosterol biosynthesis; may be a target of azole antifungal drugs | Actin cortical patch component, interacts with the actin depolymerizing factor cofilin; required to restrict cofilin localization to cortical patches; contains WD repeats | Major isoform of tropomyosin; binds to and stabilizes actin cables and filaments, which direct polarized cell growth and the distribution of several organelles; acetylated by the NatB complex and acetylated form binds actin most efficiently | Regulatory protein of unknown function, constitutively-expressed, involved in the regulation of mating-specific genes and the invasive growth pathway, required for MAP-kinase imposed repression, inhibits pheromone-responsive transcription | Type 2C protein phosphatase; dephosphorylates Hog1p (see also Ptc2p) to limit maximal kinase activity induced by osmotic stress; dephosphorylates T169 phosphorylated Cdc28p (see also Ptc2p); role in DNA checkpoint inactivation | Ubiquitin-conjugating enzyme (E2), mediates degradation of short-lived and abnormal proteins; interacts with E3-CaM in ubiquitinating calmodulin; interacts with many SCF ubiquitin protein ligases; component of the cellular stress response | Actin-binding protein of the cortical actin cytoskeleton, important for activation of the Arp2/3 complex that plays a key role actin in cytoskeleton organization | Essential component of the Arp2/3 complex, which is a highly conserved actin nucleation center required for the motility and integrity of actin patches; involved in endocytosis and membrane growth and polarity | 1-acyl-sn-gylcerol-3-phosphate acyltransferase, catalyzes the acylation of lysophosphatidic acid to form phosphatidic acid, a key intermediate in lipid metabolism; located in lipid particles and endoplasmic reticulum | Protein involved in transcription initiation at TATA-containing promoters; associates with the basal transcription factor TFIID; contains two bromodomains; corresponds to the C-terminal region of mammalian TAF1; redundant with Bdf1p | Actin-associated protein, subunit of a complex (Rvs161p-Rvs167p) involved in regulation of actin cytoskeleton, endocytosis, and viability following starvation or osmotic stress; homolog of mammalian amphiphysin | One of three possible beta-subunits of the Snf1 kinase complex, allows nuclear localization of the Snf1 kinase complex in the presence of a nonfermentable carbon source; contains glycogen-binding domain | One of two redundant DL-glycerol-3-phosphatases (RHR2/GPP1 encodes the other) involved in glycerol biosynthesis; induced in response to hyperosmotic stress and oxidative stress, and during the diauxic transition | 14-3-3 protein, major isoform; controls proteome at post-transcriptional level, binds proteins and DNA, involved in regulation of many processes including exocytosis, vesicle transport, Ras/MAPK signaling, and rapamycin-sensitive signaling | Ubiquitin-specific protease situated in the base subcomplex of the 26S proteasome, releases free ubiquitin from branched polyubiquitin chains; deletion causes hypersensitivity to cycloheximide and other toxic compounds | Protein of unknown function required for cell viability; localizes to the endoplasmic reticulum membrane | Protein of unknown function, similar to Listeria monocytogenes major sigma factor (rpoD gene product); the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; deletion mutant is viable | Protein localized to COPII-coated vesicles, involved in vesicle formation and incorporation of specific secretory cargo | Nuclear thiol peroxidase which functions as an alkyl-hydroperoxide reductase during post-diauxic growth | Protein that associates with ribosomes; putative metalloprotease | Ribosomal protein P2 beta, a component of the ribosomal stalk, which is involved in the interaction between translational elongation factors and the ribosome; regulates the accumulation of P1 (Rpp1Ap and Rpp1Bp) in the cytoplasm | Transcriptional activator involved in maintenance of ion homeostasis and protection against DNA damage caused by bleomycin and other oxidants, contains a C-terminal leucine-rich repeat | Putative protein of unknown function, similar to bacterial galactoside O-acetyltransferases; induced by oleate in an OAF1/PIP2-dependent manner; promoter contains an oleate response element consensus sequence; non-essential gene | Protein involved in 20S proteasome assembly; forms a heterodimer with Pba1p that binds to proteasome precursors; similar to human PAC2 constituent of the PAC1-PAC2 complex involved in proteasome assembly | Enzyme of 'de novo' purine biosynthesis containing both 5-aminoimidazole-4-carboxamide ribonucleotide transformylase and inosine monophosphate cyclohydrolase activities, isozyme of Ade17p; ade16 ade17 mutants require adenine and histidine | Dual function protein involved in translation initiation as a substoichiometric component of eukaryotic translation initiation factor 3 (eIF3) and required for processing of 20S pre-rRNA; binds to eIF3 subunits Rpg1p and Prt1p and 18S rRNA | Glucose-6-phosphate 1-epimerase (hexose-6-phosphate mutarotase), likely involved in carbohydrate metabolism; GFP-fusion protein localizes to both the nucleus and cytoplasm and is induced in response to the DNA-damaging agent MMS | NADP(+)-dependent dehydrogenase; acts on serine, L-allo-threonine, and other 3-hydroxy acids; green fluorescent protein fusion protein localizes to the cytoplasm and nucleus; may interact with ribosomes, based on co-purification experiments | GTP-binding protein that regulates the nitrogen starvation response, sporulation, and filamentous growth; farnesylation and palmitoylation required for activity and localization to plasma membrane; homolog of mammalian Ras proto-oncogenes | Heat shock protein regulator that binds to Hsp90p and may stimulate ATPase activity; originally identified as a high-copy number suppressor of a HSP90 loss-of-function mutation; GFP-fusion protein localizes to the cytoplasm and nucleus | Mitochondrial membrane protein that coordinates expression of mitochondrially-encoded genes; may facilitate delivery of mRNA to membrane-bound translation machinery | Ferrochelatase, a mitochondrial inner membrane protein, catalyzes the insertion of ferrous iron into protoporphyrin IX, the eighth and final step in the heme biosynthetic pathway | Protein kinase implicated in activation of the plasma membrane H(+)-ATPase Pma1p in response to glucose metabolism; plays a role in ion homeostasis | ATPase that is a component of the heat shock protein Hsp90 chaperone complex; binds unfolded proteins; member of the heat shock protein 70 (HSP70) family; localized to the cytoplasm | SET-domain lysine-N-methyltransferase, catalyzes the formation of dimethyllysine residues on the large ribsomal subunit protein L23a (RPL23A and RPL23B) |
| 283 | 53 | 440 | 0.408 | 0.588 | 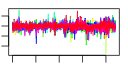 |  | 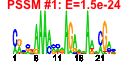 | 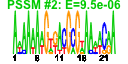 |  | YDL122W YDR110W YJL080C YMR247C YOR098C YAL024C YAR007C YAR008W YBL061C YBR038W YBR130C YBR245C YCL013W YDL093W YDL094C YDL102W YDL103C YDL163W YDL227C YDL229W YDR046C YDR138W YDR146C YDR147W YDR509W YER001W YER164W YGL016W YGL116W YGR140W YHR119W YHR153C YHR154W YHR173C YIL128W YJL018W YJL019W YJL076W YJR030C YLR032W YLR057W YLR084C YML059C YML064C YML104C YMR001C YMR277W YNL029C YNL218W YNL233W YNL289W YOR188W YPR070W | UBP1 FOB1 SCP160 RKR1 NUP1 LTE1 RFA1 SEN34 SKT5 CHS2 SHE3 ISW1 PMT5 POL3 QRI1 HO SSB1 BAP3 HPR1 SWI5 EKI1 MNN1 CHD1 KAP122 CDC20 CBF2 SET1 SPO16 RTT107 MET18 MPS3 NET1 RAD5 RAX2 NTE1 TEM1 MDM1 CDC5 FCP1 KTR5 MGS1 BNI4 PCL1 MSB1 MED1 | Ubiquitin-specific protease that removes ubiquitin from ubiquitinated proteins; cleaves at the C terminus of ubiquitin fusions irrespective of their size; capable of cleaving polyubiquitin chains | Nucleolar protein required for DNA replication fork blocking and recombinational hotspot activities; binds to the replication fork barrier site in the rDNA region; related to retroviral integrases | Essential RNA-binding G protein effector of mating response pathway, mainly associated with nuclear envelope and ER, interacts in mRNA-dependent manner with translating ribosomes via multiple KH domains, similar to vertebrate vigilins | Nuclear RING domain protein with functional connections to chromatin modification; may interact with ribosomes, based on co-purification experiments; YMR247C is not an essential gene | Nuclear pore complex (NPC) subunit, involved in protein import/export and in export of RNAs, possible karyopherin release factor that accelerates release of karyopherin-cargo complexes after transport across NPC; potential Cdc28p substrate | Putative GDP/GTP exchange factor required for mitotic exit at low temperatures; acts as a guanine nucleotide exchange factor (GEF) for Tem1p, which is a key regulator of mitotic exit; physically associates with Ras2p-GTP | Subunit of heterotrimeric Replication Protein A (RPA), which is a highly conserved single-stranded DNA binding protein involved in DNA replication, repair, and recombination | Subunit of the tRNA splicing endonuclease, which is composed of Sen2p, Sen15p, Sen34p, and Sen54p; Sen34p contains the active site for tRNA 3' splice site cleavage and has similarity to Sen2p and to Archaeal tRNA splicing endonuclease | Activator of Chs3p (chitin synthase III), recruits Chs3p to the bud neck via interaction with Bni4p; has similarity to Shc1p, which activates Chs3p during sporulation | Chitin synthase II, requires activation from zymogenic form in order to catalyze the transfer of N-acetylglucosamine (GlcNAc) to chitin; required for the synthesis of chitin in the primary septum during cytokinesis | Protein that acts as an adaptor between Myo4p and the She2p-mRNA complex; part of the mRNA localization machinery that restricts accumulation of certain proteins to the bud; also required for cortical ER inheritance | Member of the imitation-switch (ISWI) class of ATP-dependent chromatin remodeling complexes; ATPase that forms a complex with Ioc2p and Ioc4p to regulate transcription elongation, and a complex with Ioc3p to repress transcription initiation | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; acts in a complex with Pmt3p, can instead interact with Pmt2p in some conditions; target for new antifungals | Catalytic subunit of DNA polymerase delta; required for chromosomal DNA replication during mitosis and meiosis, intragenic recombination, repair of double strand DNA breaks, and DNA replication during nucleotide excision repair (NER) | UDP-N-acetylglucosamine pyrophosphorylase, catalyzes the formation of UDP-N-acetylglucosamine (UDP-GlcNAc), which is important in cell wall biosynthesis, protein N-glycosylation, and GPI anchor biosynthesis | Site-specific endonuclease required for gene conversion at the MAT locus (homothallic switching) through the generation of a ds DNA break; expression restricted to mother cells in late G1 as controlled by Swi4p-Swi6p, Swi5p and Ash1p | Cytoplasmic ATPase that is a ribosome-associated molecular chaperone, functions with J-protein partner Zuo1p; may be involved in folding of newly-made polypeptide chains; member of the HSP70 family; interacts with phosphatase subunit Reg1p | Amino acid permease involved in the uptake of cysteine, leucine, isoleucine and valine | Subunit of THO/TREX complexes that couple transcription elongation with mitotic recombination and with mRNA metabolism and export, subunit of an RNA Pol II complex; regulates lifespan; involved in telomere maintenance; similar to Top1p | Transcription factor that activates transcription of genes expressed at the M/G1 phase boundary and in G1 phase; localization to the nucleus occurs during G1 and appears to be regulated by phosphorylation by Cdc28p kinase | Ethanolamine kinase, primarily responsible for phosphatidylethanolamine synthesis via the CDP-ethanolamine pathway; exhibits some choline kinase activity, thus contributing to phosphatidylcholine synthesis via the CDP-choline pathway | Alpha-1,3-mannosyltransferase, integral membrane glycoprotein of the Golgi complex, required for addition of alpha1,3-mannose linkages to N-linked and O-linked oligosaccharides, one of five S. cerevisiae proteins of the MNN1 family | Nucleosome remodeling factor that functions in regulation of transcription elongation; contains a chromo domain, a helicase domain and a DNA-binding domain; component of both the SAGA and SILK complexes | Karyopherin beta, responsible for import of the Toa1p-Toa2p complex into the nucleus; binds to nucleoporins Nup1p and Nup2p; may play a role in regulation of pleiotropic drug resistance | Cell-cycle regulated activator of anaphase-promoting complex/cyclosome (APC/C), which is required for metaphase/anaphase transition; directs ubiquitination of mitotic cyclins, Pds1p, and other anaphase inhibitors; potential Cdc28p substrate | Essential kinetochore protein, component of the CBF3 multisubunit complex that binds to the CDEIII region of the centromere; Cbf2p also binds to the CDEII region possibly forming a different multimeric complex, ubiquitinated in vivo | Histone methyltransferase, subunit of the COMPASS (Set1C) complex which methylates histone H3 on lysine 4; required in transcriptional silencing near telomeres and at the silent mating type loci; contains a SET domain | Protein of unknown function, required for spore formation | Protein implicated in Mms22-dependent DNA repair during S phase, DNA damage induces phosphorylation by Mec1p at one or more SQ/TQ motifs; interacts with Mms22p and Slx4p; has four BRCT domains; has a role in regulation of Ty1 transposition | DNA repair and TFIIH regulator, required for both nucleotide excision repair (NER) and RNA polymerase II (RNAP II) transcription; involved in telomere maintenance | Essential integral membrane protein required for spindle pole body duplication and for nuclear fusion, localizes to the spindle pole body half bridge, interacts with DnaJ-like chaperone Jem1p and with centrin homolog Cdc31p | Core subunit of the RENT complex, which is a complex involved in nucleolar silencing and telophase exit; stimulates transcription by RNA polymerase I and regulates nucleolar structure | Putative protein of unknown function; expression repressed in carbon limited vs carbon replete chemostat cultures; YJR030C is a non-essential gene | DNA helicase proposed to promote replication fork regression during postreplication repair by template switching; contains RING finger domain | Putative protein of unknown function; YLR050C is not an essential gene | N-glycosylated protein involved in the maintenance of bud site selection during bipolar budding; localization requires Rax1p | Serine esterase that deacylates exogenous lysophospholipids, homolog of human neuropathy target esterase (NTE); mammalian NTE1 deacylates phosphatidylcholine to glycerophosphocholine | GTP-binding protein of the ras superfamily involved in termination of M-phase; controls actomyosin and septin dynamics during cytokinesis | Intermediate filament protein, required for nuclear and mitochondrial transmission to daughter buds; contains a Phox homology (PX) domain and specifically binds phosphatidylinositol 3-phosphate (PtdIns-3-P) | Polo-like kinase with similarity to Xenopus Plx1 and S. pombe Plo1p; found at bud neck, nucleus and SPBs; has multiple functions in mitosis and cytokinesis through phosphorylation of substrates; may be a Cdc28p substrate | Carboxy-terminal domain (CTD) phosphatase, essential for dephosphorylation of the repeated C-terminal domain of the RNA polymerase II large subunit (Rpo21p) | Putative mannosyltransferase involved in protein glycosylation; member of the KRE2/MNT1 mannosyltransferase family | Protein with DNA-dependent ATPase and ssDNA annealing activities involved in maintenance of genome; interacts functionally with DNA polymerase delta; homolog of human Werner helicase interacting protein (WHIP) | Targeting subunit for Glc7p protein phosphatase, localized to the bud neck, required for localization of chitin synthase III to the bud neck via interaction with the chitin synthase III regulatory subunit Skt5p | Pho85 cyclin of the Pcl1,2-like subfamily, involved in entry into the mitotic cell cycle and regulation of morphogenesis, localizes to sites of polarized cell growth | Protein involved in positive regulation of both 1,3-beta-glucan synthesis and the Pkc1p-MAPK pathway, potential Cdc28p substrate; multicopy suppressor of temperature-sensitive mutations in CDC24 and CDC42, and of mutations in BEM4 | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; essential for transcriptional regulation |
| 179 | 46 | 439 | 0.417 | 0.604 | 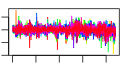 |  | 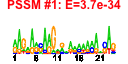 | 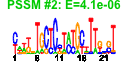 |  | YAL013W YBR059C YCR088W YDR036C YDR388W YGL036W YGL229C YIL153W YIL154C YIR003W YJL108C YJR148W YLL028W YLR132C YLR133W YML127W YMR226C YMR304W YNR049C YOL096C YOR273C YPL024W YAL015C YAL039C YAL060W YCL049C YDL019C YGL023C YGL125W YGL227W YGR097W YGR112W YHR116W YHR161C YKL039W YKL096W YLR337W YNL071W YNL241C YNR006W YNR014W YNR035C YOL067C YOR090C YOR181W YPR047W | DEP1 AKL1 ABP1 EHD3 RVS167 SAP4 RPP2B IMP2' AIM21 PRM10 BAT2 TPO1 CKI1 RSC9 TMA29 UBP15 MSO1 COQ3 TPO4 RMI1 NTG1 CYC3 BDH1 OSH2 PIB2 MET13 VID30 ASK10 SHY1 COX23 YAP1801 PTM1 CWP2 VRP1 LAT1 ZWF1 VPS27 ARC35 RTG1 PTC5 LAS17 AIM30 | Transcriptional modulator involved in regulation of structural phospholipid biosynthesis genes and metabolically unrelated genes, as well as maintenance of telomeres, mating efficiency, and sporulation | Ser-Thr protein kinase, member (with Ark1p and Prk1p) of the Ark kinase family; involved in endocytosis and actin cytoskeleton organization | Actin-binding protein of the cortical actin cytoskeleton, important for activation of the Arp2/3 complex that plays a key role actin in cytoskeleton organization | 3-hydroxyisobutyryl-CoA hydrolase, member of a family of enoyl-CoA hydratase/isomerases; non-tagged protein is detected in highly purified mitochondria in high-throughput studies; phosphorylated; mutation affects fluid-phase endocytosis | Actin-associated protein, subunit of a complex (Rvs161p-Rvs167p) involved in regulation of actin cytoskeleton, endocytosis, and viability following starvation or osmotic stress; homolog of mammalian amphiphysin | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YGL036W is not an essential gene | Protein required for function of the Sit4p protein phosphatase, member of a family of similar proteins that form complexes with Sit4p, including Sap155p, Sap185p, and Sap190p | Ribosomal protein P2 beta, a component of the ribosomal stalk, which is involved in the interaction between translational elongation factors and the ribosome; regulates the accumulation of P1 (Rpp1Ap and Rpp1Bp) in the cytoplasm | Transcriptional activator involved in maintenance of ion homeostasis and protection against DNA damage caused by bleomycin and other oxidants, contains a C-terminal leucine-rich repeat | Protein of unknown function; may interact with ribosomes; GFP-fusion protein colocalizes with Sac1p to the actin cytoskeleton; null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Pheromone-regulated protein, predicted to have 5 transmembrane segments | Cytosolic branched-chain amino acid aminotransferase, homolog of murine ECA39; highly expressed during stationary phase and repressed during logarithmic phase | Polyamine transporter that recognizes spermine, putrescine, and spermidine; catalyzes uptake of polyamines at alkaline pH and excretion at acidic pH; phosphorylation enhances activity and sorting to the plasma membrane | Protein of unknown function; green fluorescent protein (GFP)-tagged protein localizes to mitochondria and nucleus; YLR132C is an essential gene | Choline kinase, catalyzing the first step in phosphatidylcholine synthesis via the CDP-choline (Kennedy pathway); exhibits some ethanolamine kinase activity contributing to phosphatidylethanolamine synthesis via the CDP-ethanolamine pathway | Component of the RSC chromatin remodeling complex; DNA-binding protein involved in the synthesis of rRNA and in transcriptional repression and activation of genes regulated by the Target of Rapamycin (TOR) pathway | NADP(+)-dependent dehydrogenase; acts on serine, L-allo-threonine, and other 3-hydroxy acids; green fluorescent protein fusion protein localizes to the cytoplasm and nucleus; may interact with ribosomes, based on co-purification experiments | Ubiquitin-specific protease that may play a role in ubiquitin precursor processing | Probable component of the secretory vesicle docking complex, acts at a late step in secretion; shows genetic and physical interactions with Sec1p and is enriched in microsomal membrane fractions; required for sporulation | O-methyltransferase, catalyzes two different O-methylation steps in ubiquinone (Coenzyme Q) biosynthesis; component of a mitochondrial ubiquinone-synthesizing complex | Polyamine transport protein, recognizes spermine, putrescine, and spermidine; localizes to the plasma membrane; member of the major facilitator superfamily | Involved in response to DNA damage; null mutants have increased rates of recombination and delayed S phase; interacts physically and genetically with Sgs1p (RecQ family member) and Top3p (topoisomerase III) | DNA N-glycosylase and apurinic/apyrimidinic (AP) lyase involved in base excision repair, localizes to the nucleus and mitochondrion | Cytochrome c heme lyase (holocytochrome c synthase), attaches heme to apo-cytochrome c (Cyc1p or Cyc7p) in the mitochondrial intermembrane space; human ortholog may have a role in microphthalmia with linear skin defects (MLS) | NAD-dependent (R,R)-butanediol dehydrogenase, catalyzes oxidation of (R,R)-2,3-butanediol to (3R)-acetoin, oxidation of meso-butanediol to (3S)-acetoin, and reduction of acetoin; enhances use of 2,3-butanediol as an aerobic carbon source | Protein of unknown function; localizes to membrane fraction; YCL049C is not an essential gene | Member of an oxysterol-binding protein family with seven members in S. cerevisiae; family members have overlapping, redundant functions in sterol metabolism and collectively perform a function essential for viability | Protein binding phosphatidylinositol 3-phosphate, involved in telomere-proximal repression of gene expression; similar to Fab1 and Vps27 | Isozyme of methylenetetrahydrofolate reductase, catalyzes the reduction of 5,10-methylenetetrahydrofolate to 5-methyltetrahydrofolate in the methionine biosynthesis pathway | Protein involved in proteasome-dependent catabolite degradation of fructose-1,6-bisphosphatase (FBPase); shifts the balance of nitrogen metabolism toward the production of glutamate; localized to the nucleus and the cytoplasm | Component of the RNA polymerase II holoenzyme, phosphorylated in response to oxidative stress; has a role in destruction of Ssn8p, which relieves repression of stress-response genes | Mitochondrial inner membrane protein required for assembly of cytochrome c oxidase (complex IV); associates with complex IV assembly intermediates and complex III/complex IV supercomplexes; similar to human SURF1 involved in Leigh Syndrome | Mitochondrial intermembrane space protein that functions in mitochondrial copper homeostasis, essential for functional cytochrome oxidase expression; homologous to Cox17p | Protein involved in clathrin cage assembly; binds Pan1p and clathrin; homologous to Yap1802p, member of the AP180 protein family | Protein of unknown function, copurifies with late Golgi vesicles containing the v-SNARE Tlg2p | Covalently linked cell wall mannoprotein, major constituent of the cell wall; plays a role in stabilizing the cell wall; involved in low pH resistance; precursor is GPI-anchored | Proline-rich actin-associated protein involved in cytoskeletal organization and cytokinesis; related to mammalian Wiskott-Aldrich syndrome protein (WASP)-interacting protein (WIP) | Dihydrolipoamide acetyltransferase component (E2) of pyruvate dehydrogenase complex, which catalyzes the oxidative decarboxylation of pyruvate to acetyl-CoA | Glucose-6-phosphate dehydrogenase (G6PD), catalyzes the first step of the pentose phosphate pathway; involved in adapting to oxidatve stress; homolog of the human G6PD which is deficient in patients with hemolytic anemia | Endosomal protein that forms a complex with Hse1p; required for recycling Golgi proteins, forming lumenal membranes and sorting ubiquitinated proteins destined for degradation; has Ubiquitin Interaction Motifs which bind ubiquitin (Ubi4p) | Putative protein of unknown function; expression is cell-cycle regulated, Azf1p-dependent, and heat-inducible | Subunit of the ARP2/3 complex, which is required for the motility and integrity of cortical actin patches; required for cortical localization of calmodulin | Transcription factor (bHLH) involved in interorganelle communication between mitochondria, peroxisomes, and nucleus | Mitochondrially localized type 2C protein phosphatase involved in regulation of pyruvate dehydrogenase activity; contains Mg2+/Mn2+-dependent casein phosphatase activity in vitro | Actin assembly factor, activates the Arp2/3 protein complex that nucleates branched actin filaments; localizes with the Arp2/3 complex to actin patches; homolog of the human Wiskott-Aldrich syndrome protein (WASP) | Putative protein of unknown function that may be involved in intramitochondrial sorting; similar to Ups1p and to human PRELI; GFP-tagged protein localizes to mitochondria; required for wild-type respiratory growth |
| 161 | 49 | 442 | 0.429 | 0.503 |  |  | 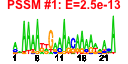 |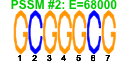  |  | YBL088C YDR102C YGR293C YHL037C YIL037C YJL182C YML047C YOL160W YAL064W YBL013W YBR040W YBR148W YCL066W YCL074W YCL075W YCR040W YDL114W YDL162C YDR009W YDR095C YDR215C YDR278C YDR446W YDR523C YDR525W YEL010W YER181C YFL003C YFL012W YFL050C YFR012W YFR057W YGL024W YGL118C YGL170C YGL262W YGR269W YGR273C YGR291C YHL041W YHR015W YHR126C YHR185C YIL174W YLR365W YOL166C YOR183W YOR268C YPL192C | TEL1 PRM2 PRM6 YAL064W-B FMT1 FIG1 YSW1 HMLALPHA1 GAL3 ECM11 SPS1 SNA2 MSH4 ALR2 SPO74 MIP6 PFS1 FYV12 PRM3 | Protein kinase primarily involved in telomere length regulation; contributes to cell cycle checkpoint control in response to DNA damage; functionally redundant with Mec1p; homolog of human ataxia telangiectasia (ATM) gene | Pheromone-regulated protein, predicted to have 4 transmembrane segments and a coiled coil domain; regulated by Ste12p | Pheromone-regulated protein, predicted to have 2 transmembrane segments; regulated by Ste12p during mating | Fungal-specific protein of unknown function | Methionyl-tRNA formyltransferase, catalyzes the formylation of initiator Met-tRNA in mitochondria; potential Cdc28p substrate | Integral membrane protein required for efficient mating; may participate in or regulate the low affinity Ca2+ influx system, which affects intracellular signaling and cell-cell fusion during mating | Protein expressed specifically in spores | Silenced copy of ALPHA1 at HML, encoding a transcriptional coactivator involved in the regulation of mating-type alpha-specific gene expression | Putative protein of unknown function with similarity to acyl-carrier-protein reductases; YDL114W is not an essential gene | Transcriptional regulator involved in activation of the GAL genes in response to galactose; forms a complex with Gal80p to relieve Gal80p inhibition of Gal4p; binds galactose and ATP but does not have galactokinase activity | Non-essential protein apparently involved in meiosis, GFP fusion protein is present in discrete clusters in the nucleus throughout mitosis; may be involved in maintaining chromatin structure | Putative protein serine/threonine kinase expressed at the end of meiosis and localized to the prospore membrane, required for correct localization of enzymes involved in spore wall synthesis | Protein of unknown function, has similarity to Pmp3p, which is involved in cation transport; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Protein involved in meiotic recombination, required for normal levels of crossing over, colocalizes with Zip2p to discrete foci on meiotic chromosomes, has homology to bacterial MutS protein | Putative protein of unknown function; transcribed during sporulation; null mutant exhibits increased resistance to rapamycin | Probable Mg(2+) transporter; overexpression confers increased tolerance to Al(3+) and Ga(3+) ions | Putative protein of unknown function | Component of the meiotic outer plaque of the spindle pole body, involved in modifying the meiotic outer plaque that is required prior to prospore membrane formation | Putative protein of unknown function; null mutant displays elevated sensitivity to expression of a mutant huntingtin fragment or of alpha-synuclein; YGL262W is not an essential gene | Putative protein of unknown function; deletion mutant has no readily detectable phenotype | Putative RNA-binding protein, interacts with Mex67p, which is a component of the nuclear pore involved in nuclear mRNA export | Putative protein of unknown function; transcription dependent upon Azf1p | Sporulation protein required for prospore membrane formation at selected spindle poles, ensures functionality of all four spindle pole bodies during meiosis II; not required for meiotic recombination or meiotic chromosome segregation | Protein of unknown function, required for survival upon exposure to K1 killer toxin | Putative protein of unknown function; sporulation is abnormal in homozygous diploid; YOR268C is not an essential gene | Pheromone-regulated protein required for karyogamy; localizes to the inner membrane of the nuclear envelope |
| 119 | 49 | 447 | 0.443 | 0.569 | 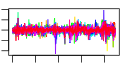 |  | 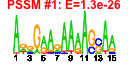 | 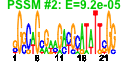 |  | YBR008C YCR025C YCR105W YDR160W YER011W YER076C YFL003C YGL046W YHR156C YIR020C YJL219W YJR146W YKL031W YKL097C YKR040C YLR054C YLR111W YLR137W YLR255C YLR296W YML037C YMR042W YMR106C YMR213W YMR306W YOR013W YOR042W YOR140W YOR181W YPL027W YPL056C YPL124W YPR085C YCR048W YCR050C YDL035C YDR501W YER018C YER019W YGL045W YJL220W YLR393W YLR402W YML089C YNL187W YOL156W YOR029W YPL058C YPR077C | FLR1 ADH7 SSY1 TIR1 MSH4 RIM8 LIN1 HXT9 OSW2 ARG80 YKU80 CEF1 FKS3 CUE5 SFL1 LAS17 SMA1 SPC29 ARE1 GPR1 PLM2 SPC25 ISC1 ATP10 HXT11 PDR12 | Plasma membrane multidrug transporter of the major facilitator superfamily, involved in efflux of fluconazole, diazaborine, benomyl, methotrexate, and other drugs | NADPH-dependent medium chain alcohol dehydrogenase with broad substrate specificity; member of the cinnamyl family of alcohol dehydrogenases; may be involved in fusel alcohol synthesis or in aldehyde tolerance | Component of the SPS plasma membrane amino acid sensor system (Ssy1p-Ptr3p-Ssy5p), which senses external amino acid concentration and transmits intracellular signals that result in regulation of expression of amino acid permease genes | Cell wall mannoprotein of the Srp1p/Tip1p family of serine-alanine-rich proteins; expression is downregulated at acidic pH and induced by cold shock and anaerobiosis; abundance is increased in cells cultured without shaking | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; analysis of HA-tagged protein suggests a membrane localization | Protein involved in meiotic recombination, required for normal levels of crossing over, colocalizes with Zip2p to discrete foci on meiotic chromosomes, has homology to bacterial MutS protein | Protein of unknown function, involved in the proteolytic activation of Rim101p in response to alkaline pH; has similarity to A. nidulans PalF | Non-essential component of U5 snRNP; nuclear protein; physically interacts with Irr1p of cohesin complex; may link together proteins involved in chromosome segregation, mRNA splicing and DNA replication | Putative hexose transporter that is nearly identical to Hxt11p, has similarity to major facilitator superfamily (MFS) transporters, expression of HXT9 is regulated by transcription factors Pdr1p and Pdr3p | Protein of unknown function proposed to be involved in the assembly of the spore wall | Putative protein of unknown function | Putative protein of unknown function with some characteristics of a transcriptional activator; may be a target of Dbf2p-Mob1p kinase; GFP-fusion protein co-localizes with clathrin-coated vesicles; YML037C is not an essential gene | Transcription factor involved in regulation of arginine-responsive genes; acts with Arg81p and Arg82p | Subunit of the telomeric Ku complex (Yku70p-Yku80p), involved in telomere length maintenance, structure and telomere position effect; relocates to sites of double-strand cleavage to promote nonhomologous end joining during DSB repair | Essential splicing factor; associated with Prp19p and the spliceosome, contains an N-terminal c-Myb DNA binding motif necessary for cell viability but not for Prp19p association, evolutionarily conserved and homologous to S. pombe Cdc5p | Protein involved in spore wall assembly, has similarity to 1,3-beta-D-glucan synthase catalytic subunits Fks1p and Gsc2p; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein containing a CUE domain that binds ubiquitin, which may facilitate intramolecular monoubiquitination; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Transcriptional repressor and activator; involved in repression of flocculation-related genes, and activation of stress responsive genes; negatively regulated by cAMP-dependent protein kinase A subunit Tpk2p | Actin assembly factor, activates the Arp2/3 protein complex that nucleates branched actin filaments; localizes with the Arp2/3 complex to actin patches; homolog of the human Wiskott-Aldrich syndrome protein (WASP) | Protein of unknown function involved in the assembly of the prospore membrane during sporulation | Putative protein of unknown function; deletion mutant is fluconazole resistant | Inner plaque spindle pole body (SPB) component, links the central plaque component Spc42p to the inner plaque component Spc110p; required for SPB duplication | Putative protein of unknown function; YPR085C is an essential gene | Acyl-CoA:sterol acyltransferase, isozyme of Are2p; endoplasmic reticulum enzyme that contributes the major sterol esterification activity in the absence of oxygen | Non-essential protein of unknown function; deletion mutant is synthetically sick or lethal with alpha-synuclein | Plasma membrane G protein coupled receptor (GPCR) that interacts with the heterotrimeric G protein alpha subunit, Gpa2p, and with Plc1p; sensor that integrates nutritional signals with the modulation of cell fate via PKA and cAMP synthesis | Protein required for partitioning of the 2-micron plasmid | Component of the evolutionarily conserved kinetochore-associated Ndc80 complex (Ndc80p-Nuf2p-Spc24p-Spc25p); involved in chromosome segregation, spindle checkpoint activity and kinetochore clustering | Mitochondrial membrane localized inositol phosphosphingolipid phospholipase C, hydrolyzes complex sphingolipids to produce ceramide; activated by phosphatidylserine, cardiolipin, and phosphatidylglycerol; mediates Na+ and Li+ halotolerance | Mitochondrial inner membrane protein required for assembly of the F0 sector of mitochondrial F1F0 ATP synthase, interacts genetically with ATP6 | Putative protein of unknown function; contains a consensus nuclear export signal (NES) sequence similar to the consensus sequence recognized by Crm1p; may interact with or be fuctionally redundant with Prp40p | Putative hexose transporter that is nearly identical to Hxt9p, has similarity to major facilitator superfamily (MFS) transporters and is involved in pleiotropic drug resistance | Plasma membrane ATP-binding cassette (ABC) transporter, weak-acid-inducible multidrug transporter required for weak organic acid resistance; induced by sorbate and benzoate and regulated by War1p; mutants exhibit sorbate hypersensitivity |
| 109 | 35 | 446 | 0.461 | 0.551 | 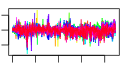 |  |  |  |  | YAR015W YAR073W YBR248C YCL030C YDR019C YDR226W YGL186C YGL234W YGL253W YGR208W YIL094C YLR180W YLR359W YMR189W YMR241W YNR050C YOR184W YOR254C YOR375C YAL044C YER056C YER091C YGL077C YGR061C YGR204W YKL029C YKR080W YLR058C YMR120C YMR243C YMR300C YNL220W YOR128C YOR335C YPR004C | ADE1 HIS7 HIS4 GCV1 ADK1 TPN1 ADE5,7 HXK2 SER2 LYS12 SAM1 ADE13 GCV2 YHM2 LYS9 SER1 SEC63 GDH1 GCV3 FCY2 MET6 HNM1 ADE6 ADE3 MAE1 MTD1 SHM2 ADE17 ZRC1 ADE4 ADE12 ADE2 ALA1 AIM45 | N-succinyl-5-aminoimidazole-4-carboxamide ribotide (SAICAR) synthetase, required for 'de novo' purine nucleotide biosynthesis; red pigment accumulates in mutant cells deprived of adenine | Imidazole glycerol phosphate synthase (glutamine amidotransferase:cyclase), catalyzes the fifth and sixth steps of histidine biosynthesis and also produces 5-aminoimidazole-4-carboxamide ribotide (AICAR), a purine precursor | Multifunctional enzyme containing phosphoribosyl-ATP pyrophosphatase, phosphoribosyl-AMP cyclohydrolase, and histidinol dehydrogenase activities; catalyzes the second, third, ninth and tenth steps in histidine biosynthesis | T subunit of the mitochondrial glycine decarboxylase complex, required for the catabolism of glycine to 5,10-methylene-THF; expression is regulated by levels of levels of 5,10-methylene-THF in the cytoplasm | Adenylate kinase, required for purine metabolism; localized to the cytoplasm and the mitochondria; lacks cleavable signal sequence | Plasma membrane pyridoxine (vitamin B6) transporter; member of the purine-cytosine permease subfamily within the major facilitator superfamily; proton symporter with similarity to Fcy21p, Fcy2p, and Fcy22p | Bifunctional enzyme of the 'de novo' purine nucleotide biosynthetic pathway, contains aminoimidazole ribotide synthetase and glycinamide ribotide synthetase activities | Hexokinase isoenzyme 2 that catalyzes phosphorylation of glucose in the cytosol; predominant hexokinase during growth on glucose; functions in the nucleus to repress expression of HXK1 and GLK1 and to induce expression of its own gene | Phosphoserine phosphatase of the phosphoglycerate pathway, involved in serine and glycine biosynthesis, expression is regulated by the available nitrogen source | Homo-isocitrate dehydrogenase, an NAD-linked mitochondrial enzyme required for the fourth step in the biosynthesis of lysine, in which homo-isocitrate is oxidatively decarboxylated to alpha-ketoadipate | S-adenosylmethionine synthetase, catalyzes transfer of the adenosyl group of ATP to the sulfur atom of methionine; one of two differentially regulated isozymes (Sam1p and Sam2p) | Adenylosuccinate lyase, catalyzes two steps in the 'de novo' purine nucleotide biosynthetic pathway; expression is repressed by adenine and activated by Bas1p and Pho2p; mutations in human ortholog ADSL cause adenylosuccinase deficiency | P subunit of the mitochondrial glycine decarboxylase complex, required for the catabolism of glycine to 5,10-methylene-THF; expression is regulated by levels of 5,10-methylene-THF in the cytoplasm | Mitochondrial DNA-binding protein, component of the mitochondrial nucleoid structure, involved in mtDNA replication and segregation of mitochondrial genomes; member of the mitochondrial carrier protein family | Saccharopine dehydrogenase (NADP+, L-glutamate-forming); catalyzes the formation of saccharopine from alpha-aminoadipate 6-semialdehyde, which is the seventh step in lysine biosynthesis pathway | 3-phosphoserine aminotransferase, catalyzes the formation of phosphoserine from 3-phosphohydroxypyruvate, required for serine and glycine biosynthesis; regulated by the general control of amino acid biosynthesis mediated by Gcn4p | Essential subunit of Sec63 complex (Sec63p, Sec62p, Sec66p and Sec72p); with Sec61 complex, Kar2p/BiP and Lhs1p forms a channel competent for SRP-dependent and post-translational SRP-independent protein targeting and import into the ER | NADP(+)-dependent glutamate dehydrogenase, synthesizes glutamate from ammonia and alpha-ketoglutarate; rate of alpha-ketoglutarate utilization differs from Gdh3p; expression regulated by nitrogen and carbon sources | H subunit of the mitochondrial glycine decarboxylase complex, required for the catabolism of glycine to 5,10-methylene-THF; expression is regulated by levels of levels of 5,10-methylene-THF in the cytoplasm | Purine-cytosine permease, mediates purine (adenine, guanine, and hypoxanthine) and cytosine accumulation | Cobalamin-independent methionine synthase, involved in amino acid biosynthesis; requires a minimum of two glutamates on the methyltetrahydrofolate substrate, similar to bacterial metE homologs | Choline/ethanolamine transporter (permease) that also controls the uptake of nitrogen mustard; expression is co-regulated with phospholipid biosynthetic genes and negatively regulated by choline and myo-inositol | Formylglycinamidine-ribonucleotide (FGAM)-synthetase, catalyzes a step in the 'de novo' purine nucleotide biosynthetic pathway | Cytoplasmic trifunctional enzyme C1-tetrahydrofolate synthase, involved in single carbon metabolism and required for biosynthesis of purines, thymidylate, methionine, and histidine | Mitochondrial malic enzyme, catalyzes the oxidative decarboxylation of malate to pyruvate, which is a key intermediate in sugar metabolism and a precursor for synthesis of several amino acids | NAD-dependent 5,10-methylenetetrahydrafolate dehydrogenase, plays a catalytic role in oxidation of cytoplasmic one-carbon units; expression is regulated by Bas1p and Bas2p, repressed by adenine, and may be induced by inositol and choline | Cytosolic serine hydroxymethyltransferase, converts serine to glycine plus 5,10 methylenetetrahydrofolate; major isoform involved in generating precursors for purine, pyrimidine, amino acid, and lipid biosynthesis | Enzyme of 'de novo' purine biosynthesis containing both 5-aminoimidazole-4-carboxamide ribonucleotide transformylase and inosine monophosphate cyclohydrolase activities, isozyme of Ade16p; ade16 ade17 mutants require adenine and histidine | Vacuolar membrane zinc transporter, transports zinc from the cytosol into the vacuole for storage; also has a role in resistance to zinc shock resulting from a sudden influx of zinc into the cytoplasm | Phosphoribosylpyrophosphate amidotransferase (PRPPAT; amidophosphoribosyltransferase), catalyzes first step of the 'de novo' purine nucleotide biosynthetic pathway | Adenylosuccinate synthase, catalyzes the first step in synthesis of adenosine monophosphate from inosine 5'monophosphate during purine nucleotide biosynthesis; exhibits binding to single-stranded autonomously replicating (ARS) core sequence | Phosphoribosylaminoimidazole carboxylase, catalyzes a step in the 'de novo' purine nucleotide biosynthetic pathway; red pigment accumulates in mutant cells deprived of adenine | Cytoplasmic alanyl-tRNA synthetase, required for protein synthesis; point mutation (cdc64-1 allele) causes cell cycle arrest at G1; lethality of null mutation is functionally complemented by human homolog | Protein with similarity to mammalian electron transfer flavoprotein complex subunit ETF-alpha; interacts with yeast frataxin Yfh1p; null mutant displays increased frequency of mitochondrial genome loss (petite formation) |
| 129 | 49 | 440 | 0.478 | 0.555 |  |  |  | 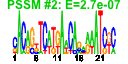 |  | YAL024C YAR035W YBR020W YBR040W YBR213W YCL001W YCR106W YFL011W YIL100W YIR019C YLR081W YLR087C YLR382C YNR056C YNR057C YNR058W YNR074C YOR030W YOR373W YPL006W YPL033C YPL121C YPR152C YPR153W YAL026C YAL048C YBR019C YHR164C YIL072W YIL073C YJL113W YKR104W YLR278C YLR308W YLR318W YML017W YMR280C YMR306W YOR366W YOR381W YOR388C YPL073C YPL110C YPL167C YPL174C YPL180W YPR095C YPR115W YPR157W | LTE1 YAT1 GAL1 FIG1 MET8 RER1 RDS1 HXT10 MUC1 GAL2 CSF1 MSL1 BIO5 BIO4 BIO3 CPD1 MET30 TRM2 NCR1 MEI5 URN1 DRS2 GEM1 GAL10 DNA2 HOP1 SPO22 NFT1 CDA2 EST2 PSP2 CAT8 FKS3 FRE3 FDH1 GDE1 REV3 NIP100 TCO89 SYT1 | Putative GDP/GTP exchange factor required for mitotic exit at low temperatures; acts as a guanine nucleotide exchange factor (GEF) for Tem1p, which is a key regulator of mitotic exit; physically associates with Ras2p-GTP | Outer mitochondrial carnitine acetyltransferase, minor ethanol-inducible enzyme involved in transport of activated acyl groups from the cytoplasm into the mitochondrial matrix; phosphorylated | Galactokinase, phosphorylates alpha-D-galactose to alpha-D-galactose-1-phosphate in the first step of galactose catabolism; expression regulated by Gal4p | Integral membrane protein required for efficient mating; may participate in or regulate the low affinity Ca2+ influx system, which affects intracellular signaling and cell-cell fusion during mating | Bifunctional dehydrogenase and ferrochelatase, involved in the biosynthesis of siroheme, a prosthetic group used by sulfite reductase; required for sulfate assimilation and methionine biosynthesis | Protein involved in retention of membrane proteins, including Sec12p, in the ER; localized to Golgi; functions as a retrieval receptor in returning membrane proteins to the ER | Zinc cluster transcription factor involved in conferring resistance to cycloheximide | Putative hexose transporter, expressed at low levels and expression is repressed by glucose | GPI-anchored cell surface glycoprotein (flocculin) required for pseudohyphal formation, invasive growth, flocculation, and biofilms; transcriptionally regulated by the MAPK pathway (via Ste12p and Tec1p) and the cAMP pathway (via Flo8p) | Galactose permease, required for utilization of galactose; also able to transport glucose | Protein required for fermentation at low temperature; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | U2B component of U2 snRNP, involved in splicing, binds the U2 snRNA stem-loop IV in vitro; does not contain the conserved C-terminal RNA binding domain found in other family members | Putative transmembrane protein involved in the biotin biosynthesis pathway; responsible for uptake of 7-keto 8-aminopelargonic acid; BIO5 is in a cluster of 3 genes (BIO3, BIO4, and BIO5) that mediate biotin synthesis | Dethiobiotin synthetase, catalyzes the third step in the biotin biosynthesis pathway; BIO4 is in a cluster of 3 genes (BIO3, BIO4, and BIO5) that mediate biotin synthesis; expression appears to be repressed at low iron levels | 7,8-diamino-pelargonic acid aminotransferase (DAPA), catalyzes the second step in the biotin biosynthesis pathway; BIO3 is in a cluster of 3 genes (BIO3, BIO4, and BIO5) that mediate biotin synthesis | Cyclic nucleotide phosphodiesterase, hydrolyzes ADP-ribose 1'', 2''-cyclic phosphate to ADP-ribose 1''-phosphate; may have a role in tRNA splicing; no detectable phenotype is conferred by null mutation or by overexpression | F-box protein containing five copies of the WD40 motif, controls cell cycle function, sulfur metabolism, and methionine biosynthesis as part of the ubiquitin ligase complex; interacts with and regulates Met4p, localizes within the nucleus | tRNA methyltransferase, 5-methylates the uridine residue at position 54 of tRNAs and may also have a role in tRNA stabilization or maturation; endo-exonuclease with a role in DNA repair | Vacuolar membrane protein that transits through the biosynthetic vacuolar protein sorting pathway, involved in sphingolipid metabolism; glycoprotein and functional orthologue of human Niemann Pick C1 (NPC1) protein | Putative protein of unknown function; may be involved in DNA metabolism; expression is induced by Kar4p | Meiosis specific protein involved in DMC1-dependent meiotic recombination, forms heterodimer with Sae3p; proposed to be an assembly factor for Dmc1p | Pre-mRNA splicing factor associated with the U2-U5-U6 snRNPs, the RES complex, and the Prp19-associated complex (NTC); null mutation displays synthetic genetic interactions with mutations affecting U2 snRNA and pre-mRNA splicing factors | Putative protein of unknown function | Aminophospholipid translocase (flippase) that maintains membrane lipid asymmetry in post-Golgi secretory vessicles; contributes to clathrin-coated vesicle formation and endocytosis; mutations in human homolog ATP8B1 result in liver disease | Evolutionarily-conserved tail-anchored outer mitochondrial membrane GTPase which regulates mitochondrial morphology; cells lacking Gem1p contain collapsed, globular, or grape-like mitochondria; not required for pheromone-induced cell death | UDP-glucose-4-epimerase, catalyzes the interconversion of UDP-galactose and UDP-D-glucose in galactose metabolism; also catalyzes the conversion of alpha-D-glucose or alpha-D-galactose to their beta-anomers | Essential tripartite DNA replication factor with single-stranded DNA-dependent ATPase, ATP-dependent nuclease, and helicase activities; required for Okazaki fragment processing; involved in DNA repair pathways; potential Cdc28p substrate | Meiosis-specific DNA binding protein that displays Red1p dependent localization to the unsynapsed axial-lateral elements of the synaptonemal complex; required for homologous chromosome synapsis and chiasma formation | Meiosis-specific protein essential for chromosome synapsis, similar to phospholipase A2, involved in completion of nuclear divisions during meiosis; induced early in meiosis | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Putative transporter of the multidrug resistance-associated protein (MRP) subfamily; adjacent ORFs YKR103W and YKR104W are merged in different strain backgrounds. | Zinc-cluster protein; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; mutant shows moderate growth defect on caffeine; YLR278C is not an essential gene | Chitin deacetylase, together with Cda1p involved in the biosynthesis ascospore wall component, chitosan; required for proper rigidity of the ascospore wall | Reverse transcriptase subunit of the telomerase holoenzyme, essential for telomerase core catalytic activity, involved in other aspects of telomerase assembly and function; mutations in human homolog are associated with aplastic anemia | Asn rich cytoplasmic protein that contains RGG motifs; high-copy suppressor of group II intron-splicing defects of a mutation in MRS2 and of a conditional mutation in POL1 (DNA polymerase alpha); possible role in mitochondrial mRNA splicing | Zinc cluster transcriptional activator necessary for derepression of a variety of genes under non-fermentative growth conditions, active after diauxic shift, binds carbon source responsive elements | Protein involved in spore wall assembly, has similarity to 1,3-beta-D-glucan synthase catalytic subunits Fks1p and Gsc2p; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Ferric reductase, reduces siderophore-bound iron prior to uptake by transporters; expression induced by low iron levels | NAD(+)-dependent formate dehydrogenase, may protect cells from exogenous formate | Glycerophosphocholine (GroPCho) phosphodiesterase; hydrolyzes GroPCho to choline and glycerolphosphate, for use as a phosphate source and as a precursor for phosphocholine synthesis; may interact with ribosomes | Catalytic subunit of DNA polymerase zeta, which is involved in DNA repair and translesion synthesis; required for mutagenesis induced by DNA damage | Large subunit of the dynactin complex, which is involved in partitioning the mitotic spindle between mother and daughter cells; putative ortholog of mammalian p150(glued) | Subunit of TORC1 (Tor1p or Tor2p-Kog1p-Lst8p-Tco89p), a complex that regulates growth in response to nutrient availability; cooperates with Ssd1p in the maintenance of cellular integrity; deletion strains are hypersensitive to rapamycin | Guanine nucleotide exchange factor (GEF) for Arf proteins; involved in vesicular transport; suppressor of ypt3 mutations; member of the Sec7-domain family | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm |
| 206 | 40 | 437 | 0.484 | 0.580 |  |  | 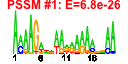 | 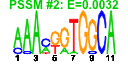 |  | YAL035W YBR011C YDL100C YFL022C YGL148W YGL238W YGR155W YKL154W YLR179C YML008C YOR202W YAL012W YBR281C YCL043C YCL050C YCL057W YDR502C YEL071W YFR009W YGR055W YGR124W YHR179W YIL074C YIL118W YJR010W YKL001C YKL054C YKL073W YKL212W YLL062C YLR044C YLR134W YLR303W YML125C YMR011W YMR049C YMR318C YOR198C YPL061W YPR080W | FUN12 IPP1 GET3 FRS2 ARO2 CSE1 CYS4 SRP102 ERG6 HIS3 CYS3 DUG2 PDI1 APA1 PRD1 SAM2 DLD3 GCN20 MUP1 ASN2 OYE2 SER33 RHO3 MET3 MET14 DEF1 LHS1 SAC1 MHT1 PDC1 PDC5 MET17 PGA3 HXT2 ERB1 ADH6 BFR1 ALD6 TEF1 | GTPase, required for general translation initiation by promoting Met-tRNAiMet binding to ribosomes and ribosomal subunit joining; homolog of bacterial IF2 | Cytoplasmic inorganic pyrophosphatase (PPase), catalyzes the rapid exchange of oxygens from Pi with water, highly expressed and essential for viability, active-site residues show identity to those from E. coli PPase | ATPase, subunit of the GET complex; required for the retrieval of HDEL proteins from the Golgi to the ER in an ERD2 dependent fashion; involved in resistance to heat and metal stress | Alpha subunit of cytoplasmic phenylalanyl-tRNA synthetase, forms a tetramer with Frs1p to form active enzyme; evolutionarily distant from mitochondrial phenylalanyl-tRNA synthetase based on protein sequence, but substrate binding is similar | Bifunctional chorismate synthase and flavin reductase, catalyzes the conversion of 5-enolpyruvylshikimate 3-phosphate (EPSP) to form chorismate, which is a precursor to aromatic amino acids | Nuclear envelope protein that mediates the nuclear export of importin alpha (Srp1p), homolog of metazoan CAS protein, required for accurate chromosome segregation | Cystathionine beta-synthase, catalyzes the synthesis of cystathionine from serine and homocysteine, the first committed step in cysteine biosynthesis | Signal recognition particle (SRP) receptor beta subunit; involved in SRP-dependent protein targeting; anchors Srp101p to the ER membrane | Protein of unknown function, transcription is activated by paralogous proteins Yrm1p and Yrr1p along with proteins involved in multidrug resistance; GFP-tagged protein localizes to the cytoplasm and nucleus; YLR179C is not essential | Delta(24)-sterol C-methyltransferase, converts zymosterol to fecosterol in the ergosterol biosynthetic pathway by methylating position C-24; localized to both lipid particles and mitochondrial outer membrane | Imidazoleglycerol-phosphate dehydratase, catalyzes the sixth step in histidine biosynthesis; mutations cause histidine auxotrophy and sensitivity to Cu, Co, and Ni salts; transcription is regulated by general amino acid control via Gcn4p | Cystathionine gamma-lyase, catalyzes one of the two reactions involved in the transsulfuration pathway that yields cysteine from homocysteine with the intermediary formation of cystathionine | Probable di- and tri-peptidase; forms a complex with Dug1p and Dug3p to degrade glutathione (GSH) and other peptides containing a gamma-glu-X bond in an alternative pathway to GSH degradation by gamma-glutamyl transpeptidase (Ecm38p) | Protein disulfide isomerase, multifunctional protein resident in the endoplasmic reticulum lumen, essential for the formation of disulfide bonds in secretory and cell-surface proteins, unscrambles non-native disulfide bonds | Diadenosine 5',5''-P1,P4-tetraphosphate phosphorylase I (AP4A phosphorylase), involved in catabolism of bis(5'-nucleosidyl) tetraphosphates; has similarity to Apa2p | Zinc metalloendopeptidase, found in the cytoplasm and intermembrane space of mitochondria; with Cym1p, involved in degradation of mitochondrial proteins and of presequence peptides cleaved from imported proteins | S-adenosylmethionine synthetase, catalyzes transfer of the adenosyl group of ATP to the sulfur atom of methionine; one of two differentially regulated isozymes (Sam1p and Sam2p) | D-lactate dehydrogenase, part of the retrograde regulon which consists of genes whose expression is stimulated by damage to mitochondria and reduced in cells grown with glutamate as the sole nitrogen source, located in the cytoplasm | Positive regulator of the Gcn2p kinase activity, forms a complex with Gcn1p; proposed to stimulate Gcn2p activation by an uncharged tRNA | High affinity methionine permease, integral membrane protein with 13 putative membrane-spanning regions; also involved in cysteine uptake | Asparagine synthetase, isozyme of Asn1p; catalyzes the synthesis of L-asparagine from L-aspartate in the asparagine biosynthetic pathway | Widely conserved NADPH oxidoreductase containing flavin mononucleotide (FMN), homologous to Oye3p with slight differences in ligand binding and catalytic properties; may be involved in sterol metabolism | 3-phosphoglycerate dehydrogenase, catalyzes the first step in serine and glycine biosynthesis; isozyme of Ser3p | Non-essential small GTPase of the Rho/Rac subfamily of Ras-like proteins involved in the establishment of cell polarity; GTPase activity positively regulated by the GTPase activating protein (GAP) Rgd1p | ATP sulfurylase, catalyzes the primary step of intracellular sulfate activation, essential for assimilatory reduction of sulfate to sulfide, involved in methionine metabolism | Adenylylsulfate kinase, required for sulfate assimilation and involved in methionine metabolism | RNAPII degradation factor, forms a complex with Rad26p in chromatin, enables ubiquitination and proteolysis of RNAPII present in an elongation complex | Molecular chaperone of the endoplasmic reticulum lumen, involved in polypeptide translocation and folding; member of the Hsp70 family; localizes to the lumen of the ER; regulated by the unfolded protein response pathway | Phosphatidylinositol (PI) phosphatase, involved in hydrolysis of PI 3-phosphate, PI 4-phosphate and PI 3,5-bisphosphate to PI; membrane protein localizes to ER and Golgi; involved in protein trafficking, secretion and inositol metabolism | S-methylmethionine-homocysteine methyltransferase, functions along with Sam4p in the conversion of S-adenosylmethionine (AdoMet) to methionine to control the methionine/AdoMet ratio | Major of three pyruvate decarboxylase isozymes, key enzyme in alcoholic fermentation, decarboxylates pyruvate to acetaldehyde; subject to glucose-, ethanol-, and autoregulation; involved in amino acid catabolism | Minor isoform of pyruvate decarboxylase, key enzyme in alcoholic fermentation, decarboxylates pyruvate to acetaldehyde, regulation is glucose- and ethanol-dependent, repressed by thiamine, involved in amino acid catabolism | Methionine and cysteine synthase (O-acetyl homoserine-O-acetyl serine sulfhydrylase), required for sulfur amino acid synthesis | Essential protein required for maturation of Gas1p and Pho8p, protein trafficking; GFP-fusion protein localizes to the endoplasmic reticulum; null mutants have a cell separation defect | High-affinity glucose transporter of the major facilitator superfamily, expression is induced by low levels of glucose and repressed by high levels of glucose | Protein required for maturation of the 25S and 5.8S ribosomal RNAs; constituent of 66S pre-ribosomal particles; homologous to mammalian Bop1 | NADPH-dependent medium chain alcohol dehydrogenase with broad substrate specificity; member of the cinnamyl family of alcohol dehydrogenases; may be involved in fusel alcohol synthesis or in aldehyde tolerance | Component of mRNP complexes associated with polyribosomes; implicated in secretion and nuclear segregation; multicopy suppressor of BFA (Brefeldin A) sensitivity | Cytosolic aldehyde dehydrogenase, activated by Mg2+ and utilizes NADP+ as the preferred coenzyme; required for conversion of acetaldehyde to acetate; constitutively expressed; locates to the mitochondrial outer surface upon oxidative stress | Translational elongation factor EF-1 alpha; also encoded by TEF2; functions in the binding reaction of aminoacyl-tRNA (AA-tRNA) to ribosomes |
| 17 | 44 | 446 | 0.488 | 0.575 |  |  | 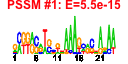 | 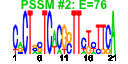 |  | YAL023C YBR029C YBR220C YCL014W YCL046W YDL095W YDL096C YDL189W YDR119W YDR451C YDR514C YDR531W YER156C YFR028C YGL094C YGL195W YGR198W YGR229C YGR233C YHR031C YHR098C YHR118C YHR119W YHR133C YHR186C YHR204W YIL009W YIL128W YIL129C YIL140W YJL076W YJL135W YKL106W YLR215C YMR006C YMR129W YOR087W YOR188W YOR342C YPL141C YPL144W YPL158C YPL242C YPR069C | PMT2 CDS1 BUD3 PMT1 RBS1 YHP1 CDC14 PAN2 GCN1 YPP1 SMI1 PHO81 RRM3 SFB3 ORC6 SET1 NSG1 KOG1 MNL1 FAA3 MET18 TAO3 AXL2 NET1 AAT1 CDC123 PLB2 POM152 YVC1 MSB1 POC4 AIM44 IQG1 SPE3 | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; acts in a complex with Pmt1p, can instead interact with Pmt5p in some conditions; target for new antifungals | Phosphatidate cytidylyltransferase (CDP-diglyceride synthetase); an enzyme that catalyzes that conversion of CTP + phosphate into diphosphate + CDP-diaclglyerol, a critical step in the synthesis of all major yeast phospholipids | Putative protein of unknown function; YBR220C is not an essential gene | Protein involved in bud-site selection and required for axial budding pattern; localizes with septins to bud neck in mitosis and may constitute an axial landmark for next round of budding | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; acts in a complex with Pmt2p, can instead interact with Pmt3p in some conditions; target for new antifungals | Protein of unknown function, identified as a high copy suppressor of psk1 psk2 mutations that confer temperature-sensitivity for galactose utilization; proposed to bind single-stranded nucleic acids via its R3H domain | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to vacuolar membrane; YDR119W is not an essential gene | One of two homeobox transcriptional repressors (see also Yox1p), that bind to Mcm1p and to early cell cycle box (ECB) elements of cell cycle regulated genes, thereby restricting ECB-mediated transcription to the M/G1 interval | Putative protein of unknown function | Pantothenate kinase (ATP:D-pantothenate 4'-phosphotransferase, EC 2.7.1.33); catalyzes the first committed step in the universal biosynthetic pathway leading to coenzyme A1 | Putative protein of unknown function; identified as interacting with Hsp82p in a high-throughput two-hybrid screen; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Protein phosphatase required for mitotic exit; located in the nucleolus until liberated by the FEAR and Mitotic Exit Network in anaphase, enabling it to act on key substrates to effect a decrease in CDK/B-cyclin activity and mitotic exit | Essential subunit of the Pan2p-Pan3p poly(A)-ribonuclease complex, which acts to control poly(A) tail length and regulate the stoichiometry and activity of postreplication repair complexes | Positive regulator of the Gcn2p kinase activity, forms a complex with Gcn20p; proposed to stimulate Gcn2p activation by an uncharged tRNA | Cargo-transport protein involved in endocytosis; may interact with ribosomes, based on co-purification experiments; GFP-fusion protein localizes to the cytoplasm; YGR198W is an essential gene | Protein involved in the regulation of cell wall synthesis; proposed to be involved in coordinating cell cycle progression with cell wall integrity | Cyclin-dependent kinase (CDK) inhibitor, regulates Pho80p-Pho85p and Pcl7p-Pho85p cyclin-CDK complexes in response to phosphate levels; inhibitory activity for Pho80p-Pho85p requires myo-D-inositol heptakisphosphate (IP7) generated by Vip1p | DNA helicase involved in rDNA replication and Ty1 transposition; relieves replication fork pauses at telomeric regions; structurally and functionally related to Pif1p | Member of the Sec24p family; forms a complex, with Sec23p, that is involved in sorting of Pma1p into COPII vesicles; peripheral ER membrane protein; potential Cdc28p substrate | Subunit of the origin recognition complex, which directs DNA replication by binding to replication origins and is also involved in transcriptional silencing; phosphorylated by Cdc28p | Histone methyltransferase, subunit of the COMPASS (Set1C) complex which methylates histone H3 on lysine 4; required in transcriptional silencing near telomeres and at the silent mating type loci; contains a SET domain | Protein involved in regulation of sterol biosynthesis; specifically stabilizes Hmg2p, one of two HMG-CoA isoenzymes that catalyze the rate-limiting step in sterol biosynthesis; homolog of mammalian INSIG proteins | Subunit of TORC1, a rapamycin-sensitive complex involved in growth control that contains Tor1p or Tor2p, Lst8p and Tco89p; contains four HEAT repeats and seven WD-40 repeats; may act as a scaffold protein to couple TOR and its effectors | Alpha mannosidase-like protein of the endoplasmic reticulum required for degradation of glycoproteins but not for processing of N-linked oligosaccharides | Long chain fatty acyl-CoA synthetase, has a preference for C16 and C18 fatty acids; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | DNA repair and TFIIH regulator, required for both nucleotide excision repair (NER) and RNA polymerase II (RNAP II) transcription; involved in telomere maintenance | Protein involved in cell morphogenesis and proliferation, associated with protein kinase Cbk1p; mutants activate OCH1 transcription | Integral plasma membrane protein required for axial budding in haploid cells, localizes to the incipient bud site and bud neck; glycosylated by Pmt4p; potential Cdc28p substrate | Core subunit of the RENT complex, which is a complex involved in nucleolar silencing and telophase exit; stimulates transcription by RNA polymerase I and regulates nucleolar structure | Mitochondrial aspartate aminotransferase, catalyzes the conversion of oxaloacetate to aspartate in aspartate and asparagine biosynthesis | Protein involved in nutritional control of the cell cycle; regulates abundance of the translation initiation factor eIF2; ortholog of human D123 protein | Phospholipase B (lysophospholipase) involved in phospholipid metabolism; displays transacylase activity in vitro; overproduction confers resistance to lysophosphatidylcholine | Nuclear pore membrane glycoprotein; may be involved in duplication of nuclear pores and nuclear pore complexes during S-phase | Vacuolar cation channel, mediates release of Ca(2+) from the vacuole in response to hyperosmotic shock | Protein involved in positive regulation of both 1,3-beta-glucan synthesis and the Pkc1p-MAPK pathway, potential Cdc28p substrate; multicopy suppressor of temperature-sensitive mutations in CDC24 and CDC42, and of mutations in BEM4 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and the nucleus | Putative protein kinase; similar to Kin4p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YPL141C is not an essential gene | Chaperone component involved in 20S proteasome assembly; forms a heterodimer with Irc25p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; null mutant is viable, exhibits shortened telomeres | Protein of unknown function; null mutant displays increased frequency of mitochondrial genome loss (petite formation) and reduced growth rate in minimal glycerol media | Essential protein required for determination of budding pattern, promotes localization of axial markers Bud4p and Cdc12p and functionally interacts with Sec3p, localizes to the contractile ring during anaphase, member of the IQGAP family | Spermidine synthase, involved in biosynthesis of spermidine and also in biosynthesis of pantothenic acid; spermidine is required for growth of wild-type cells |
| 11 | 45 | 436 | 0.488 | 0.571 |  |  | 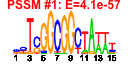 | 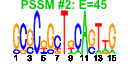 |  | YBL086C YBR184W YBR284W YDL089W YDL154W YDL239C YDR285W YDR290W YDR344C YDR374C YER153C YER179W YFL015C YFL051C YFR023W YFR026C YGL007W YGL033W YGL182C YGL183C YGR051C YGR059W YGR269W YHR079BC YHR093W YHR124W YHR157W YIL072W YIL073C YIR013C YKR104W YLL030C YLL046C YLR329W YLR394W YMR101C YMR232W YNL196C YOL104C YOL131W YOR084W YOR186W YOR351C YPL082C YPL201C | MSH5 ADY3 ZIP1 PET122 DMC1 PES4 HOP2 MND1 SPR3 NDT80 REC104 HOP1 SPO22 GAT4 NFT1 RNP1 REC102 CST9 SRT1 FUS2 SLZ1 TMT1 LPX1 MEK1 MOT1 YIG1 | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | Putative protein of unknown function; YBR184W is not an essential gene | Putative protein of unknown function; YBR284W is not an essential gene; null mutant exhibits decreased resistance to rapamycin and wortmannin | Protein of unknown function; interacts with meiotic division protein Csm1p; green fluorescent protein (GFP)-fusion protein localizes to the nuclear periphery, potential Cdc28p substrate | Protein of the MutS family, forms a dimer with Msh4p that facilitates crossovers between homologs during meiosis; msh5-Y823H mutation confers tolerance to DNA alkylating agents; homologs present in C. elegans and humans | Protein required for spore wall formation, thought to mediate assembly of a Don1p-containing structure at the leading edge of the prospore membrane via interaction with spindle pole body components; potentially phosphorylated by Cdc28p | Transverse filament protein of the synaptonemal complex; required for normal levels of meiotic recombination and pairing between homologous chromosome during meiosis; potential Cdc28p substrate | Putative protein of unknown function | Mitochondrial translational activator specific for the COX3 mRNA, acts together with Pet54p and Pet494p; located in the mitochondrial inner membrane | Meiosis-specific protein required for repair of double-strand breaks and pairing between homologous chromosomes; homolog of Rad51p and the bacterial RecA protein | Putative protein of unknown function; YFL051C is not an essential gene | Poly(A) binding protein, suppressor of DNA polymerase epsilon mutation, similar to Mip6p | Meiosis-specific protein that localizes to chromosomes, preventing synapsis between nonhomologous chromosomes and ensuring synapsis between homologs; complexes with Mnd1p to promote homolog pairing and meiotic double-strand break repair | Protein required for recombination and meiotic nuclear division; forms a complex with Hop2p, which is involved in chromosome pairing and repair of meiotic double-strand breaks | Sporulation-specific homolog of the yeast CDC3/10/11/12 family of bud neck microfilament genes; septin protein involved in sporulation; regulated by ABFI | Meiosis-specific transcription factor required for exit from pachytene and for full meiotic recombination; activates middle sporulation genes; competes with Sum1p for binding to promoters containing middle sporulation elements (MSE) | Protein involved in early stages of meiotic recombination; required for meiotic crossing over; forms a complex with Rec102p and Spo11p necessary during the initiation of recombination | Meiosis-specific DNA binding protein that displays Red1p dependent localization to the unsynapsed axial-lateral elements of the synaptonemal complex; required for homologous chromosome synapsis and chiasma formation | Meiosis-specific protein essential for chromosome synapsis, similar to phospholipase A2, involved in completion of nuclear divisions during meiosis; induced early in meiosis | Protein containing GATA family zinc finger motifs | Putative transporter of the multidrug resistance-associated protein (MRP) subfamily; adjacent ORFs YKR103W and YKR104W are merged in different strain backgrounds. | Ribonucleoprotein that contains two RNA recognition motifs (RRM) | Protein involved in early stages of meiotic recombination; required for chromosome synapsis; forms a complex with Rec104p and Spo11p necessary during the initiation of recombination | SUMO E3 ligase; required for synaptonemal complex formation; localizes to synapsis initiation sites on meiotic chromosomes; potential Cdc28p substrate | Cis-prenyltransferase involved in synthesis of long-chain dolichols (19-22 isoprene units; as opposed to Rer2p which synthesizes shorter-chain dolichols); localizes to lipid bodies; transcription is induced during stationary phase | Cytoplasmic protein localized to the shmoo tip; required for the alignment of parental nuclei before nuclear fusion during mating | Sporulation-specific protein with a leucine zipper motif | Trans-aconitate methyltransferase, cytosolic enzyme that catalyzes the methyl esterification of 3-isopropylmalate, an intermediate of the leucine biosynthetic pathway, and trans-aconitate, which inhibits the citric acid cycle | Oleic acid-inducible, peroxisomal matrix localized lipase; transcriptionally activated by Yrm1p along with genes involved in multidrug resistance; peroxisomal import is dependent on the PTS1 receptor, Pex5p and on self-interaction | Putative protein of unknown function; proper regulation of expression during heat stress is sphingolipid-dependent | Meiosis-specific serine/threonine protein kinase, functions in meiotic checkpoint, promotes recombination between homologous chromosomes by suppressing double strand break repair between sister chromatids | Essential abundant protein involved in regulation of transcription, removes Spt15p (TBP) from DNA via its C-terminal ATPase activity, forms a complex with TBP that binds TATA DNA with high affinity but with altered specificity | Protein that interacts with glycerol 3-phosphatase and plays a role in anaerobic glycerol production; localizes to the nucleus and cytosol |
| 240 | 41 | 436 | 0.489 | 0.591 |  |  | 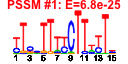 | 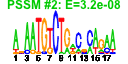 |  | YDR432W YIL114C YOR344C YAL003W YAL007C YBR011C YBR196C YCR012W YCR013C YDL192W YDR002W YDR050C YDR226W YEL027W YEL034W YEL035C YER026C YFL037W YGL189C YGR192C YGR254W YHR046C YHR123W YHR174W YIL053W YJR009C YJR047C YKL152C YKL153W YLL014W YLL024C YLR065C YLR179C YLR301W YLR354C YML094W YNL244C YOL014W YOL086C YOR375C YPR009W | NPL3 POR2 TYE7 EFB1 ERP2 IPP1 PGI1 PGK1 ARF1 YRB1 TPI1 ADK1 CUP5 HYP2 UTR5 CHO1 TUB2 RPS26A TDH3 ENO1 INM1 EPT1 ENO2 RHR2 TDH2 ANB1 GPM1 SSA2 TAL1 GIM5 SUI1 ADH1 GDH1 SUT2 | RNA-binding protein that carries poly(A)+ mRNA from the nucleus into the cytoplasm; dissociation from mRNAs is promoted by Mtr10p; phosphorylated by Sky1p in the cytoplasm | Putative mitochondrial porin (voltage-dependent anion channel), related to Por1p but not required for mitochondrial membrane permeability or mitochondrial osmotic stability | Serine-rich protein that contains a basic-helix-loop-helix (bHLH) DNA binding motif; binds E-boxes of glycolytic genes and contributes to their activation; may function as a transcriptional activator in Ty1-mediated gene expression | Translation elongation factor 1 beta; stimulates nucleotide exchange to regenerate EF-1 alpha-GTP for the next elongation cycle; part of the EF-1 complex, which facilitates binding of aminoacyl-tRNA to the ribosomal A site | Protein that forms a heterotrimeric complex with Erp1p, Emp24p, and Erv25p; member, along with Emp24p and Erv25p, of the p24 family involved in ER to Golgi transport and localized to COPII-coated vesicles | Cytoplasmic inorganic pyrophosphatase (PPase), catalyzes the rapid exchange of oxygens from Pi with water, highly expressed and essential for viability, active-site residues show identity to those from E. coli PPase | Glycolytic enzyme phosphoglucose isomerase, catalyzes the interconversion of glucose-6-phosphate and fructose-6-phosphate; required for cell cycle progression and completion of the gluconeogenic events of sporulation | 3-phosphoglycerate kinase, catalyzes transfer of high-energy phosphoryl groups from the acyl phosphate of 1,3-bisphosphoglycerate to ADP to produce ATP; key enzyme in glycolysis and gluconeogenesis | ADP-ribosylation factor, GTPase of the Ras superfamily involved in regulation of coated vesicle formation in intracellular trafficking within the Golgi; functionally interchangeable with Arf2p | Ran GTPase binding protein; involved in nuclear protein import and RNA export, ubiquitin-mediated protein degradation during the cell cycle; shuttles between the nucleus and cytoplasm; is essential; homolog of human RanBP1 | Triose phosphate isomerase, abundant glycolytic enzyme; mRNA half-life is regulated by iron availability; transcription is controlled by activators Reb1p, Gcr1p, and Rap1p through binding sites in the 5' non-coding region | Adenylate kinase, required for purine metabolism; localized to the cytoplasm and the mitochondria; lacks cleavable signal sequence | Proteolipid subunit of the vacuolar H(+)-ATPase V0 sector (subunit c; dicyclohexylcarbodiimide binding subunit); required for vacuolar acidification and important for copper and iron metal ion homeostasis | Translation initiation factor eIF-5A, promotes formation of the first peptide bond; similar to and functionally redundant with Anb1p; possible role in translation elongation; undergoes an essential hypusination modification | Protein of unknown function; transcription may be regulated by Gcr1p | Phosphatidylserine synthase, functions in phospholipid biosynthesis; catalyzes the reaction CDP-diaclyglycerol + L-serine = CMP + L-1-phosphatidylserine, transcriptionally repressed by myo-inositol and choline | Beta-tubulin; associates with alpha-tubulin (Tub1p and Tub3p) to form tubulin dimer, which polymerizes to form microtubules | Protein component of the small (40S) ribosomal subunit; nearly identical to Rps26Bp and has similarity to rat S26 ribosomal protein | Glyceraldehyde-3-phosphate dehydrogenase, isozyme 3, involved in glycolysis and gluconeogenesis; tetramer that catalyzes the reaction of glyceraldehyde-3-phosphate to 1,3 bis-phosphoglycerate; detected in the cytoplasm and cell-wall | Enolase I, a phosphopyruvate hydratase that catalyzes the conversion of 2-phosphoglycerate to phosphoenolpyruvate during glycolysis and the reverse reaction during gluconeogenesis; expression is repressed in response to glucose | Inositol monophosphatase, involved in biosynthesis of inositol and in phosphoinositide second messenger signaling; INM1 expression increases in the presence of inositol and decreases upon exposure to antibipolar drugs lithium and valproate | sn-1,2-diacylglycerol ethanolamine- and cholinephosphotranferase; not essential for viability | Enolase II, a phosphopyruvate hydratase that catalyzes the conversion of 2-phosphoglycerate to phosphoenolpyruvate during glycolysis and the reverse reaction during gluconeogenesis; expression is induced in response to glucose | Constitutively expressed isoform of DL-glycerol-3-phosphatase; involved in glycerol biosynthesis, induced in response to both anaerobic and, along with the Hor2p/Gpp2p isoform, osmotic stress | Glyceraldehyde-3-phosphate dehydrogenase, isozyme 2, involved in glycolysis and gluconeogenesis; tetramer that catalyzes the reaction of glyceraldehyde-3-phosphate to 1,3 bis-phosphoglycerate; detected in the cytoplasm and cell-wall | Translation initiation factor eIF-5A, promotes formation of the first peptide bond; similar to and functionally redundant with Hyp2p; undergoes an essential hypusination modification; expressed under anaerobic conditions | Tetrameric phosphoglycerate mutase, mediates the conversion of 3-phosphoglycerate to 2-phosphoglycerate during glycolysis and the reverse reaction during gluconeogenesis | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum; YLL014W is not an essential gene | ATP binding protein involved in protein folding and vacuolar import of proteins; member of heat shock protein 70 (HSP70) family; associated with the chaperonin-containing T-complex; present in the cytoplasm, vacuolar membrane and cell wall | Putative protein of unknown function; YLR065C is not an essential gene | Protein of unknown function, transcription is activated by paralogous proteins Yrm1p and Yrr1p along with proteins involved in multidrug resistance; GFP-tagged protein localizes to the cytoplasm and nucleus; YLR179C is not essential | Protein of unknown function that interacts with Sec72p | Transaldolase, enzyme in the non-oxidative pentose phosphate pathway; converts sedoheptulose 7-phosphate and glyceraldehyde 3-phosphate to erythrose 4-phosphate and fructose 6-phosphate | Subunit of the heterohexameric cochaperone prefoldin complex which binds specifically to cytosolic chaperonin and transfers target proteins to it | Translation initiation factor eIF1; component of a complex involved in recognition of the initiator codon; modulates translation accuracy at the initiation phase | Putative protein of unknown function | Alcohol dehydrogenase, fermentative isozyme active as homo- or heterotetramers; required for the reduction of acetaldehyde to ethanol, the last step in the glycolytic pathway | NADP(+)-dependent glutamate dehydrogenase, synthesizes glutamate from ammonia and alpha-ketoglutarate; rate of alpha-ketoglutarate utilization differs from Gdh3p; expression regulated by nitrogen and carbon sources | Putative transcription factor; multicopy suppressor of mutations that cause low activity of the cAMP/protein kinase A pathway; highly similar to Sut1p |
| 169 | 49 | 438 | 0.490 | 0.590 |  |  | 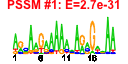 | 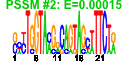 |  | YBL056W YDL033C YDR462W YGL040C YGR021W YGR207C YHL031C YHR059W YHR116W YIL070C YJR122W YKL087C YKL138C YKL195W YKR084C YLL013C YMR267W YNL185C YNR033W YOL033W YOR088W YOR354C YPL031C YPL097W YPL104W YPL105C YPL132W YPL173W YPR047W YAL019W YBR003W YBR037C YBR146W YDR041W YDR116C YDR494W YJL096W YKR006C YLR439W YMR024W YNL252C YNL315C YNR036C YOR068C YOR087W YOR158W YPL172C YPL215W YPR166C | PTC3 SLM3 MRPL28 HEM2 GOS1 FYV4 COX23 MAM33 IBA57 CYT2 HSK3 MIA40 HBS1 PUF3 PPA2 MRPL19 ABZ1 MSE1 YVC1 MSC6 PHO85 MSY1 MSD1 COX11 MRPL40 AIM30 FUN30 COQ1 SCO1 MRPS9 RSM10 MRPL1 RSM28 MRPL49 MRPL13 MRPL4 MRPL3 MRPL17 ATP11 VAM10 PET123 COX10 CBP3 MRP2 | Type 2C protein phosphatase; dephosphorylates Hog1p (see also Ptc2p) to limit maximal kinase activity induced by osmotic stress; dephosphorylates T169 phosphorylated Cdc28p (see also Ptc2p); role in DNA checkpoint inactivation | tRNA-specific 2-thiouridylase, responsible for 2-thiolation of the wobble base of mitochondrial tRNAs; human ortholog is implicated in myoclonus epilepsy associated with ragged red fibers (MERRF) | Mitochondrial ribosomal protein of the large subunit | Delta-aminolevulinate dehydratase, a homo-octameric enzyme, catalyzes the conversion of delta-aminolevulinic acid to porphobilinogen, the second step in the heme biosynthetic pathway; localizes to both the cytoplasm and nucleus | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein that interacts with frataxin (Yfh1p); putative homolog of mammalian electron transfer flavoprotein complex subunit ETF-beta; authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | v-SNARE protein involved in Golgi transport, homolog of the mammalian protein GOS-28/GS28 | Protein of unknown function, required for survival upon exposure to K1 killer toxin | Mitochondrial intermembrane space protein that functions in mitochondrial copper homeostasis, essential for functional cytochrome oxidase expression; homologous to Cox17p | Acidic protein of the mitochondrial matrix involved in oxidative phosphorylation; related to the human complement receptor gC1q-R | Mitochondrial matrix protein involved in the incorporation of iron-sulfur clusters into mitochondrial aconitase-type proteins; activates the radical-SAM family members Bio2p and Lip5p; interacts with Ccr4p in the two-hybrid system | Cytochrome c1 heme lyase, involved in maturation of cytochrome c1, which is a subunit of the mitochondrial ubiquinol-cytochrome-c reductase; links heme covalently to apocytochrome c1 | Essential subunit of the Dam1 complex (aka DASH complex), couples kinetochores to the force produced by MT depolymerization thereby aiding in chromosome segregation; is transferred to the kinetochore prior to mitosis | Essential protein of the mitochondrial intermembrane space (IMS); promotes retention of newly imported proteins; may do so by stabilizing client protein folding as part of a disulfide relay system or transferring metal to client proteins | GTP binding protein with sequence similarity to the elongation factor class of G proteins, EF-1alpha and Sup35p; associates with Dom34p, and shares a similar genetic relationship with genes that encode ribosomal protein components | Protein of the mitochondrial outer surface, links the Arp2/3 complex with the mitochore during anterograde mitochondrial movement; also binds to and promotes degradation of mRNAs for select nuclear-encoded mitochondrial proteins | Mitochondrial inorganic pyrophosphatase, required for mitochondrial function and possibly involved in energy generation from inorganic pyrophosphate | Para-aminobenzoate (PABA) synthase, has similarity to Escherichia coli PABA synthase components PabA and PabB | Mitochondrial glutamyl-tRNA synthetase, encoded by a nuclear gene | Vacuolar cation channel, mediates release of Ca(2+) from the vacuole in response to hyperosmotic shock | Protein of unknown function; mutant is defective in directing meiotic recombination events to homologous chromatids; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Cyclin-dependent kinase, with ten cyclin partners; involved in environmental stress response; in phosphate-rich conditions, Pho85p-Pho80p complex phosphorylates Pho4p which in turn represses PHO5 | Mitochondrial tyrosyl-tRNA synthetase | Mitochondrial aspartyl-tRNA synthetase, required for acylation of aspartyl-tRNA; yeast and bacterial aspartyl-, asparaginyl-, and lysyl-tRNA synthetases contain regions with high sequence similarity, suggesting a common ancestral gene | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Mitochondrial inner membrane protein required for delivery of copper to the Cox1p subunit of cytochrome c oxidase; association with mitochondrial ribosomes suggests that copper delivery may occur during translation of Cox1p | Putative protein of unknown function that may be involved in intramitochondrial sorting; similar to Ups1p and to human PRELI; GFP-tagged protein localizes to mitochondria; required for wild-type respiratory growth | Protein whose overexpression affects chromosome stability, potential Cdc28p substrate; homolog of Snf2p; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Hexaprenyl pyrophosphate synthetase, catalyzes the first step in ubiquinone (coenzyme Q) biosynthesis | Copper-binding protein of the mitochondrial inner membrane, required for cytochrome c oxidase activity and respiration; may function to deliver copper to cytochrome c oxidase; has similarity to thioredoxins | Mitochondrial ribosomal protein of the small subunit | Mitochondrial ribosomal protein of the small subunit, has similarity to E. coli S10 ribosomal protein; essential for viability, unlike most other mitoribosomal proteins | Mitochondrial ribosomal protein of the small subunit; genetic interactions suggest a possible role in promoting translation initiation | Mitochondrial ribosomal protein of the large subunit, not essential for mitochondrial translation | Molecular chaperone, required for the assembly of alpha and beta subunits into the F1 sector of mitochondrial F1F0 ATP synthase | Mitochondrial protein; may interact with ribosomes based on co-purification experiments; similar to E. coli and human mitochondrial S12 ribosomal proteins | Protein involved in vacuole morphogenesis; acts at an early step of homotypic vacuole fusion that is required for vacuole tethering | Mitochondrial ribosomal protein of the small subunit; PET123 exhibits genetic interactions with PET122, which encodes a COX3 mRNA-specific translational activator | Heme A:farnesyltransferase, catalyzes the first step in the conversion of protoheme to the heme A prosthetic group required for cytochrome c oxidase activity; human ortholog is associated with mitochondrial disorders | Mitochondrial protein required for assembly of ubiquinol cytochrome-c reductase complex (cytochrome bc1 complex); interacts with Cbp4p and function is partially redundant with that of Cbp4p |
| 292 | 45 | 440 | 0.497 | 0.542 |  |  | 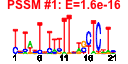 | 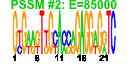 |  | YBL026W YBL062W YBR242W YCRX04W YDL157C YDR381W YDR429C YER146W YER174C YGL246C YGL255W YGR064W YGR093W YGR114C YGR276C YHR127W YIL004C YJL026W YJL169W YJL179W YJR014W YJR068W YJR133W YJR135C YJR141W YKL048C YKL122C YLR101C YLR145W YLR154C YML105C YMR132C YMR321C YNL032W YNL150W YNL188W YNR025C YOL079W YOR201C YOR251C YOR263C YOR265W YOR283W YPL199C YPR062W | LSM2 YRA1 TIF35 LSM5 GRX4 RAI1 ZRT1 DRN1 RNH70 BET1 RNR2 PFD1 TMA22 RFC2 XPT1 MCM22 ELM1 SRP21 RMP1 RNH203 SEC65 JLP2 SIW14 KAR1 MRM1 RBL2 FCY1 | Lsm (Like Sm) protein; part of heteroheptameric complexes (Lsm2p-7p and either Lsm1p or 8p): cytoplasmic Lsm1p complex involved in mRNA decay; nuclear Lsm8p complex part of U6 snRNP and possibly involved in processing tRNA, snoRNA, and rRNA | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YBR242W is not an essential gene | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Nuclear protein that binds to RNA and to Mex67p, required for export of poly(A)+ mRNA from the nucleus; member of the REF (RNA and export factor binding proteins) family; another family member, Yra2p, can substitute for Yra1p function | Subunit of the core complex of translation initiation factor 3(eIF3), which is essential for translation | Hydroperoxide and superoxide-radical responsive glutathione-dependent oxidoreductase; monothiol glutaredoxin subfamily member along with Grx3p and Grx5p; protects cells from oxidative damage | Nuclear protein that binds to and stabilizes the exoribonuclease Rat1p, required for pre-rRNA processing | High-affinity zinc transporter of the plasma membrane, responsible for the majority of zinc uptake; transcription is induced under low-zinc conditions by the Zap1p transcription factor | Putative debranching enzyme associated ribonuclease; green fluorescent protein (GFP)-fusion protein localizes to the nucleus | 3' exoribonuclease, required for 5S rRNA and tRNA-Arg3 maturation | Protein of unknown function; localizes to the nucleus | Type II membrane protein required for vesicular transport between the endoplasmic reticulum and Golgi complex; v-SNARE with similarity to synaptobrevins | Ribonucleotide-diphosphate reductase (RNR), small subunit; the RNR complex catalyzes the rate-limiting step in dNTP synthesis and is regulated by DNA replication and DNA damage checkpoint pathways via localization of the small subunits | Subunit of heterohexameric prefoldin, which binds cytosolic chaperonin and transfers target proteins to it; involved in the biogenesis of actin and of alpha- and gamma-tubulin | Protein of unknown function; associates with ribosomes and has a putative RNA binding domain; interacts with Tma20p; similar to human GRAP and human DRP1, which interacts with human Tma20p homolog MCT-1 | Subunit of heteropentameric Replication factor C (RF-C), which is a DNA binding protein and ATPase that acts as a clamp loader of the proliferating cell nuclear antigen (PCNA) processivity factor for DNA polymerases delta and epsilon | Xanthine-guanine phosphoribosyl transferase, required for xanthine utilization and for optimal utilization of guanine | Protein involved in minichromosome maintenance; component of the kinetochore; binds to centromeric DNA in a Ctf19p-dependent manner | Putative protein of unknown function; may be involved in mRNA processing; YJR141W is an essential gene | Serine/threonine protein kinase that regulates cellular morphogenesis, septin behavior, and cytokinesis; required for the regulation of other kinases; forms part of the bud neck ring | Subunit of the signal recognition particle (SRP), which functions in protein targeting to the endoplasmic reticulum membrane; not found in mammalian SRP; forms a pre-SRP structure in the nucleolus that is translocated to the cytoplasm | Subunit of RNase MRP, which processes pre-rRNA and has a role in cell cycle-regulated degradation of daughter cell-specific mRNAs; unlike most subunits, not shared between RNase MRP and nuclear RNase P | Ribonuclease H2 subunit, required for RNase H2 activity | Subunit of the signal recognition particle (SRP), involved in protein targeting to the ER; interacts with Srp54p; homolog of mammalian SRP19 | Protein of unknown function, contains sequence that closely resembles a J domain (typified by the E. coli DnaJ protein) | Putative protein of unknown function | Tyrosine phosphatase that plays a role in actin filament organization and endocytosis; localized to the cytoplasm | Essential protein involved in karyogamy during mating and in spindle pole body duplication during mitosis, localizes to the half-bridge of the spindle pole body, interacts with Spc72p during karyogamy, also interacts with Cdc31p | Ribose methyltransferase that modifies a functionally critical, conserved nucleotide in mitochondrial 21S rRNA | Putative protein of unknown function with similarity to human thiosulfate sulfurtransferase, an enzyme that catalyzes the transfer of sulfur from thiosulfate to cyanide, forming sulfite and thiocyanate; YOR251C is a non-essential gene | Protein involved in microtubule morphogenesis, required for protection from excess free beta-tubulin; proposed to be involved the folding of beta-tubulin; similar to mouse beta-tubulin cofactor A | Putative protein of unknown function with some similarity to GPM1/YKL152C, a phosphoglycerate mutase; YOR283W is not an essential gene | hypothetical protein | Cytosine deaminase, zinc metalloenzyme that catalyzes the hydrolytic deamination of cytosine to uracil; of biomedical interest because it also catalyzes the deamination of 5-fluorocytosine (5FC) to form anticancer drug 5-fluorouracil (5FU) |
| 299 | 42 | 448 | 0.503 | 0.580 | 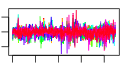 |  | 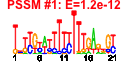 | 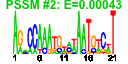 |  | YAL023C YBL004W YBR190W YCL046W YDL096C YDL112W YDL122W YDL195W YDR024W YDR234W YDR497C YDR508C YGL195W YGR090W YHR023W YHR047C YHR072W YHR143BW YHR206W YIL092W YIL093C YIR033W YJL080C YJR031C YKL182W YKR082W YLL048C YLR305C YLR384C YML075C YMR006C YMR010W YMR015C YMR129W YMR247C YNL024C YNR016C YNR075W YOR098C YOR370C YPL128C YPL231W | PMT2 UTP20 TRM3 UBP1 SEC31 LYS4 ITR1 GNP1 GCN1 UTP22 MYO1 ATP8 ERG7 SKN7 RSM25 MGA2 SCP160 GEA1 FAS1 NUP133 BAT1 STT4 IKI3 HMG1 PLB2 ERG5 POM152 RKR1 YNL024C-A ACC1 COS10 NUP1 MRS6 TBF1 FAS2 | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; acts in a complex with Pmt1p, can instead interact with Pmt5p in some conditions; target for new antifungals | Component of the small-subunit (SSU) processome, which is involved in the biogenesis of the 18S rRNA | 2'-O-ribose methyltransferase, catalyzes the ribose methylation of the guanosine nucleotide at position 18 of tRNAs | Ubiquitin-specific protease that removes ubiquitin from ubiquitinated proteins; cleaves at the C terminus of ubiquitin fusions irrespective of their size; capable of cleaving polyubiquitin chains | Essential phosphoprotein component (p150) of the COPII coat of secretory pathway vesicles, in complex with Sec13p; required for ER-derived transport vesicle formation | Homoaconitase, catalyzes the conversion of homocitrate to homoisocitrate, which is a step in the lysine biosynthesis pathway | Myo-inositol transporter with strong similarity to the minor myo-inositol transporter Itr2p, member of the sugar transporter superfamily; expression is repressed by inositol and choline via Opi1p and derepressed via Ino2p and Ino4p | High-affinity glutamine permease, also transports Leu, Ser, Thr, Cys, Met and Asn; expression is fully dependent on Grr1p and modulated by the Ssy1p-Ptr3p-Ssy5p (SPS) sensor of extracellular amino acids | Positive regulator of the Gcn2p kinase activity, forms a complex with Gcn20p; proposed to stimulate Gcn2p activation by an uncharged tRNA | Possible U3 snoRNP protein involved in maturation of pre-18S rRNA, based on computational analysis of large-scale protein-protein interaction data | Type II myosin heavy chain, required for wild-type cytokinesis and cell separation; localizes to the actomyosin ring; binds to myosin light chains Mlc1p and Mlc2p through its IQ1 and IQ2 motifs respectively | Subunit 8 of the F0 sector of mitochondrial inner membrane F1-F0 ATP synthase, encoded on the mitochondrial genome | Lanosterol synthase, an essential enzyme that catalyzes the cyclization of squalene 2,3-epoxide, a step in ergosterol biosynthesis | Nuclear response regulator and transcription factor, part of a branched two-component signaling system; required for optimal induction of heat-shock genes in response to oxidative stress; involved in osmoregulation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and to the nucleus | Mitochondrial ribosomal protein of the small subunit | ER membrane protein involved in regulation of OLE1 transcription, acts with homolog Spt23p; inactive ER form dimerizes and one subunit is then activated by ubiquitin/proteasome-dependent processing followed by nuclear targeting | Essential RNA-binding G protein effector of mating response pathway, mainly associated with nuclear envelope and ER, interacts in mRNA-dependent manner with translating ribosomes via multiple KH domains, similar to vertebrate vigilins | Guanine nucleotide exchange factor for ADP ribosylation factors (ARFs), involved in vesicular transport between the Golgi and ER, Golgi organization, and actin cytoskeleton organization; similar to but not functionally redundant with Gea2p | Beta subunit of fatty acid synthetase, which catalyzes the synthesis of long-chain saturated fatty acids; contains acetyltransacylase, dehydratase, enoyl reductase, malonyl transacylase, and palmitoyl transacylase activities | Subunit of the Nup84p subcomplex of the nuclear pore complex (NPC), localizes to both sides of the NPC, required to establish a normal nucleocytoplasmic concentration gradient of the GTPase Gsp1p | Mitochondrial branched-chain amino acid aminotransferase, homolog of murine ECA39; highly expressed during logarithmic phase and repressed during stationary phase | Phosphatidylinositol-4-kinase that functions in the Pkc1p protein kinase pathway; required for normal vacuole morphology, cell wall integrity, and actin cytoskeleton organization | Subunit of Elongator complex, which is required for modification of wobble nucleosides in tRNA; maintains structural integrity of Elongator; homolog of human IKAP, mutations in which cause familial dysautonomia (FD) | One of two isozymes of HMG-CoA reductase that catalyzes the conversion of HMG-CoA to mevalonate, which is a rate-limiting step in sterol biosynthesis; localizes to the nuclear envelope; overproduction induces the formation of karmellae | Phospholipase B (lysophospholipase) involved in phospholipid metabolism; displays transacylase activity in vitro; overproduction confers resistance to lysophosphatidylcholine | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YMR010W is not an essential gene; YMR010W mRNA is transcribed with ADI1 | C-22 sterol desaturase, a cytochrome P450 enzyme that catalyzes the formation of the C-22(23) double bond in the sterol side chain in ergosterol biosynthesis; may be a target of azole antifungal drugs | Nuclear pore membrane glycoprotein; may be involved in duplication of nuclear pores and nuclear pore complexes during S-phase | Nuclear RING domain protein with functional connections to chromatin modification; may interact with ribosomes, based on co-purification experiments; YMR247C is not an essential gene | Putative protein of unknown function; YNL024C-A is an essential gene | Acetyl-CoA carboxylase, biotin containing enzyme that catalyzes the carboxylation of acetyl-CoA to form malonyl-CoA; required for de novo biosynthesis of long-chain fatty acids | Protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Nuclear pore complex (NPC) subunit, involved in protein import/export and in export of RNAs, possible karyopherin release factor that accelerates release of karyopherin-cargo complexes after transport across NPC; potential Cdc28p substrate | Rab escort protein, forms a complex with the Ras-like small GTPase Ypt1p that is required for the prenylation of Ypt1p by protein geranylgeranyltransferase type II (Bet2p-Bet4p) | Telobox-containing general regulatory factor; binds to TTAGGG repeats within subtelomeric anti-silencing regions (STARs) and possibly throughout the genome and mediates their insulating capacity by blocking silent chromatin propagation | Alpha subunit of fatty acid synthetase, which catalyzes the synthesis of long-chain saturated fatty acids; contains beta-ketoacyl reductase and beta-ketoacyl synthase activities; phosphorylated |
| 103 | 36 | 442 | 0.518 | 0.597 | 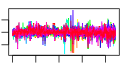 |  | 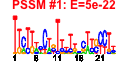 | 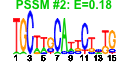 |  | YBR145W YCR023C YDR009W YDR082W YDR386W YDR532C YER051W YER152C YGL005C YGL251C YGR006W YGR224W YGR258C YHR022C YIL167W YIL168W YIR032C YJR152W YKL220C YNL097C YDL194W YDR282C YDR456W YDR481C YDR482C YEL005C YGL240W YHL044W YHR176W YIL122W YIL146C YIR015W YJL070C YKR075C YKR101W YOR244W | ADH5 GAL3 STN1 MUS81 JHD1 COG7 HFM1 PRP18 AZR1 RAD2 DAL3 DAL5 FRE2 YNL097C-B SNF3 NHX1 PHO8 CWC21 VAB2 DOC1 FMO1 POG1 ECM37 RPR2 SIR1 ESA1 | Alcohol dehydrogenase isoenzyme V; involved in ethanol production | Putative protein of unknown function; YCR023C is not an essential gene | Transcriptional regulator involved in activation of the GAL genes in response to galactose; forms a complex with Gal80p to relieve Gal80p inhibition of Gal4p; binds galactose and ATP but does not have galactokinase activity | Telomere end-binding and capping protein, plays a key role with Pol12p in linking telomerase action with completion of lagging strand synthesis, and in a regulatory step required for telomere capping | Helix-hairpin-helix protein, involved in DNA repair and replication fork stability; functions as an endonuclease in complex with Mms4p; interacts with Rad54p | Protein of unknown function that localizes to the nuclear side of the spindle pole body and along short spindles; deletion mutants have short telomeres; forms a complex with Spc105p | JmjC domain family histone demethylase specific for H3-K36, similar to proteins found in human, mouse, drosophila, X. laevis, C. elegans, and S. pombe | Putative protein of unknown function; shares amino acid similarity with the aminotransferases Aro8p and Aro9p; YER152C is not an essential gene | Component of the conserved oligomeric Golgi complex (Cog1p through Cog8p), a cytosolic tethering complex that functions in protein trafficking to mediate fusion of transport vesicles to Golgi compartments | Meiosis specific DNA helicase involved in the conversion of double-stranded breaks to later recombination intermediates and in crossover control; catalyzes the unwinding of Holliday junctions; has ssDNA and dsDNA stimulated ATPase activity | Splicing factor involved in the positioning of the 3' splice site during the second catalytic step of splicing, part of snRNP U5, interacts with Slu7p | Plasma membrane transporter of the major facilitator superfamily, involved in resistance to azole drugs such as ketoconazole and fluconazole | Single-stranded DNA endonuclease, cleaves single-stranded DNA during nucleotide excision repair to excise damaged DNA; subunit of Nucleotide Excision Repair Factor 3 (NEF3); homolog of human XPG protein | Putative protein of unknown function; YHR022C is not an essential gene. | Ureidoglycolate hydrolase, converts ureidoglycolate to glyoxylate and urea in the third step of allantoin degradation; expression sensitive to nitrogen catabolite repression | Allantoin permease; ureidosuccinate permease; expression is constitutive but sensitive to nitrogen catabolite repression | Ferric reductase and cupric reductase, reduces siderophore-bound iron and oxidized copper prior to uptake by transporters; expression induced by low iron levels but not by low copper levels | Putative protein of unknown function | Plasma membrane glucose sensor that regulates glucose transport; has 12 predicted transmembrane segments; long cytoplasmic C-terminal tail is required for low glucose induction of hexose transporter genes HXT2 and HXT4 | Endosomal Na+/H+ exchanger, required for intracellular sequestration of Na+; required for osmotolerance to acute hypertonic shock | Repressible alkaline phosphatase, a glycoprotein localized to the vacuole; regulated by levels of inorganic phosphate and by a system consisting of Pho4p, Pho9p, Pho80p, Pho81p and Pho85p; dephosphorylates phosphotyrosyl peptides | Component of a complex containing Cef1p, putatively involved in pre-mRNA splicing; may bind RNA; has similarity to S. pombe Cwf21p | Protein with a potential role in vacuolar function, as suggested by its ability to bind Vac8p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Processivity factor required for the ubiquitination activity of the anaphase promoting complex (APC), mediates the activity of the APC by contributing to substrate recognition; involved in cyclin proteolysis | Putative integral membrane protein, member of DUP240 gene family; green fluorescent protein (GFP)-fusion protein localizes to the plasma membrane in a punctate pattern | Flavin-containing monooxygenase, localized to the cytoplasmic face of the ER membrane; catalyzes oxidation of biological thiols to maintain the ER redox buffer ratio for correct folding of disulfide-bonded proteins | Putative transcriptional activator that promotes recovery from pheromone induced arrest; inhibits both alpha-factor induced G1 arrest and repression of CLN1 and CLN2 via SCB/MCB promoter elements; potential Cdc28p substrate; SBF regulated | Non-essential protein of unknown function | Subunit of nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends; not shared between RNase MRP and RNase P, in contrast to all other RNase P protein subunits | Putative protein of unknown function with similarity to AMP deaminases; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YJL070C is a non-essential gene | Protein of unknown function; similar to YOR062Cp and Reg1p; expression regulated by glucose and Rgt1p; GFP-fusion protein is induced in response to the DNA-damaging agent MMS | Protein involved in repression of transcription at the silent mating-type loci HML and HMR; recruitment to silent chromatin requires interactions with Orc1p and with Sir4p, through a common Sir1p domain; binds to centromeric chromatin | Histone acetyltransferase catalytic subunit of the native multisubunit complex (NuA4) that acetylates four conserved internal lysines of histone H4 N-terminal tail; required for cell cycle progression |
| 293 | 45 | 447 | 0.527 | 0.578 |  |  | 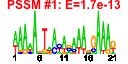 | 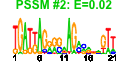 |  | YBL001C YBL006C YBL050W YBR062C YBR137W YCR075C YCR082W YCR096C YCR097W YDL002C YDL013W YDL059C YDL115C YDR005C YDR008C YDR079W YDR327W YDR452W YDR468C YDR486C YER044C YER048C YER058W YER059W YER080W YGL005C YGL091C YGR009C YGR028W YGR048W YGR222W YGR230W YHL020C YHR054C YIR024C YJL031C YJL032W YJL165C YKL002W YKR055W YLR025W YNL266W YOL053W YOR075W YOR327C | ECM15 LDB7 SEC17 ERS1 AHC2 HMRA2 HMRA1 NHP10 SLX5 RAD59 IWR1 MAF1 PET100 PPN1 TLG1 VPS60 ERG28 CAJ1 PET117 PCL6 AIM9 COG7 NBP35 SEC9 MSP1 UFD1 PET54 BNS1 OPI1 BET4 HAL5 DID4 RHO4 SNF7 AIM39 UFE1 SNC2 | Non-essential protein of unknown function, likely exists as tetramer, may be regulated by the binding of small-molecule ligands (possibly sulfate ions), may have a role in yeast cell-wall biogenesis | Component of the RSC chromatin remodeling complex; interacts with Rsc3p, Rsc30p, Npl6p, and Htl1p to form a module important for a broad range of RSC functions | Peripheral membrane protein required for vesicular transport between ER and Golgi and for the 'priming' step in homotypic vacuole fusion, part of the cis-SNARE complex; has similarity to alpha-SNAP | hypothetical protein | Protein of unknown function; localized to the cytoplasm; binds to Replication Protein A (RPA); YBR137W is not an essential gene | Protein with similarity to human cystinosin, which is a H(+)-driven transporter involved in L-cystine export from lysosomes and implicated in the disease cystinosis; contains seven transmembrane domains | Protein of unknown function, putative transcriptional regulator; proposed to be a Ada Histone acetyltransferase complex component; GFP tagged protein is localized to the cytoplasm and nucleus | Silenced copy of a2 at HMR; similarity to Alpha2p; required along with a1p for inhibiting expression of the HO endonuclease in a/alpha HO/HO diploid cells with an active mating-type interconversion system | Silenced copy of a1 at HMR; homeobox corepressor that interacts with Alpha2p to repress haploid-specific gene transcription in diploid cells | Protein related to mammalian high mobility group proteins; likely component of the INO80 complex, which is an ATP-dependent chromatin-remodeling complex | Subunit of the Slx5-Slx8 substrate-specific ubiquitin ligase complex; stimulated by prior attachment of SUMO to the substrate | Protein involved in the repair of double-strand breaks in DNA during vegetative growth via recombination and single-strand annealing; anneals complementary single-stranded DNA; homologous to Rad52p | Protein of unknown function, deletion causes hypersensitivity to the K1 killer toxin | Negative regulator of RNA polymerase III; component of several signaling pathways that repress polymerase III transcription in response to changes in cellular environment; targets the initiation factor TFIIIB | Chaperone that specifically facilitates the assembly of cytochrome c oxidase, integral to the mitochondrial inner membrane; interacts with a subcomplex of subunits VII, VIIa, and VIII (Cox7p, Cox9p, and Cox8p) but not with the holoenzyme | Vacuolar endopolyphosphatase with a role in phosphate metabolism; functions as a homodimer | Essential t-SNARE that forms a complex with Tlg2p and Vti1p and mediates fusion of endosome-derived vesicles with the late Golgi; binds the docking complex VFT (Vps fifty-three) through interaction with Vps51p | Cytoplasmic and vacuolar membrane protein involved in late endosome to vacuole transport; required for normal filament maturation during pseudohyphal growth; may function in targeting specific cargo proteins for degradation | Endoplasmic reticulum membrane protein, may facilitate protein-protein interactions between the Erg26p dehydrogenase and the Erg27p 3-ketoreductase and/or tether these enzymes to the ER, also interacts with Erg6p | Nuclear type II J heat shock protein of the E. coli dnaJ family, contains a leucine zipper-like motif, binds to non-native substrates for presentation to Ssa3p, may function during protein translocation, assembly and disassembly | Protein required for assembly of cytochrome c oxidase | Pho85p cyclin of the Pho80p subfamily; forms the major Glc8p kinase together with Pcl7p and Pho85p; involved in the control of glycogen storage by Pho85p; stabilized by Elongin C binding | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) | Component of the conserved oligomeric Golgi complex (Cog1p through Cog8p), a cytosolic tethering complex that functions in protein trafficking to mediate fusion of transport vesicles to Golgi compartments | Essential iron-sulfur cluster binding protein localized in the cytoplasm; forms a complex with Cfd1p that is involved in iron-sulfur protein assembly in the cytosol; similar to P-loop NTPases | t-SNARE protein important for fusion of secretory vesicles with the plasma membrane; similar to but not functionally redundant with Spo20p; SNAP-25 homolog | Mitochondrial protein involved in sorting of proteins in the mitochondria; putative membrane-spanning ATPase | Protein that interacts with Cdc48p and Npl4p, involved in recognition of polyubiquitinated proteins and their presentation to the 26S proteasome for degradation; involved in transporting proteins from the ER to the cytosol | Mitochondrial translational activator specific for the COX3 mRNA, acts together with Pet122p and Pet494p; also required for splicing of the COX1 intron AI5 beta; located in the mitochondrial inner membrane | Protein with some similarity to Spo12p; overexpression bypasses need for Spo12p, but not required for meiosis | Transcriptional regulator of a variety of genes; phosphorylation by protein kinase A stimulates Opi1p function in negative regulation of phospholipid biosynthetic genes; involved in telomere maintenance | Putative protein of unknown function | Protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; interacts with Arh1p, a mitochondrial oxidoreductase; deletion mutant has a respiratory growth defect | Alpha subunit of Type II geranylgeranyltransferase required for vesicular transport between the endoplasmic reticulum and the Golgi; provides a membrane attachment moiety to Rab-like proteins Ypt1p and Sec4p | Putative protein kinase; overexpression increases sodium and lithium tolerance, whereas gene disruption increases cation and low pH sensitivity and impairs potassium uptake, suggesting a role in regulation of Trk1p and/or Trk2p transporters | Class E Vps protein of the ESCRT-III complex, required for sorting of integral membrane proteins into lumenal vesicles of multivesicular bodies, and for delivery of newly synthesized vacuolar enzymes to the vacuole, involved in endocytosis | Non-essential small GTPase of the Rho/Rac subfamily of Ras-like proteins, likely to be involved in the establishment of cell polarity | One of four subunits of the endosomal sorting complex required for transport III (ESCRT-III); involved in the sorting of transmembrane proteins into the multivesicular body (MVB) pathway; recruited from the cytoplasm to endosomal membranes | Putative protein of unknown function; YOL053W is not an essential gene; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) | t-SNARE required for ER membrane fusion and vesicular traffic, integral membrane protein that constitutes with Sec20p and Use1p the trimeric acceptor for R/v-SNAREs on Golgi-derived vesicles at the ER; part of Dsl1p complex | Vesicle membrane receptor protein (v-SNARE) involved in the fusion between Golgi-derived secretory vesicles with the plasma membrane; member of the synaptobrevin/VAMP family of R-type v-SNARE proteins |
| 152 | 49 | 433 | 0.528 | 0.547 |  |  | 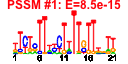 | 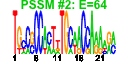 |  | YAL067C YCL023C YCL074W YCR081W YCR101C YDL187C YDR111C YDR131C YDR401W YFR057W YGR140W YHL022C YIL012W YIL032C YIL171W YJL028W YJL038C YKL115C YKL177W YKL223W YKR034W YLR013W YLR235C YLR307W YNR066C YOR183W YOR235W YPR064W YPR096C YAL037W YAR037W YBL108W YBR300C YCL076W YCR006C YCR049C YCR080W YCRX12W YDR136C YFL011W YHL018W YHR079BC YHR177W YIL132C YJR094C YJR146W YLR352W YLR445W YMR082C | SEO1 SRB8 ALT2 CBF2 SPO11 DAL80 GAT3 CDA1 FYV12 HXT10 CSM2 IME1 | Putative permease, member of the allantoate transporter subfamily of the major facilitator superfamily; mutation confers resistance to ethionine sulfoxide | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; essential for transcriptional regulation; involved in glucose repression | Putative protein of unknown function; localizes to the membrane fraction; YCR101C is not an essential gene | Putative alanine transaminase (glutamic pyruvic transaminase) | F-box protein, substrate-specific adaptor subunit that recruits substrates to a core ubiquitination complex | Putative protein of unknown function | Essential kinetochore protein, component of the CBF3 multisubunit complex that binds to the CDEIII region of the centromere; Cbf2p also binds to the CDEII region possibly forming a different multimeric complex, ubiquitinated in vivo | Meiosis-specific protein that initiates meiotic recombination by catalyzing the formation of double-strand breaks in DNA via a transesterification reaction; required for homologous chromosome pairing and synaptonemal complex formation | Protein of unknown function; may interact with ribosomes, based on co-purification experiments | Putative protein of unknown function; expression induced during sporulation and repressed during vegetative growth by Sum1p and Hst1p; similar to adjacent open reading frame, YJL037W | Negative regulator of genes in multiple nitrogen degradation pathways; expression is regulated by nitrogen levels and by Gln3p; member of the GATA-binding family, forms homodimers and heterodimers with Deh1p | Protein containing GATA family zinc finger motifs | Chitin deacetylase, together with Cda2p involved in the biosynthesis ascospore wall component, chitosan; required for proper rigidity of the ascospore wall | Putative membrane-localized protein of unknown function | Protein of unknown function, required for survival upon exposure to K1 killer toxin | Protein of unknown function that may interact with ribosomes, based on co-purification experiments | Putative hexose transporter, expressed at low levels and expression is repressed by glucose | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to mitochondria and is induced in response to the DNA-damaging agent MMS | Protein required for accurate chromosome segregation during meiosis | Master regulator of meiosis that is active only during meiotic events, activates transcription of early meiotic genes through interaction with Ume6p, degraded by the 26S proteasome following phosphorylation by Ime2p | Putative protein of unknown function with similarity to F-box proteins; interacts with Skp1p and Cdc53p; YLR352W is not an essential gene | Putative protein of unknown function; transcription is regulated by Ume6p and induced in response to alpha factor |
| 72 | 39 | 440 | 0.545 | 0.622 |  |  |  |  |  | YAR019C YAR066W YBR045C YBR226C YCR073C YDL206W YDL207W YDL216C YDR430C YGL163C YGR033C YGR212W YHL025W YHR014W YIL074W YIL076W YKL205W YKL206C YLL007C YLR173W YLR205C YLR392C YML107C YMR155W YNR005C YNR060W YNR068C YOR009W YOR044W YPL017C YPL089C YPR116W YDL217C YDR072C YIL067C YJR053W YMR154C YPL052W YPL148C | CDC15 GIP1 SSK22 GLE1 RRI1 CYM1 RAD54 TIM21 SLI1 SNF6 SPO13 SEC28 LOS1 ADD66 HMX1 PML39 FRE4 TIR4 IRC23 IRC15 RLM1 TIM22 IPT1 BFA1 RIM13 OAZ1 PPT2 | Protein kinase of the Mitotic Exit Network that is localized to the spindle pole bodies at late anaphase; promotes mitotic exit by directly switching on the kinase activity of Dbf2p | Putative GPI protein | Meiosis-specific regulatory subunit of the Glc7p protein phosphatase, regulates spore wall formation and septin organization, required for expression of some late meiotic genes and for normal localization of Glc7p | MAP kinase kinase kinase of the HOG1 mitogen-activated signaling pathway; functionally redundant with, and homologous to, Ssk2p; interacts with and is activated by Ssk1p; phosphorylates Pbs2p | Putative protein of unknown function; YDL206W is not an essential protein | Cytoplasmic nucleoporin required for polyadenylated RNA export but not for protein import; component of Nup82p nuclear pore subcomplex; contains a nuclear export signal | Catalytic subunit of the COP9 signalosome (CSN) complex that acts as an isopeptidase in cleaving the ubiquitin-like protein Nedd8 from SCF ubiquitin ligases; metalloendopeptidase involved in the adaptation to pheromone signaling | Lysine-specific metalloprotease of the mitochondrial intermembrane space, member of the pitrilysin family; degrades proteins and presequence peptides cleaved from imported proteins; required for normal mitochondrial morphology | DNA-dependent ATPase, stimulates strand exchange by modifying the topology of double-stranded DNA; involved in the recombinational repair of double-strand breaks in DNA during vegetative growth and meiosis; member of the SWI/SNF family | Constituent of the mitochondrial inner membrane presequence translocase (TIM23 complex); may regulate protein import by binding to both the translocase of the outer membrane (TOM) and presequence-associated motor (PAM) complexes | N-acetyltransferase, confers resistance to the sphingolipid biosynthesis inhibitor myriocin (ISP-1) by converting it into N-acetyl-myriocin, co-operates with Ypk1p in mediating resistance to myriocin | Subunit of the SWI/SNF chromatin remodeling complex involved in transcriptional regulation; functions interdependently in transcriptional activation with Snf2p and Snf5p | Meiosis-specific protein, involved in maintaining sister chromatid cohesion during meiosis I as well as promoting proper attachment of kinetochores to the spindle during meiosis I and meiosis II | Epsilon-COP subunit of the coatomer; regulates retrograde Golgi-to-ER protein traffic; stabilizes Cop1p, the alpha-COP and the coatomer complex; non-essential for cell growth | Nuclear pore protein involved in nuclear export of pre-tRNA | Protein involved in 20S proteasome assembly; forms a heterodimer with Pba1p that binds to proteasome precursors; similar to human PAC2 constituent of the PAC1-PAC2 complex involved in proteasome assembly | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLL007C is not an essential gene | Putative protein of unknown function | ER localized, heme-binding peroxidase involved in the degradation of heme; does not exhibit heme oxygenase activity despite similarity to heme oxygenases; expression regulated by AFT1 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR392C is not an essential gene | Protein required for nuclear retention of unspliced pre-mRNAs along with Mlp1p and Pml1p; anchored to nuclear pore complex via Mlp1p and Mlp2p; found with the subset of nuclear pores farthest from the nucleolus; may interact with ribosomes | Putative protein of unknown function; identified as interacting with Hsp82p in a high-throughput two-hybrid screen | Ferric reductase, reduces a specific subset of siderophore-bound iron prior to uptake by transporters; expression induced by low iron levels | Cell wall mannoprotein of the Srp1p/Tip1p family of serine-alanine-rich proteins; expressed under anaerobic conditions and required for anaerobic growth; transcription is also induced by cold shock | Putative protein of unknown function; green fluorescent protein (GFP)-fusion localizes to the ER; null mutant displays increased levels of spontaneous Rad52 foci | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; null mutant displays increased levels of spontaneous Rad52 foci | MADS-box transcription factor, component of the protein kinase C-mediated MAP kinase pathway involved in the maintenance of cell integrity; phosphorylated and activated by the MAP-kinase Slt2p | Component of the mitochondrial Tim54p-Tim22p complex involved in insertion of polytopic proteins into the inner membrane | Inositolphosphotransferase 1, involved in synthesis of mannose-(inositol-P)2-ceramide (M(IP)2C), which is the most abundant sphingolipid in cells, mutation confers resistance to the antifungals syringomycin E and DmAMP1 in some growth media | Uncharacterized protein of unknown function | Component of the GTPase-activating Bfa1p-Bub2p complex involved in multiple cell cycle checkpoint pathways that control exit from mitosis | Calpain-like cysteine protease involved in proteolytic activation of Rim101p in response to alkaline pH; has similarity to A. nidulans palB | Regulator of ornithine decarboxylase (Spe1p), antizyme that binds to Spe1p to regulate ubiquitin-independent degradation; ribosomal frameshifting during synthesis of Oaz1p and its ubiquitin-mediated degradation are both polyamine-regulated | Phosphopantetheine:protein transferase (PPTase), activates mitochondrial acyl carrier protein (Acp1p) by phosphopantetheinylation |
| 98 | 49 | 439 | 0.549 | 0.528 |  |  | 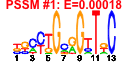 | 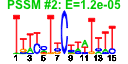 |  | YCL026C YDR520C YHR164C YJL019W YJL056C YJL113W YJL214W YJR108W YKL095W YKR005C YKR031C YKR101W YKR105C YLR213C YLR458W YML099C YMR057C YMR254C YNL179C YNR004W YOL074C YOL078W YOR029W YOR050C YOR190W YOR192C YOR268C YOR364W YOR366W YPL062W YPL174C YPL180W YPL187W YPL276W YPR122W YDL087C YKR102W YKR103W YLR072W YLR124W YML122C YMR320W YMR324C YNR060W YOL024W YOR107W YOR376W YPL130W YPL162C | HBN1 DNA2 MPS3 ZAP1 HXT8 ABM1 YJU2 SPO14 SIR1 CRR1 ARG81 AVO1 SPR1 THI72 NIP100 TCO89 MF(ALPHA)1 AXL1 LUC7 FLO10 NFT1 FRE4 RGS2 YOR376W-A SPO19 | Putative protein of unknown function; similar to bacterial nitroreductases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Putative transcription factor; contains the (Zn(II)2Cys6 motif; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; null mutant is viable and sensitive to caffeine | Essential tripartite DNA replication factor with single-stranded DNA-dependent ATPase, ATP-dependent nuclease, and helicase activities; required for Okazaki fragment processing; involved in DNA repair pathways; potential Cdc28p substrate | Essential integral membrane protein required for spindle pole body duplication and for nuclear fusion, localizes to the spindle pole body half bridge, interacts with DnaJ-like chaperone Jem1p and with centrin homolog Cdc31p | Zinc-regulated transcription factor, binds to zinc-responsive promoter elements to induce transcription of certain genes in the presence of zinc; regulates its own transcription; contains seven zinc-finger domains | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Protein of unknown function with similarity to hexose transporter family members, expression is induced by low levels of glucose and repressed by high levels of glucose | Protein of unknown function, required for normal microtubule organization | Essential protein required for pre-mRNA splicing; associates transiently with the spliceosomal NTC ('nineteen complex') and acts after Prp2p to promote the first catalytic reaction of splicing | Putative protein of unknown function | Phospholipase D, catalyzes the hydrolysis of phosphatidylcholine, producing choline and phosphatidic acid; involved in Sec14p-independent secretion; required for meiosis and spore formation; differently regulated in secretion and meiosis | Protein involved in repression of transcription at the silent mating-type loci HML and HMR; recruitment to silent chromatin requires interactions with Orc1p and with Sir4p, through a common Sir1p domain; binds to centromeric chromatin | Putative transporter of the Major Facilitator Superfamily (MFS) | Putative glycoside hydrolase of the spore wall envelope; required for normal spore wall assembly, possibly for cross-linking between the glucan and chitosan layers; expressed during sporulation | Zinc-finger transcription factor of the Zn(2)-Cys(6) binuclear cluster domain type, involved in the regulation of arginine-responsive genes; acts with Arg80p and Arg82p | Putative protein of unknown function; haploid disruptant exhibits slow growth rate on glucose-minimal medium at 15 C | Component of a membrane-bound complex containing the Tor2p kinase and other proteins, which may have a role in regulation of cell growth | Sporulation-specific exo-1,3-beta-glucanase; contributes to ascospore thermoresistance | Transporter of thiamine or related compound; shares sequence similarity with Thi7p | Putative protein of unknown function; sporulation is abnormal in homozygous diploid; YOR268C is not an essential gene | Large subunit of the dynactin complex, which is involved in partitioning the mitotic spindle between mother and daughter cells; putative ortholog of mammalian p150(glued) | Subunit of TORC1 (Tor1p or Tor2p-Kog1p-Lst8p-Tco89p), a complex that regulates growth in response to nutrient availability; cooperates with Ssd1p in the maintenance of cellular integrity; deletion strains are hypersensitive to rapamycin | Mating pheromone alpha-factor, made by alpha cells; interacts with mating type a cells to induce cell cycle arrest and other responses leading to mating; also encoded by MF(ALPHA)2, although MF(ALPHA)1 produces most alpha-factor | Haploid specific endoprotease that performs one of two N-terminal cleavages during maturation of a-factor mating pheromone; required for axial budding pattern of haploid cells | Essential protein associated with the U1 snRNP complex; splicing factor involved in recognition of 5' splice site; contains two zinc finger motifs; N-terminal zinc finger binds pre-mRNA | Lectin-like protein with similarity to Flo1p, thought to be involved in flocculation | Putative transporter of the multidrug resistance-associated protein (MRP) subfamily; adjacent ORFs YKR103W and YKR104W are merged in different strain backgrounds. | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern; YLR072W is not an esssential gene | Ferric reductase, reduces a specific subset of siderophore-bound iron prior to uptake by transporters; expression induced by low iron levels | Putative protein of unknown function, predicted to have thiol-disulfide oxidoreductase active site | Negative regulator of glucose-induced cAMP signaling; directly activates the GTPase activity of the heterotrimeric G protein alpha subunit Gpa2p | Putative protein of unknown function; identified by fungal homology and RT-PCR | Meiosis-specific protein of unknown function, involved in completion of nuclear divisions; identified as a weak high-copy suppressor of the spo1-1 ts mutation; putative GPI-dependent cell-wall protein | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the membrane of vacuole with cell cycle-correlated morphology |
| 134 | 56 | 440 | 0.555 | 0.540 |  |  | 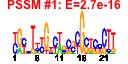 |  |  | YBL044W YBL073W YBR144C YCL075W YCLX03C YDL196W YDR028C YDR220C YEL010W YER135C YER181C YGL024W YGL034C YGL088W YGL118C YGR291C YHL005C YHR095W YHR212C YIL025C YIL149C YJL007C YJL083W YJL086C YKL198C YKL199C YLR162W YLR184W YLR233C YLR234W YLR366W YNL205C YOL166C YPR117W YAR040C YBL065W YCL026C YCLX07W YCRX09C YDL032W YDR320C YDR360W YGL176C YHL043W YJL056C YJL214W YKL031W YKL102C YKR041W YKR073C YLR054C YMR023C YMR069W YMR232W YOL074C YPR049C | REG1 MLP2 TAX4 POT1 PTK1 EST1 TOP3 HBN1 SWA2 ECM34 ZAP1 HXT8 OSW2 MSS1 NAT4 FUS2 ATG11 | Putative protein of unknown function; YBL044W is not an essential protein | Regulatory subunit of type 1 protein phosphatase Glc7p, involved in negative regulation of glucose-repressible genes | Myosin-like protein associated with the nuclear envelope, connects the nuclear pore complex with the nuclear interior; involved in the Tel1p pathway that controls telomere length | Protein involved in regulation of phosphatidylinositol 4,5-bisphosphate concentrations; Irs4p and Tax4p bind and activate the phosphatase Inp51p | 3-ketoacyl-CoA thiolase with broad chain length specificity, cleaves 3-ketoacyl-CoA into acyl-CoA and acetyl-CoA during beta-oxidation of fatty acids | Putative serine/threonine protein kinase that regulates spermine uptake; involved in polyamine transport; possible mitochondrial protein | Putative protein of unknown function; overexpression confers resistance to the antimicrobial peptide MiAMP1 | TLC1 RNA-associated factor involved in telomere length regulation as the recruitment subunit of the telomerase holoenzyme, has a possible role in activating Est2p-TLC1-RNA bound to the telomere | DNA Topoisomerase III, conserved protein that functions in a complex with Sgs1p and Rmi1p to relax single-stranded negatively-supercoiled DNA preferentially, involved in telomere stability and regulation of mitotic recombination | Putative protein of unknown function | Putative protein of unknown function; similar to bacterial nitroreductases; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Auxilin-like protein involved in vesicular transport; clathrin-binding protein required for uncoating of clathrin-coated vesicles | Putative protein of unknown function; deletion mutant is viable and has no detectable phenotype | Putative protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Zinc-regulated transcription factor, binds to zinc-responsive promoter elements to induce transcription of certain genes in the presence of zinc; regulates its own transcription; contains seven zinc-finger domains | Protein of unknown function with similarity to hexose transporter family members, expression is induced by low levels of glucose and repressed by high levels of glucose | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Protein of unknown function proposed to be involved in the assembly of the spore wall | Mitochondrial protein, forms a heterodimer complex with Mto1p that performs the 5-carboxymethylaminomethyl modification of the wobble uridine base in mitochondrial tRNAs; similar to human GTPBP3 | N alpha-acetyl-transferase, involved in acetylation of the N-terminal residues of histones H4 and H2A | Cytoplasmic protein localized to the shmoo tip; required for the alignment of parental nuclei before nuclear fusion during mating | Peripheral membrane protein required for delivery of aminopeptidase I (Lap4p) to the vacuole in the cytoplasm-to-vacuole targeting pathway; also required for peroxisomal degradation (pexophagy) |
| 189 | 44 | 445 | 0.557 | 0.572 |  |  |  |  |  | YBR138C YDR538W YER081W YER116C YFR016C YGL197W YHL010C YHR160C YIL156W YMR257C YNR047W YPR027C YPR160W YAR035W YBL084C YDR082W YDR131C YDR176W YDR181C YDR386W YDR458C YEL072W YER024W YGL073W YGL164C YGL180W YGR125W YGR247W YHL022C YHR131C YHR150W YIL066C YIR028W YIR031C YJL106W YJR152W YLL003W YLR082C YOR034C YOR156C YOR328W YPL092W YPR007C YPR083W | PAD1 SER3 SLX8 MDS3 PEX18 YIL156W-B PET111 GPH1 YAT1 CDC27 STN1 NGG1 SAS4 MUS81 HEH2 RMD6 YAT2 HSF1 YRB30 ATG1 CPD1 SPO11 PEX28 RNR3 DAL4 DAL7 IME2 DAL5 SFI1 SRL2 AKR2 NFI1 PDR10 SSU1 REC8 MDM36 | Cytoplasmic protein of unknown function, potentially phosphorylated by Cdc28p; YBR138C is not an essential gene | Phenylacrylic acid decarboxylase, confers resistance to cinnamic acid, decarboxylates aromatic carboxylic acids to the corresponding vinyl derivatives; homolog of E. coli UbiX | 3-phosphoglycerate dehydrogenase, catalyzes the first step in serine and glycine biosynthesis; isozyme of Ser33p | Subunit of the Slx5-Slx8 substrate-specific ubiquitin ligase complex; stimulated by prior attachment of SUMO to the substrate | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and bud; interacts with Spa2p; YFL016C is not an essential gene | Protein with an N-terminal kelch-like domain, putative negative regulator of early meiotic gene expression; required, with Pmd1p, for growth under alkaline conditions | Putative protein of unknown function, contains a zinc finger region and has homology to human BRAP2, which is a cytoplasmic protein that binds nuclear localization sequences | Peroxin required for targeting of peroxisomal matrix proteins containing PTS2; interacts with Pex7p; partially redundant with Pex21p | Putative protein of unknown function, originally identified based on homology to Ashbya gossypii and other related yeasts | Mitochondrial translational activator specific for the COX2 mRNA; located in the mitochondrial inner membrane | Putative protein kinase that, when overexpressed, interferes with pheromone-induced growth arrest; localizes to the cytoplasm; potential Cdc28p substrate | Putative protein of unknown function | Non-essential glycogen phosphorylase required for the mobilization of glycogen, activity is regulated by cyclic AMP-mediated phosphorylation, expression is regulated by stress-response elements and by the HOG MAP kinase pathway | Outer mitochondrial carnitine acetyltransferase, minor ethanol-inducible enzyme involved in transport of activated acyl groups from the cytoplasm into the mitochondrial matrix; phosphorylated | Subunit of the Anaphase-Promoting Complex/Cyclosome (APC/C), which is a ubiquitin-protein ligase required for degradation of anaphase inhibitors, including mitotic cyclins, during the metaphase/anaphase transition | Telomere end-binding and capping protein, plays a key role with Pol12p in linking telomerase action with completion of lagging strand synthesis, and in a regulatory step required for telomere capping | F-box protein, substrate-specific adaptor subunit that recruits substrates to a core ubiquitination complex | Transcriptional regulator involved in glucose repression of Gal4p-regulated genes; component of transcriptional adaptor and histone acetyltransferase complexes, the ADA complex, the SAGA complex, and the SLIK complex | Subunit of the SAS complex (Sas2p, Sas4p, Sas5p), which acetylates free histones and nucleosomes and regulates transcriptional silencing; required for the HAT activity of Sas2p | Helix-hairpin-helix protein, involved in DNA repair and replication fork stability; functions as an endonuclease in complex with Mms4p; interacts with Rad54p | Inner nuclear membrane (INM) protein; contains helix-extension-helix (HEH) motif, nuclear localization signal sequence; targeting to the INM requires the Srp1p-Kap95p karyopherins and the Ran cycle | Protein required for sporulation | Carnitine acetyltransferase; has similarity to Yat1p, which is a carnitine acetyltransferase associated with the mitochondrial outer membrane | Trimeric heat shock transcription factor, activates multiple genes in response to stresses that include hyperthermia; recognizes variable heat shock elements (HSEs) consisting of inverted NGAAN repeats; posttranslationally regulated | RanGTP-binding protein, inhibits RanGAP1 (Rna1p)-mediated GTP hydrolysis of RanGTP (Gsp1p); shares similarity to proteins in other fungi but not in higher eukaryotes | Protein serine/threonine kinase, required for vesicle formation during autophagy and the cytoplasm-to-vacuole targeting (Cvt) pathway; structurally required for pre-autophagosome formation; forms a complex with Atg13p and Atg17p | Putative protein of unknown function; deletion mutant has decreased rapamycin resistance but normal wormannin resistance; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Cyclic nucleotide phosphodiesterase, hydrolyzes ADP-ribose 1'', 2''-cyclic phosphate to ADP-ribose 1''-phosphate; may have a role in tRNA splicing; no detectable phenotype is conferred by null mutation or by overexpression | Meiosis-specific protein that initiates meiotic recombination by catalyzing the formation of double-strand breaks in DNA via a transesterification reaction; required for homologous chromosome pairing and synaptonemal complex formation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Peroxisomal integral membrane peroxin, involved in the regulation of peroxisomal size, number and distribution; genetic interactions suggest that Pex28p and Pex29p act at steps upstream of those mediated by Pex30p, Pex31p, and Pex32p | Ribonucleotide-diphosphate reductase (RNR), large subunit; the RNR complex catalyzes the rate-limiting step in dNTP synthesis and is regulated by DNA replication and DNA damage checkpoint pathways via localization of the small subunits | Allantoin permease; expression sensitive to nitrogen catabolite repression and induced by allophanate, an intermediate in allantoin degradation | Malate synthase, role in allantoin degradation unknown; expression sensitive to nitrogen catabolite repression and induced by allophanate, an intermediate in allantoin degradation | Serine/threonine protein kinase involved in activation of meiosis, associates with Ime1p and mediates its stability, activates Ndt80p; IME2 expression is positively regulated by Ime1p | Allantoin permease; ureidosuccinate permease; expression is constitutive but sensitive to nitrogen catabolite repression | Centrin (Cdc31p)-binding protein required for spindle pole body (SPB) duplication, localizes to the half-bridge of the SPB, required for progression through G(2)-M transition, has similarity to Xenopus laevis XCAP-C | Protein of unknown function; overexpression suppresses the lethality caused by a rad53 null mutation | Ankyrin repeat-containing protein similar to Akr1p; member of a family of putative palmitoyltransferases containing an Asp-His-His-Cys-cysteine rich (DHHC-CRD) domain; possibly involved in constitutive endocytosis of Ste3p | SUMO ligase, catalyzes the covalent attachment of SUMO (Smt3p) to proteins; involved in maintenance of proper telomere length | ATP-binding cassette (ABC) transporter, multidrug transporter involved in the pleiotropic drug resistance network; regulated by Pdr1p and Pdr3p | Plasma membrane sulfite pump involved in sulfite metabolism and required for efficient sulfite efflux; major facilitator superfamily protein | Meiosis-specific component of sister chromatid cohesion complex; maintains cohesion between sister chromatids during meiosis I; maintains cohesion between centromeres of sister chromatids until meiosis II; homolog of S. pombe Rec8p | Protein required for normal mitochondrial morphology and inheritance |
| 269 | 43 | 444 | 0.570 | 0.559 | 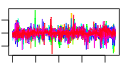 |  | 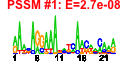 | 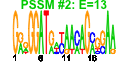 |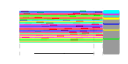  | YLR411W YAR018C YAR019C YAR042W YBR097W YDL146W YDR217C YGL075C YIL013C YIL100W YIR019C YJR128W YKL115C YKL132C YKL198C YKL199C YLR040C YLR087C YML083C YMR117C YMR254C YNL128W YNL198C YNL278W YNL319W YNR057C YNR058W YNR070W YOL078W YOR024W YOR050C YOR072W YOR083W YOR291W YOR343C YOR364W YPL016W YPL021W YPL027W YPL040C YPL062W YPL121C YPR117W | CTR3 KIN3 CDC15 SWH1 VPS15 LDB17 RAD9 MPS2 PDR11 MUC1 RMA1 POT1 PTK1 CSF1 SPC24 TEP1 CAF120 BIO4 BIO3 AVO1 YOR072W-B WHI5 SWI1 ECM23 SMA1 ISM1 MEI5 | High-affinity copper transporter of the plasma membrane, acts as a trimer; gene is disrupted by a Ty2 transposon insertion in many laboratory strains of S. cerevisiae | Nonessential protein kinase with unknown cellular role | Protein kinase of the Mitotic Exit Network that is localized to the spindle pole bodies at late anaphase; promotes mitotic exit by directly switching on the kinase activity of Dbf2p | Protein similar to mammalian oxysterol-binding protein; contains ankyrin repeats; localizes to the Golgi and the nucleus-vacuole junction | Myristoylated serine/threonine protein kinase involved in vacuolar protein sorting; functions as a membrane-associated complex with Vps34p; active form recruits Vps34p to the Golgi membrane; interacts with the GDP-bound form of Gpa1p | Protein of unknown function; GFP-fusion protein localizes to the cell periphery, cytoplasm, bud, and bud neck; null mutant shows a reduced affinity for the alcian blue dye suggesting a decreased net negative charge of the cell surface | DNA damage-dependent checkpoint protein, required for cell-cycle arrest in G1/S, intra-S, and G2/M; transmits checkpoint signal by activating Rad53p and Chk1p; hyperphosphorylated by Mec1p and Tel1p; potential Cdc28p substrate | Essential membrane protein localized at the nuclear envelope and spindle pole body (SPB), required for insertion of the newly duplicated SPB into the nuclear envelope; potentially phosphorylated by Cdc28p | ATP-binding cassette (ABC) transporter, multidrug transporter involved in multiple drug resistance; mediates sterol uptake when sterol biosynthesis is compromisedregulated by Pdr1p; required for anaerobic growth | GPI-anchored cell surface glycoprotein (flocculin) required for pseudohyphal formation, invasive growth, flocculation, and biofilms; transcriptionally regulated by the MAPK pathway (via Ste12p and Tec1p) and the cAMP pathway (via Flo8p) | Putative dihydrofolate synthetase; has similarity to Fol3p and to E. coli folylpolyglutamate synthetase/dihydrofolate synthetase; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | 3-ketoacyl-CoA thiolase with broad chain length specificity, cleaves 3-ketoacyl-CoA into acyl-CoA and acetyl-CoA during beta-oxidation of fatty acids | Putative serine/threonine protein kinase that regulates spermine uptake; involved in polyamine transport; possible mitochondrial protein | Putative protein of unknown function; localizes to the cell wall; predicted to be a GPI-attached protein; upregulated by Mcm1p-Alpha1p transcription factor; partially overlaps the dubious ORF YLR041W; YLR040C is not essential | Protein required for fermentation at low temperature; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; strong increase in transcript abundance during anaerobic growth compared to aerobic growth; cells deleted for YML083C do not exhibit growth defects in anerobic or anaerobic conditions | Component of the evolutionarily conserved kinetochore-associated Ndc80 complex (Ndc80p-Nuf2p-Spc24p-Spc25p); involved in chromosome segregation, spindle checkpoint activity and kinetochore clustering | Homolog of human tumor suppressor gene PTEN/MMAC1/TEP1 that has lipid phosphatase activity and is linked to the phosphatidylinositol signaling pathway; plays a role in normal sporulation | Part of the evolutionarily-conserved CCR4-NOT transcriptional regulatory complex involved in controlling mRNA initiation, elongation, and degradation | Dethiobiotin synthetase, catalyzes the third step in the biotin biosynthesis pathway; BIO4 is in a cluster of 3 genes (BIO3, BIO4, and BIO5) that mediate biotin synthesis; expression appears to be repressed at low iron levels | 7,8-diamino-pelargonic acid aminotransferase (DAPA), catalyzes the second step in the biotin biosynthesis pathway; BIO3 is in a cluster of 3 genes (BIO3, BIO4, and BIO5) that mediate biotin synthesis | Putative transporter of the ATP-binding cassette (ABC) family, implicated in pleiotropic drug resistance; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Component of a membrane-bound complex containing the Tor2p kinase and other proteins, which may have a role in regulation of cell growth | Putative protein of unknown function; identified by expression profiling and mass spectrometry | Repressor of G1 transcription that binds to SCB binding factor (SBF) at SCB target promoters in early G1; phosphorylation of Whi5p by the CDK, Cln3p/Cdc28p relieves repression and promoter binding by Whi5; periodically expressed in G1 | Putative protein of unknown function; shares sequence similarity with the type V P-type ATPase Spf1p; YOR291W is not an essential protein | Subunit of the SWI/SNF chromatin remodeling complex, which regulates transcription by remodeling chromosomes; required for transcription of many genes, including ADH1, ADH2, GAL1, HO, INO1 and SUC2 | Non-essential protein of unconfirmed function; affects pre-rRNA processing, may act as a negative regulator of the transcription of genes involved in pseudohyphal growth; homologous to Srd1p | Protein of unknown function involved in the assembly of the prospore membrane during sporulation | Mitochondrial isoleucyl-tRNA synthetase, null mutant is deficient in respiratory growth | Meiosis specific protein involved in DMC1-dependent meiotic recombination, forms heterodimer with Sae3p; proposed to be an assembly factor for Dmc1p | Putative protein of unknown function |
| 254 | 40 | 436 | 0.575 | 0.584 | 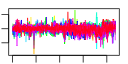 |  |  |  |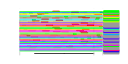  | YGL255W YJL157C YMR202W YOL136C YBR085W YBR086C YBR187W YDR033W YDR297W YDR302W YEL009C YEL021W YFL045C YGL055W YGR108W YGR175C YGR189C YGR191W YGR282C YHR133C YHR143W YJL186W YLR056W YLR229C YLR300W YMR016C YMR079W YNL043C YNL044W YNL145W YNL190W YNL217W YOL056W YOL061W YOL128C YOR344C YOR355W YPL036W YPL117C YPR053C | ZRT1 FAR1 ERG2 PFK27 AAC3 IST2 GDT1 MRH1 SUR2 GPI11 GCN4 URA3 SEC53 OLE1 CLB1 ERG1 CRH1 HIP1 BGL2 NSG1 DSE2 MNN5 ERG3 CDC42 EXG1 SOK2 SEC14 YIP3 MFA2 GPM3 PRS5 YGK3 TYE7 GDS1 PMA2 IDI1 | High-affinity zinc transporter of the plasma membrane, responsible for the majority of zinc uptake; transcription is induced under low-zinc conditions by the Zap1p transcription factor | Cyclin-dependent kinase inhibitor that mediates cell cycle arrest in response to pheromone; also forms a complex with Cdc24p, Ste4p, and Ste18p that may specify the direction of polarized growth during mating; potential Cdc28p substrate | C-8 sterol isomerase, catalyzes the isomerization of the delta-8 double bond to the delta-7 position at an intermediate step in ergosterol biosynthesis | 6-phosphofructo-2-kinase, catalyzes synthesis of fructose-2,6-bisphosphate; inhibited by phosphoenolpyruvate and sn-glycerol 3-phosphate, expression induced by glucose and sucrose, transcriptional regulation involves protein kinase A | Mitochondrial inner membrane ADP/ATP translocator, exchanges cytosolic ADP for mitochondrially synthesized ATP; expressed under anaerobic conditions; similar to Pet9p and Aac1p; has roles in maintenance of viability and in respiration | Plasma membrane protein that may be involved in osmotolerance, localizes to the mother cell in small-budded cells and to the bud in medium- and large-budded cells; mRNA is transported to the bud tip by an actomyosin-driven process | Putative protein of unknown function; expression is reduced in a gcr1 null mutant; GFP-fusion protein localizes to the vacuole; expression pattern and physical interactions suggest a possible role in ribosome biogenesis | Protein that localizes primarily to the plasma membrane, also found at the nuclear envelope; the authentic, non-tagged protein is detected in mitochondria in a phosphorylated state; has similarity to Hsp30p and Yro2p | Sphinganine C4-hydroxylase, catalyses the conversion of sphinganine to phytosphingosine in sphingolipid biosyntheis | ER membrane protein involved in a late step of glycosylphosphatidylinositol (GPI) anchor assembly; involved in the addition of phosphoethanolamine to the multiply mannosylated GPI intermediate; human PIG-Fp is a functional homolog | Transcriptional activator of amino acid biosynthetic genes in response to amino acid starvation; expression is tightly regulated at both the transcriptional and translational levels | Orotidine-5'-phosphate (OMP) decarboxylase, catalyzes the sixth enzymatic step in the de novo biosynthesis of pyrimidines, converting OMP into uridine monophosphate (UMP); converts 5-FOA into 5-fluorouracil, a toxic compound | Phosphomannomutase, involved in synthesis of GDP-mannose and dolichol-phosphate-mannose; required for folding and glycosylation of secretory proteins in the ER lumen | Delta(9) fatty acid desaturase, required for monounsaturated fatty acid synthesis and for normal distribution of mitochondria | B-type cyclin involved in cell cycle progression; activates Cdc28p to promote the transition from G2 to M phase; accumulates during G2 and M, then targeted via a destruction box motif for ubiquitin-mediated degradation by the proteasome | Squalene epoxidase, catalyzes the epoxidation of squalene to 2,3-oxidosqualene; plays an essential role in the ergosterol-biosynthesis pathway and is the specific target of the antifungal drug terbinafine | Cell wall protein that functions in the transfer of chitin to beta(1-6)glucan, putative chitin transglycosidase | High-affinity histidine permease, also involved in the transport of manganese ions | Endo-beta-1,3-glucanase, major protein of the cell wall, involved in cell wall maintenance | Protein involved in regulation of sterol biosynthesis; specifically stabilizes Hmg2p, one of two HMG-CoA isoenzymes that catalyze the rate-limiting step in sterol biosynthesis; homolog of mammalian INSIG proteins | Daughter cell-specific secreted protein with similarity to glucanases, degrades cell wall from the daughter side causing daughter to separate from mother; expression is repressed by cAMP | Alpha-1,2-mannosyltransferase, responsible for addition of the second alpha-1,2-linked mannose of the branches on the mannan backbone of oligosaccharides, localizes to an early Golgi compartment | C-5 sterol desaturase, catalyzes the introduction of a C-5(6) double bond into episterol, a precursor in ergosterol biosynthesis; mutants are viable, but cannot grow on non-fermentable carbon sources | Small rho-like GTPase, essential for establishment and maintenance of cell polarity; mutants have defects in the organization of actin and septins | Major exo-1,3-beta-glucanase of the cell wall, involved in cell wall beta-glucan assembly; exists as three differentially glycosylated isoenzymes | Nuclear protein that plays a regulatory role in the cyclic AMP (cAMP)-dependent protein kinase (PKA) signal transduction pathway; negatively regulates pseudohyphal differentiation; homologous to several transcription factors | Phosphatidylinositol/phosphatidylcholine transfer protein involved in coordinate regulation of PtdIns and PtdCho metabolism, products of which are regulators in Golgi to plasma membrane transport; functionally homologous to mammalian PITPs | Protein localized to COPII vesicles, proposed to be involved in ER to Golgi transport; interacts with members of the Rab GTPase family and Yip1p; also interacts with Rtn1p | Mating pheromone a-factor, made by a cells; interacts with alpha cells to induce cell cycle arrest and other responses leading to mating; biogenesis involves C-terminal modification, N-terminal proteolysis, and export; also encoded by MFA1 | Cell wall protein of unknown function; proposed role as a hydrophilin induced by osmotic stress; contains a putative GPI-attachment site | hypothetical protein | Homolog of Gpm1p phosphoglycerate mutase, which converts 3-phosphoglycerate to 2-phosphoglycerate in glycolysis; may be non-functional derivative of a gene duplication event | 5-phospho-ribosyl-1(alpha)-pyrophosphate synthetase, involved in nucleotide, histidine, and tryptophan biosynthesis; one of five related enzymes, which are active as heteromultimeric complexes | Protein kinase related to mammalian glycogen synthase kinases of the GSK-3 family; GSK-3 homologs (Mck1p, Rim11p, Mrk1p, Ygk3p) are involved in control of Msn2p-dependent transcription of stress responsive genes and in protein degradation | Serine-rich protein that contains a basic-helix-loop-helix (bHLH) DNA binding motif; binds E-boxes of glycolytic genes and contributes to their activation; may function as a transcriptional activator in Ty1-mediated gene expression | Protein of unknown function, required for growth on glycerol as a carbon source; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Plasma membrane H+-ATPase, isoform of Pma1p, involved in pumping protons out of the cell; regulator of cytoplasmic pH and plasma membrane potential | Isopentenyl diphosphate:dimethylallyl diphosphate isomerase (IPP isomerase), catalyzes an essential activation step in the isoprenoid biosynthetic pathway; required for viability |
| 243 | 30 | 440 | 0.575 | 0.591 |  |  |  | 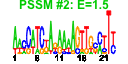 |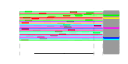  | YJR119C YAL062W YBR099C YBR201W YDL009C YDL011C YDL026W YDL054C YDL113C YDR203W YDR357C YDR511W YDR542W YEL073C YGR102C YGR196C YHL027W YHR022C YJL022W YJL042W YJL199C YJR115W YKL123W YML002W YMR007W YMR253C YOL055C YOR134W YPL159C YPL177C | JHD2 GDH3 DER1 MCH1 ATG20 ACN9 PAU10 FYV8 RIM1 MHP1 THI20 BAG7 PET20 CUP9 | JmjC domain family histone demethylase specific for H3-K4 (lysine at position 4 of the histone H3 protein); removes methyl groups specifically added by Set1p methyltransferase | NADP(+)-dependent glutamate dehydrogenase, synthesizes glutamate from ammonia and alpha-ketoglutarate; rate of alpha-ketoglutarate utilization differs from Gdh1p; expression regulated by nitrogen and carbon sources | Endoplasmic reticulum membrane protein, required for ER-associated protein degradation of misfolded or unassembled proteins; N- and C- termini protrude into the cytoplasm, has similarity to Dfm1p | Protein with similarity to mammalian monocarboxylate permeases, which are involved in transport of monocarboxylic acids across the plasma membrane; mutant is not deficient in monocarboxylate transport | Protein required for transport of aminopeptidase I (Lap4p) through the cytoplasm-to-vacuole targeting pathway; binds phosphatidylinositol-3-phosphate, involved in localization of membranes to the preautophagosome, potential Cdc28p substrate | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Protein of the mitochondrial intermembrane space, required for acetate utilization and gluconeogenesis; has orthologs in higher eukaryotes | hypothetical protein | Putative protein of unknown function; located adjacent to ARS503 and the telomere on the left arm of chromosome V; regulated by inositol/choline | Putative protein of unknown function; transposon insertion mutant is salt sensitive and deletion mutant has growth defects; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein of unknown function, required for survival upon exposure to K1 killer toxin | Single-stranded DNA-binding protein essential for mitochondrial genome maintenance; involved in mitochondrial DNA replication | Putative protein of unknown function; YHR022C is not an essential gene. | Microtubule-associated protein involved in assembly and stabilization of microtubules; overproduction results in cell cycle arrest at G2 phase; similar to Drosophila protein MAP and to mammalian MAP4 proteins | Putative protein of unknown function | Putative protein of unknown function; expression induced by heat and by calcium shortage | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern; YMR253C is not an essential gene | Multifunctional protein with both hydroxymethylpyrimidine kinase and thiaminase activities; involved in thiamine biosynthesis and also in thiamine degradation; member of a gene family with THI21 and THI22; functionally redundant with Thi21p | Rho GTPase activating protein (RhoGAP), stimulates the intrinsic GTPase activity of Rho1p, which plays a role in actin cytoskeleton organization and control of cell wall synthesis; structurally and functionally related to Sac7p | Mitochondrial protein, required for respiratory growth under some conditions and for stability of the mitochondrial genome | Homeodomain-containing transcriptional repressor of PTR2, which encodes a major peptide transporter; imported peptides activate ubiquitin-dependent proteolysis, resulting in degradation of Cup9p and de-repression of PTR2 transcription |
| 19 | 46 | 448 | 0.587 | 0.652 |  |  | 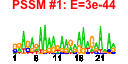 | 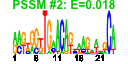 |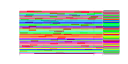  | YAL005C YAL026C YAL040C YBR263W YCR053W YDL040C YDL166C YDR508C YEL046C YER036C YER072W YFL004W YFR037C YGR210C YGR211W YHR005C YHR047C YHR061C YHR084W YIL011W YIL131C YIR021W YJL196C YKL184W YKR042W YKR043C YKR044W YLL018C YLR023C YLR172C YLR190W YLR237W YLR274W YMR012W YMR058W YMR246W YMR296C YNR018W YNR044W YOR047C YOR212W YOR306C YOR311C YPR088C YPR089W YPR114W | SSA1 DRS2 CLN3 SHM1 THR4 NAT1 FAP7 GNP1 GLY1 ARB1 VTC1 VTC2 RSC8 ZPR1 GPA1 ATP8 GIC1 STE12 TIR3 FKH1 MRS1 ELO1 SPE1 UTH1 UIP5 COX19 IZH3 DPH5 MMR1 THI7 BOI1 CLU1 FET3 FAA4 LCB1 AIM38 AGA1 STD1 STE4 MCH5 HSD1 SRP54 | ATPase involved in protein folding and nuclear localization signal (NLS)-directed nuclear transport; member of heat shock protein 70 (HSP70) family; forms a chaperone complex with Ydj1p; localized to the nucleus, cytoplasm, and cell wall | Aminophospholipid translocase (flippase) that maintains membrane lipid asymmetry in post-Golgi secretory vessicles; contributes to clathrin-coated vesicle formation and endocytosis; mutations in human homolog ATP8B1 result in liver disease | G1 cyclin involved in cell cycle progression; activates Cdc28p kinase to promote the G1 to S phase transition; plays a role in regulating transcription of the other G1 cyclins, CLN1 and CLN2; regulated by phosphorylation and proteolysis | Mitochondrial serine hydroxymethyltransferase, converts serine to glycine plus 5,10 methylenetetrahydrofolate; involved in generating precursors for purine, pyrimidine, amino acid, and lipid biosynthesis; reverse reaction generates serine | Threonine synthase, conserved protein that catalyzes formation of threonine from 0-phosphohomoserine; expression is regulated by the GCN4-mediated general amino acid control pathway | Subunit of the N-terminal acetyltransferase NatA (Nat1p, Ard1p, Nat5p); N-terminally acetylates many proteins, which influences multiple processes such as the cell cycle, heat-shock resistance, mating, sporulation, and telomeric silencing | Essential NTPase required for small ribosome subunit synthesis, mediates processing of the 20S pre-rRNA at site D in the cytoplasm but associates only transiently with 43S preribosomes via Rps14p, may be the endonuclease for site D | High-affinity glutamine permease, also transports Leu, Ser, Thr, Cys, Met and Asn; expression is fully dependent on Grr1p and modulated by the Ssy1p-Ptr3p-Ssy5p (SPS) sensor of extracellular amino acids | Threonine aldolase, catalyzes the cleavage of L-allo-threonine and L-threonine to glycine; involved in glycine biosynthesis | ATPase of the ATP-binding cassette (ABC) family involved in 40S and 60S ribosome biogenesis, has similarity to Gcn20p; shuttles from nucleus to cytoplasm, physically interacts with Tif6p, Lsg1p | Vacuolar transporter chaperon (VTC) involved in distributing V-ATPase and other membrane proteins; together with other VTC proteins, forms a heterotetrameric complex that associates with the SNARE Nyv1p and the V0 sector of the V-ATPase | Vacuolar membrane protein involved in vacuolar polyphosphate accumulation; functions as a regulator of vacuolar H+-ATPase activity and vacuolar transporter chaperones; involved in protein localization and non-autophagic vacuolar fusion | Component of the RSC chromatin remodeling complex; essential for viability and mitotic growth; homolog of SWI/SNF subunit Swi3p, but unlike Swi3p, does not activate transcription of reporters | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Essential protein with two zinc fingers, present in the nucleus of growing cells but relocates to the cytoplasm in starved cells via a process mediated by Cpr1p; binds to translation elongation factor eEF-1 (Tef1p) | GTP-binding alpha subunit of the heterotrimeric G protein that couples to pheromone receptors; negatively regulates the mating pathway by sequestering G(beta)gamma and by triggering an adaptive response; activates Vps34p at the endosome | Subunit 8 of the F0 sector of mitochondrial inner membrane F1-F0 ATP synthase, encoded on the mitochondrial genome | Protein of unknown function involved in initiation of budding and cellular polarization, interacts with Cdc42p via the Cdc42/Rac-interactive binding (CRIB) domain | Transcription factor that is activated by a MAP kinase signaling cascade, activates genes involved in mating or pseudohyphal/invasive growth pathways; cooperates with Tec1p transcription factor to regulate genes specific for invasive growth | Cell wall mannoprotein of the Srp1p/Tip1p family of serine-alanine-rich proteins; expressed under anaerobic conditions and required for anaerobic growth | Forkhead family transcription factor with a minor role in the expression of G2/M phase genes; negatively regulates transcriptional elongation; positive role in chromatin silencing at HML and HMR; regulates donor preference during switching | Protein required for the splicing of two mitochondrial group I introns (BI3 in COB and AI5beta in COX1); forms a splicing complex, containing four subunits of Mrs1p and two subunits of the BI3-encoded maturase, that binds to the BI3 RNA | Elongase I, medium-chain acyl elongase, catalyzes carboxy-terminal elongation of unsaturated C12-C16 fatty acyl-CoAs to C16-C18 fatty acids | Ornithine decarboxylase, catalyzes the first step in polyamine biosynthesis; degraded in a proteasome-dependent manner in the presence of excess polyamines | Mitochondrial outer membrane and cell wall localized SUN family member required for mitochondrial autophagy; involved in the oxidative stress response, life span during starvation, mitochondrial biogenesis, and cell death | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Protein of unknown function that interacts with Ulp1p, a Ubl (ubiquitin-like protein)-specific protease for Smt3p protein conjugates | Protein required for cytochrome c oxidase assembly, located in the cytosol and mitochondrial intermembrane space; putative copper metallochaperone that delivers copper to cytochrome c oxidase | Membrane protein involved in zinc metabolism, member of the four-protein IZH family, expression induced by zinc deficiency; deletion reduces sensitivity to elevated zinc and shortens lag phase, overexpression reduces Zap1p activity | Methyltransferase required for synthesis of diphthamide, which is a modified histidine residue of translation elongation factor 2 (Eft1p or Eft2p); not essential for viability; GFP-Dph5p fusion protein localizes to the cytoplasm | Phosphorylated protein of the mitochondrial outer membrane, localizes only to mitochondria of the bud; interacts with Myo2p to mediate mitochondrial distribution to buds; mRNA is targeted to the bud via the transport system involving She2p | Plasma membrane transporter responsible for the uptake of thiamine, member of the major facilitator superfamily of transporters; mutation of human ortholog causes thiamine-responsive megaloblastic anemia | Protein implicated in polar growth, functionally redundant with Boi2p; interacts with bud-emergence protein Bem1p; contains an SH3 (src homology 3) domain and a PH (pleckstrin homology) domain | eIF3 component of unknown function; deletion causes defects in mitochondrial organization but not in growth or translation initiation, can rescue cytokinesis and mitochondrial organization defects of the Dictyostelium cluA- mutant | Ferro-O2-oxidoreductase required for high-affinity iron uptake and involved in mediating resistance to copper ion toxicity, belongs to class of integral membrane multicopper oxidases | Long chain fatty acyl-CoA synthetase, regulates protein modification during growth in the presence of ethanol, functions to incorporate palmitic acid into phospholipids and neutral lipids | Component of serine palmitoyltransferase, responsible along with Lcb2p for the first committed step in sphingolipid synthesis, which is the condensation of serine with palmitoyl-CoA to form 3-ketosphinganine | Putative protein of unknown function; non-tagged protein is detected in purified mitochondria; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Anchorage subunit of a-agglutinin of a-cells, highly O-glycosylated protein with N-terminal secretion signal and C-terminal signal for addition of GPI anchor to cell wall, linked to adhesion subunit Aga2p via two disulfide bonds | Protein involved in control of glucose-regulated gene expression; interacts with protein kinase Snf1p, glucose sensors Snf3p and Rgt2p, and TATA-binding protein Spt15p; acts as a regulator of the transcription factor Rgt1p | G protein beta subunit, forms a dimer with Ste18p to activate the mating signaling pathway, forms a heterotrimer with Gpa1p and Ste18p to dampen signaling; may recruit Rho1p to the polarized growth site during mating; contains WD40 repeats | Plasma membrane riboflavin transporter; facilitates the uptake of vitamin B2; required for FAD-dependent processes; sequence similarity to mammalian monocarboxylate permeases, however mutants are not deficient in monocarboxylate transport | Endoplasmic reticulum (ER)-resident membrane protein, overproduction induces enlargement of ER-like membrane structures and suppresses a temperature-sensitive sly1 mutation | Signal recognition particle (SRP) subunit (homolog of mammalian SRP54); contains the signal sequence-binding activity of SRP, interacts with the SRP RNA, and mediates binding of SRP to signal receptor; contains GTPase domain | Protein of unknown function; exhibits genetic interaction with ERG11 and protein-protein interaction with Hsp82p | Putative protein of unknown function |
| 144 | 53 | 441 | 0.594 | 0.570 | 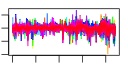 |  | 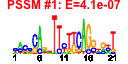 | 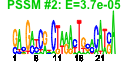 |  | YBR148W YDL139C YGL164C YGL175C YIR009W YKL033W YKL089W YKL132C YLL061W YLR004C YLR170C YMR198W YMR231W YMR294W YNL128W YNL278W YNR062C YNR064C YOL043C YOL063C YOL091W YOL164W YOR339C YOR378W YPL138C YPR086W YPR179C YBR150C YDL220C YDR076W YIL080W YIR023W YLL033W YLR070C YML005W YMR063W YMR106C YNL179C YNL279W YNR004W YNR008W YNR056C YOL017W YOL089C YOR012W YOR062C YOR114W YOR196C YOR197W YPL033C YPL194W YPR031W YPR085C | YSW1 SCM3 YRB30 SAE2 MSL1 YKL033W-A MAS2 RMA1 MMP1 THI73 APS1 CIK1 PEP5 JNM1 TEP1 CAF120 NTG2 CRT10 SPO21 YOL164W-A UBC11 SPP1 SUA7 HDA3 TBS1 CDC13 RAD55 DAL81 IRC19 XYL2 TRM12 RIM9 YKU80 PRM1 LRO1 BIO5 ESC8 HAL9 LIP5 MCA1 DDC1 NTO1 | Protein expressed specifically in spores | Nonhistone component of centromeric chromatin that binds stoichiometrically to CenH3-H4 histones, required for kinetochore assembly; contains nuclear export signal (NES); required for G2/M progression and localization of Cse4p | RanGTP-binding protein, inhibits RanGAP1 (Rna1p)-mediated GTP hydrolysis of RanGTP (Gsp1p); shares similarity to proteins in other fungi but not in higher eukaryotes | Protein with a role in accurate meiotic and mitotic double-strand break repair; phosphorylated in response to DNA damage and required for normal resistance to DNA-damaging agents | U2B component of U2 snRNP, involved in splicing, binds the U2 snRNA stem-loop IV in vitro; does not contain the conserved C-terminal RNA binding domain found in other family members | Putative protein of unknown function; similar to uncharacterized proteins from other fungi | Larger subunit of the mitochondrial processing protease (MPP), essential processing enzyme that cleaves the N-terminal targeting sequences from mitochondrially imported proteins | Putative dihydrofolate synthetase; has similarity to Fol3p and to E. coli folylpolyglutamate synthetase/dihydrofolate synthetase; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | High-affinity S-methylmethionine permease, required for utilization of S-methylmethionine as a sulfur source; has similarity to S-adenosylmethionine permease Sam3p | Putative plasma membrane permease proposed to be involved in carboxylic acid uptake and repressed by thiamine; substrate of Dbf2p/Mob1p kinase; transcription is altered if mitochondrial dysfunction occurs | Small subunit of the clathrin-associated adaptor complex AP-1, which is involved in protein sorting at the trans-Golgi network; homolog of the sigma subunit of the mammalian clathrin AP-1 complex | Kinesin-associated protein required for both karyogamy and mitotic spindle organization, interacts stably and specifically with Kar3p and may function to target this kinesin to a specific cellular role; has similarity to Vik1p | Component of CORVET tethering complex; peripheral vacuolar membrane protein required for protein trafficking and vacuole biogenesis; interacts with Pep7p | Component of the yeast dynactin complex, consisting of Nip100p, Jnm1p, and Arp1p; required for proper nuclear migration and spindle partitioning during mitotic anaphase B | Homolog of human tumor suppressor gene PTEN/MMAC1/TEP1 that has lipid phosphatase activity and is linked to the phosphatidylinositol signaling pathway; plays a role in normal sporulation | Part of the evolutionarily-conserved CCR4-NOT transcriptional regulatory complex involved in controlling mRNA initiation, elongation, and degradation | Putative membrane protein of unknown function | Epoxide hydrolase, member of the alpha/beta hydrolase fold family; may have a role in detoxification of epoxides | DNA N-glycosylase and apurinic/apyrimidinic (AP) lyase involved in base excision repair, localizes to the nucleus | Protein involved in transcriptional regulation of RNR2 and RNR3; expression of the gene is induced by DNA damage and null mutations confer increased resistance to hydroxyurea; N-terminal region has a leucine repeat and a WD40 repeat | Component of the meiotic outer plaque of the spindle pole body, involved in modifying the meiotic outer plaque that is required prior to prospore membrane formation | Putative protein of unknown function; identified by fungal homology and RT-PCR | Ubiquitin-conjugating enzyme most similar in sequence to Xenopus ubiquitin-conjugating enzyme E2-C, but not a true functional homolog of this E2; unlike E2-C, not required for the degradation of mitotic cyclin Clb2 | Putative protein of unknown function; belongs to the DHA2 family of drug:H+ antiporters; YOR378W is not an essential gene | Subunit of COMPASS (Set1C), a complex which methylates histone H3 on lysine 4 and is required in telomeric transcriptional silencing; PHD finger domain protein similar to human CGBP, an unmethylated CpG binding protein | Transcription factor TFIIB, a general transcription factor required for transcription initiation and start site selection by RNA polymerase II | Subunit of a possibly tetrameric trichostatin A-sensitive class II histone deacetylase complex that contains an Hda1p homodimer and an Hda2p-Hda3p heterodimer; required for the activity of the complex; has similarity to Hda2p | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Single stranded DNA-binding protein found at TG1-3 telomere G-tails; regulates telomere replication through recruitment of specific sub-complexes, but the essential function is telomere capping | Protein that stimulates strand exchange by stabilizing the binding of Rad51p to single-stranded DNA; involved in the recombinational repair of double-strand breaks in DNA during vegetative growth and meiosis; forms heterodimer with Rad57p | Positive regulator of genes in multiple nitrogen degradation pathways; contains DNA binding domain but does not appear to bind the dodecanucleotide sequence present in the promoter region of many genes involved in allantoin catabolism | Putative protein of unknown function; YLL033W is not an essential gene but mutant is defective in spore formation; null mutant displays increased levels of spontaneous Rad52 foci | Xylitol dehydrogenase, converts xylitol to D-xylulose in the endogenous xylose utilization pathway | S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; required for wybutosine formation in phenylalanine-accepting tRNA | Protein of unknown function, involved in the proteolytic activation of Rim101p in response to alkaline pH; has similarity to A. nidulans PalI; putative membrane protein | Subunit of the telomeric Ku complex (Yku70p-Yku80p), involved in telomere length maintenance, structure and telomere position effect; relocates to sites of double-strand cleavage to promote nonhomologous end joining during DSB repair | Pheromone-regulated multispanning membrane protein involved in membrane fusion during mating; predicted to have 5 transmembrane segments and a coiled coil domain; localizes to the shmoo tip; regulated by Ste12p | Putative protein of unknown function; haploid disruptant exhibits slow growth rate on glucose-minimal medium at 15 C | Acyltransferase that catalyzes diacylglycerol esterification; one of several acyltransferases that contribute to triglyceride synthesis; putative homolog of human lecithin cholesterol acyltransferase | Putative transmembrane protein involved in the biotin biosynthesis pathway; responsible for uptake of 7-keto 8-aminopelargonic acid; BIO5 is in a cluster of 3 genes (BIO3, BIO4, and BIO5) that mediate biotin synthesis | Protein involved in telomeric and mating-type locus silencing, interacts with Sir2p and also interacts with the Gal11p, which is a component of the RNA pol II mediator complex | Putative transcription factor containing a zinc finger; overexpression increases salt tolerance through increased expression of the ENA1 (Na+/Li+ extrusion pump) gene while gene disruption decreases both salt tolerance and ENA1 expression | Putative protein of unknown function | Protein of unknown function; similar to YKR075Cp and Reg1p; expression regulated by glucose and Rgt1p; GFP-fusion protein is induced in response to the DNA-damaging agent MMS | hypothetical protein | Protein involved in biosynthesis of the coenzyme lipoic acid, has similarity to E. coli lipoic acid synthase | Putative cysteine protease similar to mammalian caspases, involved in regulation of apoptosis upon hydrogen peroxide treatment | Putative protein of unknown function; may be involved in DNA metabolism; expression is induced by Kar4p | DNA damage checkpoint protein, part of a PCNA-like complex required for DNA damage response, required for pachytene checkpoint to inhibit cell cycle in response to unrepaired recombination intermediates; potential Cdc28p substrate | Subunit of the NuA3 histone acetyltransferase complex that acetylates histone H3; contains PHD finger domain that interacts with methylated histone H3 | Putative protein of unknown function; YPR085C is an essential gene |
| 184 | 26 | 447 | 0.626 | 0.545 | 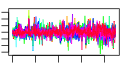 |  | 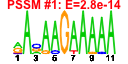 |  |  | YBL048W YBR050C YBR051W YCR005C YCR021C YDR256C YJL116C YKL109W YLR408C YOR223W YOR338W YPL119C YPL185W YAL055W YBR286W YDR313C YER065C YHR198C YIL111W YJL048C YJL066C YJL067W YLL020C YLR142W YOR186W YPR015C | REG2 CIT2 HSP30 CTA1 NCA3 HAP4 YPL119C-A PEX22 APE3 PIB1 ICL1 AIM18 COX5B UBX6 MPM1 PUT1 | Regulatory subunit of the Glc7p type-1 protein phosphatase; involved with Reg1p, Glc7p, and Snf1p in regulation of glucose-repressible genes, also involved in glucose-induced proteolysis of maltose permease | Citrate synthase, catalyzes the condensation of acetyl coenzyme A and oxaloacetate to form citrate, peroxisomal isozyme involved in glyoxylate cycle; expression is controlled by Rtg1p and Rtg2p transcription factors | Hydrophobic plasma membrane localized, stress-responsive protein that negatively regulates the H(+)-ATPase Pma1p; induced by heat shock, ethanol treatment, weak organic acid, glucose limitation, and entry into stationary phase | Catalase A, breaks down hydrogen peroxide in the peroxisomal matrix formed by acyl-CoA oxidase (Pox1p) during fatty acid beta-oxidation | Protein that functions with Nca2p to regulate mitochondrial expression of subunits 6 (Atp6p) and 8 (Atp8p ) of the Fo-F1 ATP synthase; member of the SUN family | Subunit of the heme-activated, glucose-repressed Hap2p/3p/4p/5p CCAAT-binding complex, a transcriptional activator and global regulator of respiratory gene expression; provides the principal activation function of the complex | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endosome; YLR408C is not an essential gene | Putative protein of unknown function | Putative protein of unknown function; YOR338W transcription is regulated by Azf1p and its transcript is a specific target of the G protein effector Scp160p; identified as being required for sporulation in a high-throughput mutant screen | Putative protein of unknown function; identified by expression profiling and mass spectrometry | Putative peroxisomal membrane protein required for import of peroxisomal proteins, functionally complements a Pichia pastoris pex22 mutation | Vacuolar aminopeptidase Y, processed to mature form by Prb1p | RING-type ubiquitin ligase of the endosomal and vacuolar membranes, binds phosphatidylinositol(3)-phosphate; contains a FYVE finger domain | Isocitrate lyase, catalyzes the formation of succinate and glyoxylate from isocitrate, a key reaction of the glyoxylate cycle; expression of ICL1 is induced by growth on ethanol and repressed by growth on glucose | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Subunit Vb of cytochrome c oxidase, which is the terminal member of the mitochondrial inner membrane electron transport chain; predominantly expressed during anaerobic growth while its isoform Va (Cox5Ap) is expressed during aerobic growth | UBX (ubiquitin regulatory X) domain-containing protein that interacts with Cdc48p, transcription is repressed when cells are grown in media containing inositol and choline | Mitochondrial membrane protein of unknown function, contains no hydrophobic stretches | Proline oxidase, nuclear-encoded mitochondrial protein involved in utilization of proline as sole nitrogen source; PUT1 transcription is induced by Put3p in the presence of proline and the absence of a preferred nitrogen source | Putative protein of unknown function; proper regulation of expression during heat stress is sphingolipid-dependent |
| 71 | 47 | 439 | 0.629 | 0.556 |  |  | 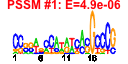 |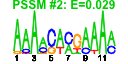  |  | YAL007C YBR017C YBR130C YDL052C YDL143W YDR002W YDR158W YDR353W YDR400W YEL071W YGL026C YGL039W YGR037C YHL034C YHR063C YHR163W YHR179W YHR183W YJL008C YJL014W YJL145W YJR040W YKR071C YLR192C YLR314C YML031W YML074C YML085C YML124C YMR208W YMR318C YNL121C YOL123W YOR198C YOR209C YOR232W YOR236W YOR323C YPL010W YBR025C YCR052W YDL040C YGL105W YGL207W YHR013C YNL064C YOL049W | ERP2 KAP104 SHE3 SLC1 PET9 YRB1 HOM2 TRR1 URH1 DLD3 TRP5 ACB1 SSB1 PAN5 SOL3 OYE2 GND1 CCT8 CCT3 SFH5 GEF1 DRE2 HCR1 CDC3 NDC1 FPR3 TUB1 TUB3 ERG12 ADH6 TOM70 HRP1 BFR1 NPT1 MGE1 DFR1 PRO2 RET3 OLA1 RSC6 NAT1 ARC1 SPT16 ARD1 YDJ1 GSH2 | Protein that forms a heterotrimeric complex with Erp1p, Emp24p, and Erv25p; member, along with Emp24p and Erv25p, of the p24 family involved in ER to Golgi transport and localized to COPII-coated vesicles | Transportin, cytosolic karyopherin beta 2 involved in delivery of heterogeneous nuclear ribonucleoproteins to the nucleoplasm, binds rg-nuclear localization signals on Nab2p and Hrp1p, plays a role in cell-cycle progression | Protein that acts as an adaptor between Myo4p and the She2p-mRNA complex; part of the mRNA localization machinery that restricts accumulation of certain proteins to the bud; also required for cortical ER inheritance | 1-acyl-sn-gylcerol-3-phosphate acyltransferase, catalyzes the acylation of lysophosphatidic acid to form phosphatidic acid, a key intermediate in lipid metabolism; located in lipid particles and endoplasmic reticulum | Major ADP/ATP carrier of the mitochondrial inner membrane, exchanges cytosolic ADP for mitochondrially synthesized ATP; phosphorylated; required for viability in many common lab strains carrying a mutation in the polymorphic SAL1 gene | Ran GTPase binding protein; involved in nuclear protein import and RNA export, ubiquitin-mediated protein degradation during the cell cycle; shuttles between the nucleus and cytoplasm; is essential; homolog of human RanBP1 | Aspartic beta semi-aldehyde dehydrogenase, catalyzes the second step in the common pathway for methionine and threonine biosynthesis; expression regulated by Gcn4p and the general control of amino acid synthesis | Cytoplasmic thioredoxin reductase, key regulatory enzyme that determines the redox state of the thioredoxin system, which acts as a disulfide reductase system and protects cells against both oxidative and reductive stress | Uridine nucleosidase (uridine-cytidine N-ribohydrolase), cleaves N-glycosidic bonds in nucleosides; involved in the pyrimidine salvage and nicotinamide riboside salvage pathways | D-lactate dehydrogenase, part of the retrograde regulon which consists of genes whose expression is stimulated by damage to mitochondria and reduced in cells grown with glutamate as the sole nitrogen source, located in the cytoplasm | Tryptophan synthase involved in tryptophan biosynthesis, regulated by the general control system of amino acid biosynthesis | Oxidoreductase, catalyzes NADPH-dependent reduction of the bicyclic diketone bicyclo[2.2.2]octane-2,6-dione (BCO2,6D) to the chiral ketoalcohol (1R,4S,6S)-6-hydroxybicyclo[2.2.2]octane-2-one (BCO2one6ol) | Acyl-CoA-binding protein, transports newly synthesized acyl-CoA esters from fatty acid synthetase (Fas1p-Fas2p) to acyl-CoA-consuming processes | Cytoplasmic ATPase that is a ribosome-associated molecular chaperone, functions with J-protein partner Zuo1p; may be involved in folding of newly-made polypeptide chains; member of the HSP70 family; interacts with phosphatase subunit Reg1p | 2-dehydropantoate 2-reductase, part of the pantothenic acid pathway, structurally homologous to E. coli panE | 6-phosphogluconolactonase, catalyzes the second step of the pentose phosphate pathway; weak multicopy suppressor of los1-1 mutation; homologous to Sol2p and Sol1p | Widely conserved NADPH oxidoreductase containing flavin mononucleotide (FMN), homologous to Oye3p with slight differences in ligand binding and catalytic properties; may be involved in sterol metabolism | 6-phosphogluconate dehydrogenase (decarboxylating), catalyzes an NADPH regenerating reaction in the pentose phosphate pathway; required for growth on D-glucono-delta-lactone and adaptation to oxidative stress | Subunit of the cytosolic chaperonin Cct ring complex, related to Tcp1p, required for the assembly of actin and tubulins in vivo | Putative phosphatidylinositol transfer protein (PITP), exhibits phosphatidylinositol- but not phosphatidylcholine-transfer activity, localized to cytosol and microsomes, similar to Sec14p; may be PITP regulator rather than actual PITP | Chloride channel localized to late- or post-Golgi vesicles, involved in iron metabolism; highly homologous to voltage-gated chloride channels in vertebrates | Protein of unknown function; mutation displays synthetic lethal interaction with the pol3-13 allele of CDC2 | Dual function protein involved in translation initiation as a substoichiometric component of eukaryotic translation initiation factor 3 (eIF3) and required for processing of 20S pre-rRNA; binds to eIF3 subunits Rpg1p and Prt1p and 18S rRNA | Component of the septin ring of the mother-bud neck that is required for cytokinesis; septins recruit proteins to the neck and can act as a barrier to diffusion at the membrane, and they comprise the 10nm filaments seen with EM | Nuclear envelope protein with multiple putative transmembrane domains, required for nuclear pore complex assembly and spindle pole body duplication; required for meiosis II | Nucleolar peptidyl-prolyl cis-trans isomerase (PPIase); FK506 binding protein; phosphorylated by casein kinase II (Cka1p-Cka2p-Ckb1p-Ckb2p) and dephosphorylated by Ptp1p | Alpha-tubulin; associates with beta-tubulin (Tub2p) to form tubulin dimer, which polymerizes to form microtubules | Alpha-tubulin; associates with beta-tubulin (Tub2p) to form tubulin dimer, which polymerizes to form microtubules; expressed at lower level than Tub1p | Mevalonate kinase, acts in the biosynthesis of isoprenoids and sterols, including ergosterol, from mevalonate | NADPH-dependent medium chain alcohol dehydrogenase with broad substrate specificity; member of the cinnamyl family of alcohol dehydrogenases; may be involved in fusel alcohol synthesis or in aldehyde tolerance | Component (70 kDa) of the TOM (translocase of outer membrane) complex responsible for recognition and initial import steps for all mitochondrially directed proteins; acts as a receptor for incoming precursor proteins | Subunit of cleavage factor I, a five-subunit complex required for the cleavage and polyadenylation of pre-mRNA 3' ends; RRM-containing heteronuclear RNA binding protein and hnRNPA/B family member that binds to poly (A) signal sequences | Component of mRNP complexes associated with polyribosomes; implicated in secretion and nuclear segregation; multicopy suppressor of BFA (Brefeldin A) sensitivity | Nicotinate phosphoribosyltransferase, acts in the salvage pathway of NAD+ biosynthesis; required for silencing at rDNA and telomeres and has a role in silencing at mating-type loci; localized to the nucleus | Protein of the mitochondrial matrix involved in protein import into mitochondria; acts as a cochaperone and a nucleotide release factor for Ssc1p; homolog of E. coli GrpE | Dihydrofolate reductase, part of the dTTP biosynthetic pathway, involved in folate metabolism, possibly required for mitochondrial function | Gamma-glutamyl phosphate reductase, catalyzes the second step in proline biosynthesis | Zeta subunit of the coatomer complex (COPI), which coats Golgi-derived transport vesicles; involved in retrograde transport between Golgi and ER | P-loop ATPase with similarity to human OLA1 and bacterial YchF; identified as specifically interacting with the proteasome; protein levels are induced by hydrogen peroxide | Component of the RSC chromatin remodeling complex; essential for mitotic growth; homolog of SWI/SNF subunit Swp73p | Subunit of the N-terminal acetyltransferase NatA (Nat1p, Ard1p, Nat5p); N-terminally acetylates many proteins, which influences multiple processes such as the cell cycle, heat-shock resistance, mating, sporulation, and telomeric silencing | Protein that binds tRNA and methionyl- and glutamyl-tRNA synthetases (Mes1p and Gus1p), delivering tRNA to them, stimulating catalysis, and ensuring their localization to the cytoplasm; also binds quadruplex nucleic acids | Subunit of the heterodimeric FACT complex (Spt16p-Pob3p), facilitates RNA Polymerase II transcription elongation through nucleosomes by destabilizing and then reassembling nucleosome structure | Protein chaperone involved in regulation of the HSP90 and HSP70 functions; involved in protein translocation across membranes; member of the DnaJ family | Glutathione synthetase, catalyzes the ATP-dependent synthesis of glutathione (GSH) from gamma-glutamylcysteine and glycine; induced by oxidative stress and heat shock |
| 279 | 43 | 438 | 0.640 | 0.576 |  |  |  |  |  | YJL012C YJR014W YJR031C YKL209C YMR220W YNL029C YPR169W YBR021W YBR060C YDR172W YDR300C YFL026W YGL106W YGR228W YHR005C YIL015W YJL130C YJR032W YLR088W YLR244C YLR274W YLR291C YLR314C YLR335W YLR420W YLR452C YML021C YML038C YMR211W YMR296C YNL035C YNL059C YNL109W YNL123W YNR044W YOL076W YOR026W YOR074C YOR143C YOR239W YOR390W YPL094C YPL279C | VTC4 TMA22 GEA1 STE6 ERG8 KTR5 JIP5 FUR4 ORC2 SUP35 PRO1 STE2 MLC1 GPA1 BAR1 URA2 CPR7 GAA1 MAP1 BOI1 GCD7 CDC3 NUP2 URA4 SST2 UNG1 YMD8 DML1 LCB1 ARP5 NMA111 AGA1 MDM20 BUB3 CDC21 THI80 ABP140 SEC62 | Vacuolar membrane protein involved in vacuolar polyphosphate accumulation; functions as a regulator of vacuolar H+-ATPase activity and vacuolar transporter chaperones; involved in non-autophagic vacuolar fusion | Protein of unknown function; associates with ribosomes and has a putative RNA binding domain; interacts with Tma20p; similar to human GRAP and human DRP1, which interacts with human Tma20p homolog MCT-1 | Guanine nucleotide exchange factor for ADP ribosylation factors (ARFs), involved in vesicular transport between the Golgi and ER, Golgi organization, and actin cytoskeleton organization; similar to but not functionally redundant with Gea2p | Plasma membrane ATP-binding cassette (ABC) transporter required for the export of a-factor, catalyzes ATP hydrolysis coupled to a-factor transport; contains 12 transmembrane domains and two ATP binding domains; expressed only in MATa cells | Phosphomevalonate kinase, an essential cytosolic enzyme that acts in the biosynthesis of isoprenoids and sterols, including ergosterol, from mevalonate | Putative mannosyltransferase involved in protein glycosylation; member of the KRE2/MNT1 mannosyltransferase family | Nucleolar protein of unknown function, exhibits a physical interaction with Bre1p | Uracil permease, localized to the plasma membrane; expression is tightly regulated by uracil levels and environmental cues | Subunit of the origin recognition complex, which directs DNA replication by binding to replication origins and is also involved in transcriptional silencing; phosphorylated by Cdc28p | Translation termination factor eRF3; altered protein conformation creates the [PSI(+)] prion, a dominant cytoplasmically inherited protein aggregate that alters translational fidelity and creates a nonsense suppressor phenotype | Gamma-glutamyl kinase, catalyzes the first step in proline biosynthesis | Receptor for alpha-factor pheromone; seven transmembrane-domain GPCR that interacts with both pheromone and a heterotrimeric G protein to initiate the signaling response that leads to mating between haploid a and alpha cells | Essential light chain for Myo1p, light chain for Myo2p; stabilizes Myo2p by binding to the neck region; interacts with Myo1p, Iqg1p, and Myo2p to coordinate formation and contraction of the actomyosin ring with targeted membrane deposition | GTP-binding alpha subunit of the heterotrimeric G protein that couples to pheromone receptors; negatively regulates the mating pathway by sequestering G(beta)gamma and by triggering an adaptive response; activates Vps34p at the endosome | Aspartyl protease secreted into the periplasmic space of mating type a cells, helps cells find mating partners, cleaves and inactivates alpha factor allowing cells to recover from alpha-factor-induced cell cycle arrest | Bifunctional carbamoylphosphate synthetase (CPSase)-aspartate transcarbamylase (ATCase), catalyzes the first two enzymatic steps in the de novo biosynthesis of pyrimidines; both activities are subject to feedback inhibition by UTP | Peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; binds to Hsp82p and contributes to chaperone activity | Subunit of the GPI (glycosylphosphatidylinositol):protein transamidase complex, removes the GPI-anchoring signal and attaches GPI to proteins in the ER | Methionine aminopeptidase, catalyzes the cotranslational removal of N-terminal methionine from nascent polypeptides; function is partially redundant with that of Map2p | Protein implicated in polar growth, functionally redundant with Boi2p; interacts with bud-emergence protein Bem1p; contains an SH3 (src homology 3) domain and a PH (pleckstrin homology) domain | Beta subunit of the translation initiation factor eIF2B, the guanine-nucleotide exchange factor for eIF2; activity subsequently regulated by phosphorylated eIF2; first identified as a negative regulator of GCN4 expression | Component of the septin ring of the mother-bud neck that is required for cytokinesis; septins recruit proteins to the neck and can act as a barrier to diffusion at the membrane, and they comprise the 10nm filaments seen with EM | Nucleoporin involved in nucleocytoplasmic transport, binds to either the nucleoplasmic or cytoplasmic faces of the nuclear pore complex depending on Ran-GTP levels; also has a role in chromatin organization | Dihydroorotase, catalyzes the third enzymatic step in the de novo biosynthesis of pyrimidines, converting carbamoyl-L-aspartate into dihydroorotate | GTPase-activating protein for Gpa1p, regulates desensitization to alpha factor pheromone; also required to prevent receptor-independent signaling of the mating pathway; member of the RGS (regulator of G-protein signaling) family | Uracil-DNA glycosylase, required for repair of uracil in DNA formed by spontaneous cytosine deamination, not required for strand-specific mismatch repair, cell-cycle regulated, expressed in late G1, localizes to mitochondria and nucleus | Putative nucleotide sugar transporter, has similarity to Vrg4p | Essential protein involved in mtDNA inheritance, may also function in the partitioning of the mitochondrial organelle or in the segregation of chromosomes, exhibits regions similar to members of a GTPase family | Component of serine palmitoyltransferase, responsible along with Lcb2p for the first committed step in sphingolipid synthesis, which is the condensation of serine with palmitoyl-CoA to form 3-ketosphinganine | Putative protein of unknown function with similarity to proteins containing WD-40 domains; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; YNL035C is not an essential gene | Nuclear actin-related protein involved in chromatin remodeling, component of chromatin-remodeling enzyme complexes | Protein of unknown function which may contribute to lipid homeostasis and/or apoptosis; sequence similarity to the mammalian Omi/HtrA2 family of serine proteases | Anchorage subunit of a-agglutinin of a-cells, highly O-glycosylated protein with N-terminal secretion signal and C-terminal signal for addition of GPI anchor to cell wall, linked to adhesion subunit Aga2p via two disulfide bonds | Non-catalytic subunit of the NatB N-terminal acetyltransferase, which catalyzes N-acetylation of proteins with specific N-terminal sequences; involved in mitochondrial inheritance and actin assembly | Kinetochore checkpoint WD40 repeat protein that localizes to kinetochores during prophase and metaphase, delays anaphase in the presence of unattached kinetochores; forms complexes with Mad1p-Bub1p and with Cdc20p, binds Mad2p and Mad3p | Thymidylate synthase, required for de novo biosynthesis of pyrimidine deoxyribonucleotides; expression is induced at G1/S | Thiamine pyrophosphokinase, phosphorylates thiamine to produce the coenzyme thiamine pyrophosphate (thiamine diphosphate) | Nonessential protein that binds actin filaments and localizes to actin patches and cables, has similarity to S-adenosylmethionine (AdoMet)-dependent methyltransferases | Putative protein of unknown function | Essential subunit of Sec63 complex (Sec63p, Sec62p, Sec66p and Sec72p); with Sec61 complex, Kar2p/BiP and Lhs1p forms a channel competent for SRP-dependent and post-translational SRP-independent protein targeting and import into the ER |
| 154 | 32 | 446 | 0.661 | 0.588 |  |  |  | 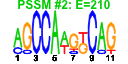 |  | YAL008W YBL001C YBR183W YDL223C YEL073C YGR182C YHR054C YHR113W YHR150W YJL048C YJL067W YJL153C YBR054W YCR079W YDL018C YDL046W YDL142C YDR230W YDR287W YDR406W YDR504C YDR530C YER037W YGL010W YGL046W YGL193C YGR154C YHR053C YHR055C YIL050W YIL098C YKL222C | FUN14 ECM15 YPC1 HBT1 PEX28 UBX6 INO1 YRO2 PTC6 ERP3 NPC2 CRD1 INM2 PDR15 SPG3 APA2 PHM8 RIM8 GTO1 CUP1-1 PCL7 FMC1 | Mitochondrial protein of unknown function | Non-essential protein of unknown function, likely exists as tetramer, may be regulated by the binding of small-molecule ligands (possibly sulfate ions), may have a role in yeast cell-wall biogenesis | Alkaline ceramidase that also has reverse (CoA-independent) ceramide synthase activity, catalyzes both breakdown and synthesis of phytoceramide; overexpression confers fumonisin B1 resistance | Substrate of the Hub1p ubiquitin-like protein that localizes to the shmoo tip (mating projection); mutants are defective for mating projection formation, thereby implicating Hbt1p in polarized cell morphogenesis | Putative protein of unknown function; located adjacent to ARS503 and the telomere on the left arm of chromosome V; regulated by inositol/choline | Putative protein of unknown function | Cytoplasmic aspartyl aminopeptidase; cleaves unblocked N-terminal acidic amino acid residues from peptide substrates; forms a 12 subunit homo-oligomeric complex; M18 metalloprotease family member; may interact with ribosomes | Peroxisomal integral membrane peroxin, involved in the regulation of peroxisomal size, number and distribution; genetic interactions suggest that Pex28p and Pex29p act at steps upstream of those mediated by Pex30p, Pex31p, and Pex32p | UBX (ubiquitin regulatory X) domain-containing protein that interacts with Cdc48p, transcription is repressed when cells are grown in media containing inositol and choline | Inositol 1-phosphate synthase, involved in synthesis of inositol phosphates and inositol-containing phospholipids; transcription is coregulated with other phospholipid biosynthetic genes by Ino2p and Ino4p, which bind the UASINO DNA element | Putative protein of unknown function; the authentic, non-tagged protein is detected in a phosphorylated state in highly purified mitochondria in high-throughput studies; transcriptionally regulated by Haa1p | Phosphoprotein phosphatase type 2C similar to mammalian PP1Ks; involved in mitophagy; localized to mitochondrial inner membrane space; null mutant is sensitive to rapamycin | Protein with similarity to Emp24p and Erv25p, member of the p24 family involved in ER to Golgi transport | Functional homolog of human NPC2/He1, which is a cholesterol-binding protein whose deficiency causes Niemann-Pick type C2 disease involving retention of cholesterol in lysosomes | Cardiolipin synthase; produces cardiolipin, which is an important constituent of mitochondrial membranes; required for normal mitochondrial membrane potential and function | Inositol monophosphatase, involved in biosynthesis of inositol; enzymatic activity requires magnesium ions and is inhibited by lithium and sodium ions; inm1 inm2 double mutant lacks inositol auxotrophy | Plasma membrane ATP binding cassette (ABC) transporter, multidrug transporter and general stress response factor implicated in cellular detoxification; regulated by Pdr1p, Pdr3p and Pdr8p; promoter contains a PDR responsive element | Protein required for survival at high temperature during stationary phase; not required for growth on nonfermentable carbon sources | Diadenosine 5',5''-P1,P4-tetraphosphate phosphorylase II (AP4A phosphorylase), involved in catabolism of bis(5'-nucleosidyl) tetraphosphates; has similarity to Apa1p | Protein of unknown function, expression is induced by low phosphate levels and by inactivation of Pho85p | Putative protein of unknown function; YGL010W is not an essential gene | Protein of unknown function, involved in the proteolytic activation of Rim101p in response to alkaline pH; has similarity to A. nidulans PalF | Haploid-specific gene repressed by a1-alpha2, turned off in sir3 null strains, absence enhances the sensitivity of rad52-327 cells to campothecin almost 100-fold | Omega-class glutathione transferase; induced under oxidative stress; putative peroxisomal localization | Metallothionein, binds copper and mediates resistance to high concentrations of copper and cadmium; locus is variably amplified in different strains, with two copies, CUP1-1 and CUP1-2, in the genomic sequence reference strain S288C | Pho85p cyclin of the Pho80p subfamily, forms a functional kinase complex with Pho85p which phosphorylates Mmr1p and is regulated by Pho81p; involved in glycogen metabolism, expression is cell-cycle regulated | Mitochondrial matrix protein, required for assembly or stability at high temperature of the F1 sector of mitochondrial F1F0 ATP synthase; null mutant temperature sensitive growth on glycerol is suppressed by multicopy expression of Odc1p | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; similar to transcriptional regulators from the zinc cluster (binuclear cluster) family; null mutant is sensitive to caffeine |
| 166 | 34 | 445 | 0.670 | 0.619 |  |  | 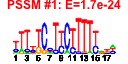 | 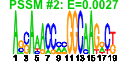 |  | YDL129W YDL149W YDR169C YEL019C YGL160W YGL179C YGR010W YGR222W YIL010W YIL137C YKL161C YKL204W YKR053C YKR061W YLR120C YMR177W YNR006W YNR010W YNR059W YOR125C YOR137C YAR023C YAR031W YBR160W YDL089W YDL206W YFR047C YGR133W YJL083W YKL159C YLR118C YML102W YMR295C YPL264C | ATG9 STB3 MMS21 AIM14 TOS3 NMA2 PET54 DOT5 TMA108 EAP1 YSR3 KTR2 YAP3 MMT1 VPS27 CSE2 MNT4 CAT5 SIA1 PRM9 CDC28 BNA6 PEX4 TAX4 RCN1 CAC2 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and the nucleus; YDL129W is not an essential gene | Transmembrane protein involved in formation of Cvt and autophagic vesicles; cycles between the pre-autophagosomal structure and other cytosolic punctate structures, not found in autophagosomes | Protein that binds Sin3p in a two-hybrid assay | SUMO ligase involved in chromosomal organization and DNA repair; essential subunit of the Mms21-Smc5-Smc6 complex; mutants are sensitive to methyl methanesulfonate and show increased spontaneous mutation and mitotic recombination | Putative protein of unknown function; similar to iron/copper reductases (FRE1-8), possibly involved in iron homeostasis; may interact with ribosomes, null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Protein kinase, related to and functionally redundant with Elm1p and Sak1p for the phosphorylation and activation of Snf1p; functionally orthologous to LKB1, a mammalian kinase associated with Peutz-Jeghers cancer-susceptibility syndrome | Nicotinic acid mononucleotide adenylyltransferase, involved in NAD(+) salvage pathway | Mitochondrial translational activator specific for the COX3 mRNA, acts together with Pet122p and Pet494p; also required for splicing of the COX1 intron AI5 beta; located in the mitochondrial inner membrane | Nuclear thiol peroxidase which functions as an alkyl-hydroperoxide reductase during post-diauxic growth | Protein that associates with ribosomes; putative metalloprotease | Protein kinase implicated in the Slt2p mitogen-activated (MAP) kinase signaling pathway; associates with Rlm1p | eIF4E-associated protein, binds eIF4E and inhibits cap-dependent translation, also functions independently of eIF4E to maintain genetic stability; plays a role in cell growth, implicated in the TOR signaling cascade | Dihydrosphingosine 1-phosphate phosphatase, membrane protein involved in sphingolipid metabolism; has similarity to Lcb3p | Mannosyltransferase involved in N-linked protein glycosylation; member of the KRE2/MNT1 mannosyltransferase family | Basic leucine zipper (bZIP) transcription factor | Putative metal transporter involved in mitochondrial iron accumulation; closely related to Mmt2p | Endosomal protein that forms a complex with Hse1p; required for recycling Golgi proteins, forming lumenal membranes and sorting ubiquitinated proteins destined for degradation; has Ubiquitin Interaction Motifs which bind ubiquitin (Ubi4p) | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; component of the Med9/10 module; required for regulation of RNA polymerase II activity | Putative alpha-1,3-mannosyltransferase, not required for protein O-glycosylation | Protein required for ubiquinone (Coenzyme Q) biosynthesis; localizes to the matrix face of the mitochondrial inner membrane in a large complex with ubiquinone biosynthetic enzymes; required for gluconeogenic gene activation | Protein of unassigned function involved in activation of the Pma1p plasma membrane H+-ATPase by glucose | Putative integral membrane protein, member of DUP240 gene family | Pheromone-regulated protein with 3 predicted transmembrane segments and an FF sequence, a motif involved in COPII binding; member of DUP240 gene family | Catalytic subunit of the main cell cycle cyclin-dependent kinase (CDK); alternately associates with G1 cyclins (CLNs) and G2/M cyclins (CLBs) which direct the CDK to specific substrates | Protein of unknown function; interacts with meiotic division protein Csm1p; green fluorescent protein (GFP)-fusion protein localizes to the nuclear periphery, potential Cdc28p substrate | Putative protein of unknown function; YDL206W is not an essential protein | Quinolinate phosphoribosyl transferase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Peroxisomal ubiquitin conjugating enzyme required for peroxisomal matrix protein import and peroxisome biogenesis | Protein involved in regulation of phosphatidylinositol 4,5-bisphosphate concentrations; Irs4p and Tax4p bind and activate the phosphatase Inp51p | Protein involved in calcineurin regulation during calcium signaling; has similarity to H. sapiens DSCR1 which is found in the Down Syndrome candidate region | Acyl-protein thioesterase responsible for depalmitoylation of Gpa1p; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and nucleus and is induced in response to the DNA-damaging agent MMS | Component of the chromatin assembly complex (with Rlf2p and Msi1p) that assembles newly synthesized histones onto recently replicated DNA, required for building functional kinetochores, conserved from yeast to humans | Protein of unknown function that associates with ribosomes; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery and bud; YMR295C is not an essential gene | Putative membrane protein of unknown function; physically interacts with Hsp82p; YPL264C is not an essential gene |
| 247 | 50 | 447 | 0.676 | 0.610 |  |  | 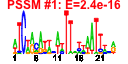 | 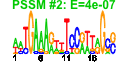 |  | YAR014C YBR075W YDL171C YDR300C YEL053C YHR072W YHR218W YLR138W YLR463C YML006C YMR127C YNL023C YNL123W YOL130W YOR116C YOR390W YPL036W YPL101W YPL108W YPL279C YBR069C YBR202W YCR035C YHL012W YHL049C YHR032W YHR204W YJL078C YJR043C YJR132W YLR322W YLR328W YLR353W YLR398C YLR419W YLR434C YLR462W YLR465C YLR466W YML123C YMR233W YNL108C YNL329C YNR065C YOR302W YPL019C YPL144W YPR016C YPR048W YPR203W | BUD14 YBR074W GLT1 PRO1 MAK10 ERG7 NHA1 GIS4 SAS2 FAP1 NMA111 SWC3 RPO31 PMA2 HAP1 TAT1 MCM7 RRP43 MNL1 PRY3 POL32 NMD5 NMA1 BUD8 SKI2 YRF1-1 PHO84 TRI1 PEX6 VTC3 POC4 TIF6 TAH18 | Protein involved in bud-site selection, Bud14p-Glc7p complex functions as a cortical regulator of dynein; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Putative metalloprotease | NAD(+)-dependent glutamate synthase (GOGAT), synthesizes glutamate from glutamine and alpha-ketoglutarate; with Gln1p, forms the secondary pathway for glutamate biosynthesis from ammonia; expression regulated by nitrogen source | Gamma-glutamyl kinase, catalyzes the first step in proline biosynthesis | Non-catalytic subunit of N-terminal acetyltransferase of the NatC type, required for replication of dsRNA virus; expression is glucose-repressible | Lanosterol synthase, an essential enzyme that catalyzes the cyclization of squalene 2,3-epoxide, a step in ergosterol biosynthesis | Helicase-like protein encoded within the telomeric Y' element | Na+/H+ antiporter involved in sodium and potassium efflux through the plasma membrane; required for alkali cation tolerance at acidic pH | CAAX box containing protein of unknown function, proposed to be involved in the RAS/cAMP signaling pathway | Histone acetyltransferase (HAT) catalytic subunit of the SAS complex (Sas2p-Sas4p-Sas5p), which acetylates free histones and nucleosomes and regulates transcriptional silencing; member of the MYSTacetyltransferase family | Protein that binds to Fpr1p (FKBP12), conferring rapamycin resistance by competing with rapamycin for Fpr1p binding; has similarity to putative transcription factors, including D. melanogaster shuttle craft and human NFX1 | Protein of unknown function which may contribute to lipid homeostasis and/or apoptosis; sequence similarity to the mammalian Omi/HtrA2 family of serine proteases | Protein of unknown function, component of the SWR1 complex, which exchanges histone variant H2AZ (Htz1p) for chromatin-bound histone H2A; required for formation of nuclear-associated array of smooth endoplasmic reticulum known as karmellae | RNA polymerase III subunit C160, part of core enzyme; similar to bacterial beta-prime subunit | Putative protein of unknown function | Plasma membrane H+-ATPase, isoform of Pma1p, involved in pumping protons out of the cell; regulator of cytoplasmic pH and plasma membrane potential | Zinc finger transcription factor involved in the complex regulation of gene expression in response to levels of heme and oxygen; the S288C sequence differs from other strain backgrounds due to a Ty1 insertion in the carboxy terminus | Cytoplasmic protein of unknown function; non-essential gene that is induced in a GDH1 deleted strain with altered redox metabolism; GFP-fusion protein is induced in response to the DNA-damaging agent MMS | Amino acid transport protein for valine, leucine, isoleucine, and tyrosine, low-affinity tryptophan and histidine transporter; overexpression confers FK506 and FTY720 resistance | Component of the hexameric MCM complex, which is important for priming origins of DNA replication in G1 and becomes an active ATP-dependent helicase that promotes DNA melting and elongation when activated by Cdc7p-Dbf4p in S-phase | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex with Rrp41p, Rrp42p, Rrp4p and Dis3p; required for efficient maturation of 5.8S, 18S and 25S rRNA | Putative protein of unknown function, has some homology to Ugp1p, which encodes UDP-glucose pyrophosphorylase | Putative protein of unknown function; putative substrate of the cAMP-dependent protein kinase (PKA) | Alpha mannosidase-like protein of the endoplasmic reticulum required for degradation of glycoproteins but not for processing of N-linked oligosaccharides | Protein of unknown function, has similarity to Pry1p and Pry2p and to the plant PR-1 class of pathogen related proteins | Third subunit of DNA polymerase delta, involved in chromosomal DNA replication; required for error-prone DNA synthesis in the presence of DNA damage and processivity; interacts with Hys2p, PCNA (Pol30p), and Pol1p | Karyopherin, a carrier protein involved in nuclear import of proteins; importin beta homolog | Nicotinic acid mononucleotide adenylyltransferase, involved in NAD(+) salvage pathway | Protein involved in bud-site selection; diploid mutants display a unipolar budding pattern instead of the wild-type bipolar pattern, and bud at the proximal pole | Putative RNA helicase, involved in exosome mediated 3' to 5' mRNA degradation and translation inhibition of non-poly(A) mRNAs; forms complex with Ski3p and Ski8p; required for repressing propagation of dsRNA viruses | Putative helicase with limited sequence similarity to human Rb protein; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YLR419W is not an essential gene | Putative protein of unknown function with similarity to helicases; YLR462W is within the telomere on the right arm of chromosome XII | Helicase encoded by the Y' element of subtelomeric regions, highly expressed in the mutants lacking the telomerase component TLC1; potentially phosphorylated by Cdc28p | High-affinity inorganic phosphate (Pi) transporter and low-affinity manganese transporter; regulated by Pho4p and Spt7p; mutation confers resistance to arsenate; exit from the ER during maturation requires Pho86p | Non-essential sumoylated protein of unknown function with similarity to components of human SWI/SNF complex including SMRD3; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm, nucleus and nucleolus | Putative protein of unknown function with similarity to Tfc7p and prokaryotic phosphotransfer enzymes; null mutant shows alterations in glucose metabolism; GFP-fusion protein localizes to the cytoplasm and nucleus | AAA-peroxin that heterodimerizes with AAA-peroxin Pex1p and participates in the recycling of peroxisomal signal receptor Pex5p from the peroxisomal membrane to the cystosol | Protein of unknown function; YNR065C is not an essential gene | CPA1 uORF , Arginine attenuator peptide, regulates translation of the CPA1 mRNA | Vacuolar membrane protein involved in vacuolar polyphosphate accumulation; functions as a regulator of vacuolar H+-ATPase activity and vacuolar transporter chaperones; involved in non-autophagic vacuolar fusion | Chaperone component involved in 20S proteasome assembly; forms a heterodimer with Irc25p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; null mutant is viable, exhibits shortened telomeres | Constituent of 66S pre-ribosomal particles, has similarity to human translation initiation factor 6 (eIF6); may be involved in the biogenesis and or stability of 60S ribosomal subunits | Protein with a potential role in DNA replication; displays synthetic lethal genetic interaction with the pol3-13 allele of POL3, which encodes DNA polymerase delta |
| 194 | 42 | 449 | 0.719 | 0.592 |  |  | 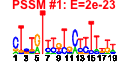 | 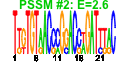 |  | YDL105W YDR253C YDR521W YGL089C YHR035W YJL084C YKR029C YLR457C YNL311C YNR063W YOL142W YOR076C YOR147W YPL113C YPR092W YPR111W YPR196W YBL097W YBR293W YDR088C YDR351W YGR224W YIR005W YIR009W YJL110C YKL105C YKR005C YKR069W YLR425W YMR025W YMR111C YMR138W YMR204C YMR255W YNR031C YOL102C YOR077W YOR086C YOR365C YOR371C YPL112C YPL261C | NSE4 MET32 MF(ALPHA)2 ALY2 SET3 NBP1 SKP2 RRP40 SKI7 MDM32 DBF20 BRN1 VBA2 SLU7 SBE2 AZR1 IST3 MSL1 GZF3 MET1 TUS1 CSI1 CIN4 INP1 GFD1 SSK2 TPT1 RTS2 TCB1 GPB1 PEX25 | Nuclear protein that plays a role in the function of the Smc5p-Rhc18p complex | Zinc-finger DNA-binding protein, involved in transcriptional regulation of the methionine biosynthetic genes, similar to Met31p | Mating pheromone alpha-factor, made by alpha cells; interacts with mating type a cells to induce cell cycle arrest and other responses leading to mating; also encoded by MF(ALPHA)1, which is more highly expressed than MF(ALPHA)2 | Putative protein of unknown function; not an essential gene | Cytoplasmic protein of unknown function that interacts with the cyclin Pcl7p; phosphorylated in vitro by the cyclin-CDK complex, Pcl7p-Pho85p; identified as a potential Cdc28p substrate; mRNA is cell cycle regulated, peaking in M phase | Defining member of the SET3 histone deacetylase complex which is a meiosis-specific repressor of sporulation genes; necessary for efficient transcription by RNAPII; one of two yeast proteins that contains both SET and PHD domains | Spindle pole body (SPB) component, required for the insertion of the duplication plaque into the nuclear membrane during SPB duplication; essential for bipolar spindle formation; component of the Mps2p-Bbp1p complex | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; putative F-box protein; analysis of integrated high-throughput datasets predicts involvement in ubiquitin-dependent protein catabolism | Putative zinc-cluster protein of unknown function | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex | Putative GTPase that mediates interactions via its N-terminus between the exosome and Ski complex (Ski2p, Ski3p, Ski8p) which operate in the 3'-to-5' mRNA-decay pathway; cytoplasmic protein required for degrading nonstop mRNAs | Mitochondrial inner membrane protein with similarity to Mdm31p, required for normal mitochondrial morphology and inheritance; interacts genetically with MMM1, MDM10, MDM12, and MDM34 | Glyoxylate reductase; acts on glyoxylate and hydroxypyruvate substrates; YPL113C is not an essential gene | Ser/Thr kinase involved in late nuclear division, one of the mitotic exit network (MEN) proteins; necessary for the execution of cytokinesis | Putative maltose activator | Essential protein required for chromosome condensation, likely to function as an intrinsic component of the condensation machinery, may influence multiple aspects of chromosome transmission and dynamics | Permease of basic amino acids in the vacuolar membrane | RNA splicing factor, required for ATP-independent portion of 2nd catalytic step of spliceosomal RNA splicing; interacts with Prp18p; contains zinc knuckle domain | Protein involved in the transport of cell wall components from the Golgi to the cell surface; required for bud growth | Plasma membrane transporter of the major facilitator superfamily, involved in resistance to azole drugs such as ketoconazole and fluconazole | Component of the U2 snRNP, required for the first catalytic step of splicing and for spliceosomal assembly; interacts with Rds3p and is required for Mer1p-activated splicing | U2B component of U2 snRNP, involved in splicing, binds the U2 snRNA stem-loop IV in vitro; does not contain the conserved C-terminal RNA binding domain found in other family members | GATA zinc finger protein and Dal80p homolog that negatively regulates nitrogen catabolic gene expression by competing with Gat1p for GATA site binding; function requires a repressive carbon source; dimerizes with Dal80p and binds to Tor1p | Putative protein of unknown function | S-adenosyl-L-methionine uroporphyrinogen III transmethylase, involved in the biosynthesis of siroheme, a prosthetic group used by sulfite reductase; required for sulfate assimilation and methionine biosynthesis | Guanine nucleotide exchange factor (GEF) that functions to modulate Rho1p activity as part of the cell integrity signaling pathway; multicopy suppressor of tor2 mutation and ypk1 ypk2 double mutation; potential Cdc28p substrate | Subunit of the Cop9 signalosome, which is required for deneddylation, or removal of the ubiquitin-like protein Rub1p from Cdc53p (cullin); involved in adaptation to pheromone signaling | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; YMR111C is not an essential gene | GTP-binding protein involved in beta-tubulin (Tub2p) folding; isolated as mutant with increased chromosome loss and sensitivity to benomyl | Peripheral membrane protein of peroxisomes involved in peroxisomal inheritance | Coiled-coiled protein of unknown function, identified as a high-copy suppressor of a dbp5 mutation | MAP kinase kinase kinase of the HOG1 mitogen-activated signaling pathway; interacts with Ssk1p, leading to autophosphorylation and activation of Ssk2p which phosphorylates Pbs2p; also mediates actin cytoskeleton recovery from osmotic stress | tRNA 2'-phosphotransferase, catalyzes the final step in yeast tRNA splicing: the transfer of the 2'-PO(4) from the splice junction to NAD(+) to form ADP-ribose 1''-2''cyclic phosphate and nicotinamide | Basic zinc-finger protein, similar to human and mouse Kin17 proteins which are chromatin-associated proteins involved in UV response and DNA replication | Lipid-binding protein containing three calcium and lipid binding domains; non-tagged protein localizes to mitochondria and GFP-fusion protein localizes to the cell periphery; C-termini of Tcb1p, Tcb2p and Tcb3p interact | Putative protein of unknown function; YOR365C is not an essential protein | Multistep regulator of cAMP-PKA signaling; inhibits PKA downstream of Gpa2p and Cyr1p, thereby increasing cAMP dependency; inhibits Ras activity through direct interactions with Ira1p/2p; regulated by G-alpha protein Gpa2p; homolog of Gpb2p | Peripheral peroxisomal membrane peroxin required for the regulation of peroxisome size and maintenance, recruits GTPase Rho1p to peroxisomes, induced by oleate, interacts with homologous protein Pex27p |
| 128 | 34 | 451 | 0.727 | 0.603 |  |  | 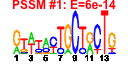 | 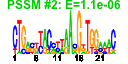 |  | YDL215C YDR535C YER188W YFR022W YHR080C YIL001W YIL028W YIL086C YIL165C YJR038C YKL147C YKL221W YLR094C YML005W YMR018W YOR049C YOR055W YOR389W YPL275W YPL277C YAL017W YCLX04W YCR068W YGR022C YHL006C YIL006W YIL007C YJR020W YKR098C YLR279W YLR280C YNR071C YOR348C YPR013C | GDH2 ROG3 MCH2 GIS3 TRM12 RSB1 PSK1 ATG15 SHU1 YIA6 NAS2 UBP11 PUT4 | NAD(+)-dependent glutamate dehydrogenase, degrades glutamate to ammonia and alpha-ketoglutarate; expression sensitive to nitrogen catabolite repression and intracellular ammonia levels | Protein that binds to Rsp5p, which is a hect-type ubiquitin ligase, via its 2 PY motifs; has similarity to Rod1p; mutation suppresses the temperature sensitivity of an mck1 rim11 double mutant | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | hypothetical protein | Putative protein of unknown function; in closely related species and other S. cerevisiae strain backgrounds YIL165C and adjacent ORF, YIL164C, likely constitute a single ORF encoding a nitrilase gene | Protein with similarity to mammalian monocarboxylate permeases, which are involved in transport of monocarboxylic acids across the plasma membrane; mutant is not deficient in monocarboxylate transport | Protein of unknown function | S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; required for wybutosine formation in phenylalanine-accepting tRNA | Putative protein of unknown function with similarity to human PEX5Rp (peroxin protein 5 related protein); transcription increases during colony development similar to genes involved in peroxisome biogenesis; YMR018W is not an essential gene | Suppressor of sphingoid long chain base (LCB) sensitivity of an LCB-lyase mutation; putative integral membrane transporter or flippase that may transport LCBs from the cytoplasmic side toward the extracytoplasmic side of the membrane | Putative protein of unknown function; expression regulated by copper levels | Putative protein of unknown function; localized to the membranes; gene expression regulated by copper levels | One of two (see also PSK2) PAS domain containing S/T protein kinases; coordinately regulates protein synthesis and carbohydrate metabolism and storage in response to a unknown metabolite that reflects nutritional status | Lipase, required for intravacuolar lysis of autophagic bodies; located in the endoplasmic reticulum membrane and targeted to intravacuolar vesicles during autophagy via the multivesicular body (MVB) pathway | Protein of unassigned function involved in mutation suppression, important for error-free repair of spontaneous and induced DNA lesions to protect the genome from mutation; associates with Shu2p, Psy3p, and Csm2p | Mitochondrial NAD+ transporter, involved in the transport of NAD+ into the mitochondria (see also YEA6); member of the mitochondrial carrier subfamily; disputed role as a pyruvate transporter; has putative mouse and human orthologs | Protein with similarity to the p27 subunit of mammalian proteasome modulator; not essential; interacts with Rpn4p | Ubiquitin-specific protease that cleaves ubiquitin from ubiquitinated proteins | Putative protein of unknown function | Proline permease, required for high-affinity transport of proline; also transports the toxic proline analog azetidine-2-carboxylate (AzC); PUT4 transcription is repressed in ammonia-grown cells | Putative zinc finger protein; YPR013C is not an essential gene |
| 246 | 45 | 447 | 0.727 | 0.563 |  |  | 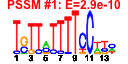 | 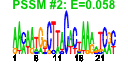 |  | YCL039W YDR488C YHR143BW YJL037W YJL039C YOR010C YAL066W YAR020C YAR070C YBL031W YBR213W YBR224W YCL001W YCR022C YCR085W YCR089W YCRX03C YDL210W YDR480W YER011W YGL034C YGR293C YHL005C YHR048W YIL011W YJL027C YJL150W YJL207C YJR150C YJR154W YKL131W YKR033C YLL015W YLR013W YLR415C YLR416C YMR057C YNL018C YOR009W YOR192C YOR314W YPL102C YPL136W YPL189W YPR096C | GID7 PAC11 IRC18 NUP192 TIR2 PAU7 MYO4 MET8 RER1 FIG2 UGA4 DIG2 TIR1 YHK8 TIR3 LAA1 DAN1 BPT1 GAT3 TIR4 THI72 GUP2 | Protein of unknown function, involved in proteasome-dependent catabolite inactivation of fructose-1,6-bisphosphatase; contains six WD40 repeats; computational analysis suggests that Gid7p and Moh1p have similar functions | Dynein intermediate chain, acts in the cytoplasmic dynein pathway, forms cortical cytoplasmic microtubule capture site with Num1p; null mutant is defective in nuclear migration, essential in the absence of CIN8 | Putative protein of unknown function; expression induced in respiratory-deficient cells and in carbon-limited chemostat cultures; similar to adjacent ORF, YJL038C; null mutant displays increased levels of spontaneous Rad52 foci | Essential structural subunit of the nuclear pore complex (NPC), localizes to the nuclear periphery of nuclear pores, homologous to human p205 | Putative cell wall mannoprotein of the Srp1p/Tip1p family of serine-alanine-rich proteins; transcription is induced by cold shock and anaerobiosis | Part of 23-member seripauperin multigene family, active during alcoholic fermentation, regulated by anaerobiosis, inhibited by oxygen, repressed by heme | One of two type V myosin motors (along with MYO2) involved in actin-based transport of cargos; required for mRNA transport, including ASH1 mRNA, and facilitating the growth and movement of ER tubules into the growing bud along with She3p | Bifunctional dehydrogenase and ferrochelatase, involved in the biosynthesis of siroheme, a prosthetic group used by sulfite reductase; required for sulfate assimilation and methionine biosynthesis | Protein involved in retention of membrane proteins, including Sec12p, in the ER; localized to Golgi; functions as a retrieval receptor in returning membrane proteins to the ER | Cell wall adhesin, expressed specifically during mating; may be involved in maintenance of cell wall integrity during mating | Permease that serves as a gamma-aminobutyrate (GABA) transport protein involved in the utilization of GABA as a nitrogen source; catalyzes the transport of putrescine and delta-aminolevulinic acid (ALA); localized to the vacuolar membrane | Regulatory protein of unknown function, pheromone-inducible, involved in the regulation of mating-specific genes and the invasive growth pathway, required for MAP-kinase imposed repression, inhibits pheromone-responsive transcription | Cell wall mannoprotein of the Srp1p/Tip1p family of serine-alanine-rich proteins; expression is downregulated at acidic pH and induced by cold shock and anaerobiosis; abundance is increased in cells cultured without shaking | Presumed antiporter of the DHA1 family of multidrug resistance transporters; contains 12 predicted transmembrane spans; expression of gene is up-regulated in cells exhibiting reduced susceptibility to azoles | Cell wall mannoprotein of the Srp1p/Tip1p family of serine-alanine-rich proteins; expressed under anaerobic conditions and required for anaerobic growth | Putative protein of unknown function | AP-1 accessory protein; colocalizes with clathrin to the late-Golgi apparatus; involved in TGN-endosome transport; physically interacts with AP-1; similar to the mammalian p200; may interact with ribosomes; YJL207C is a non-essential gene | Cell wall mannoprotein with similarity to Tir1p, Tir2p, Tir3p, and Tir4p; expressed under anaerobic conditions, completely repressed during aerobic growth | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | ABC type transmembrane transporter of MRP/CFTR family, found in vacuolar membrane, involved in the transport of unconjugated bilirubin and in heavy metal detoxification via glutathione conjugates, along with Ycf1p | Protein containing GATA family zinc finger motifs | Putative protein of unknown function; YLR415C is not an essential gene | Cell wall mannoprotein of the Srp1p/Tip1p family of serine-alanine-rich proteins; expressed under anaerobic conditions and required for anaerobic growth; transcription is also induced by cold shock | Transporter of thiamine or related compound; shares sequence similarity with Thi7p | Probable membrane protein with a possible role in proton symport of glycerol; member of the MBOAT family of putative membrane-bound O-acyltransferases; Gup1p homolog | Protein of unknown function that may interact with ribosomes, based on co-purification experiments |
| 149 | 41 | 445 | 0.728 | 0.606 | 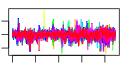 |  | 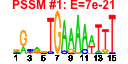 | 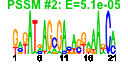 |  | YCLX01W YCR043C YCR051W YCR056W YDL112W YDR156W YDR157W YER154W YGL209W YGR081C YGR286C YHR042W YHR100C YKL125W YML125C YNL310C YNR024W YBL042C YBL069W YDR384C YDR514C YDR526C YGR082W YGR083C YGR109C YGR210C YHR062C YJL075C YJL115W YJR147W YKR043C YKR044W YLR172C YNL282W YNL292W YNL309W YNR018W YOL093W YOR236W YPL080C YPR183W | TRM3 RPA14 OXA1 MIG2 SLX9 BIO2 CPR1 RRN3 PGA3 ZIM17 FUI1 AST1 ATO3 TOM20 GCD2 CLB6 RPP1 CIA1 HMS2 UIP5 DPH5 POP3 PUS4 STB1 AIM38 TRM10 DFR1 DPM1 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the Golgi apparatus; YCR043C is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; contains ankyrin (Ank) repeats; YCR051W is not an essential gene | 2'-O-ribose methyltransferase, catalyzes the ribose methylation of the guanosine nucleotide at position 18 of tRNAs | RNA polymerase I subunit A14 | Mitochondrial inner membrane insertase, mediates the insertion of both mitochondrial- and nuclear-encoded proteins from the matrix into the inner membrane, interacts with mitochondrial ribosomes; conserved from bacteria to animals | Protein containing zinc fingers, involved in repression, along with Mig1p, of SUC2 (invertase) expression by high levels of glucose; binds to Mig1p-binding sites in SUC2 promoter | Protein required for pre-rRNA processing; associated with the 90S pre-ribosome and 43S small ribosomal subunit precursor; interacts with U3 snoRNA; deletion mutant has synthetic fitness defect with an sgs1 deletion mutant | Biotin synthase, catalyzes the conversion of dethiobiotin to biotin, which is the last step of the biotin biosynthesis pathway; complements E. coli bioB mutant | Cytoplasmic peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; binds the drug cyclosporin A | Putative protein of unknown function, required for respiratory growth; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Protein required for transcription of rDNA by RNA polymerase I; transcription factor independent of DNA template; involved in recruitment of RNA polymerase I to rDNA | Essential protein required for maturation of Gas1p and Pho8p, protein trafficking; GFP-fusion protein localizes to the endoplasmic reticulum; null mutants have a cell separation defect | Heat shock protein with a zinc finger motif; essential for protein import into mitochondria; may act with Pam18p to facilitate recognition and folding of imported proteins by Ssc1p (mtHSP70) in the mitochondrial matrix | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the nucleus | High affinity uridine permease, localized to the plasma membrane; not involved in uracil transport | Peripheral membrane protein that interacts with the plasma membrane ATPase Pma1p and has a role in its targeting to the plasma membrane, possibly by influencing its incorporation into lipid rafts | Plasma membrane protein, regulation pattern suggests a possible role in export of ammonia from the cell; phosphorylated in mitochondria; member of the TC 9.B.33 YaaH family of putative transporters | Putative protein of unknown function | Component of the TOM (translocase of outer membrane) complex responsible for recognition and initial import steps for all mitochondrially directed proteins; acts as a receptor for incoming precursor proteins | Delta subunit of the translation initiation factor eIF2B, the guanine-nucleotide exchange factor for eIF2; activity subsequently regulated by phosphorylated eIF2; first identified as a negative regulator of GCN4 expression | B-type cyclin involved in DNA replication during S phase; activates Cdc28p to promote initiation of DNA synthesis; functions in formation of mitotic spindles along with Clb3p and Clb4p; most abundant during late G1 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends | Essential protein involved in assembly of cytosolic and nuclear iron-sulfur proteins | Protein with similarity to heat shock transcription factors; overexpression suppresses the pseudohyphal filamentation defect of a diploid mep1 mep2 homozygous null mutant | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | Protein of unknown function that interacts with Ulp1p, a Ubl (ubiquitin-like protein)-specific protease for Smt3p protein conjugates | Methyltransferase required for synthesis of diphthamide, which is a modified histidine residue of translation elongation factor 2 (Eft1p or Eft2p); not essential for viability; GFP-Dph5p fusion protein localizes to the cytoplasm | Pseudouridine synthase, catalyzes only the formation of pseudouridine-55 (Psi55), a highly conserved tRNA modification, in mitochondrial and cytoplasmic tRNAs; PUS4 overexpression leads to translational derepression of GCN4 (Gcd- phenotype) | Protein with a role in regulation of MBF-specific transcription at Start, phosphorylated by Cln-Cdc28p kinases in vitro; unphosphorylated form binds Swi6p and binding is required for Stb1p function; expression is cell-cycle regulated | Putative protein of unknown function; non-tagged protein is detected in purified mitochondria; null mutant displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | tRNA methyltransferase, methylates the N-1 position of guanosine in tRNAs | Dihydrofolate reductase, part of the dTTP biosynthetic pathway, involved in folate metabolism, possibly required for mitochondrial function | Dolichol phosphate mannose (Dol-P-Man) synthase of the ER membrane, catalyzes the formation of Dol-P-Man from Dol-P and GDP-Man; required for glycosyl phosphatidylinositol membrane anchoring, O mannosylation, and protein glycosylation |
| 271 | 38 | 454 | 0.737 | 0.603 | 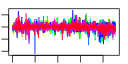 |  | 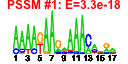 |  |  | YBR137W YLR151C YBR105C YDR248C YDR259C YHL036W YIL001W YIL002C YIL068C YJL020C YJL021C YJL046W YJL047C YJL107C YJL213W YJR048W YJR061W YJR110W YKL038W YKR003W YKR039W YLL006W YLL007C YLR001C YLR228C YLR231C YLR247C YLR309C YML076C YMR317W YNL040W YNL180C YOR124C YOR125C YOR223W YOR363C YOR387C YPL005W | PCD1 VID24 YAP6 MUP3 INP51 SEC6 BBC1 AIM22 YJL047C-A CYC1 YMR1 RGT1 OSH6 GAP1 YLL006W-A ECM22 BNA5 IRC20 IMH1 WAR1 RHO5 UBP2 CAT5 PIP2 AEP3 | Protein of unknown function; localized to the cytoplasm; binds to Replication Protein A (RPA); YBR137W is not an essential gene | Peroxisomal nudix pyrophosphatase with specificity for coenzyme A and CoA derivatives, may function to remove potentially toxic oxidized CoA disulfide from peroxisomes to maintain the capacity for beta-oxidation of fatty acids | Peripheral membrane protein located at Vid (vacuole import and degradation) vesicles; regulates fructose-1,6-bisphosphatase (FBPase) targeting to the vacuole; involved in proteasome-dependent catabolite degradation of FBPase | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative basic leucine zipper (bZIP) transcription factor; overexpression increases sodium and lithium tolerance | Low affinity methionine permease, similar to Mup1p | hypothetical protein | Phosphatidylinositol 4,5-bisphosphate 5-phosphatase, synaptojanin-like protein with an N-terminal Sac1 domain, plays a role in phosphatidylinositol 4,5-bisphosphate homeostasis and in endocytosis; null mutation confers cold-tolerant growth | Essential 88kDa subunit of the exocyst complex, which mediates polarized targeting of secretory vesicles to active sites of exocytosis; dimeric form of Sec6p interacts with Sec9p in vitro and inhibits t-SNARE assembly | Protein possibly involved in assembly of actin patches; interacts with an actin assembly factor Las17p and with the SH3 domains of Type I myosins Myo3p and Myo5p; localized predominantly to cortical actin patches | Putative lipoate-protein ligase A family member; null mutant displays respiratory growth defect, decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Putative protein of unknown function | Putative protein of unknown function; expression is induced by activation of the HOG1 mitogen-activated signaling pathway and this induction is Hog1p/Pbs2p dependent; YJL107C and adjacent ORF, YJL108C are merged in related fungi | Protein of unknown function that may interact with ribosomes; periodically expressed during the yeast metabolic cycle; phosphorylated in vitro by the mitotic exit network (MEN) kinase complex, Dbf2p/Mob1p | Cytochrome c, isoform 1; electron carrier of the mitochondrial intermembrane space that transfers electrons from ubiquinone-cytochrome c oxidoreductase to cytochrome c oxidase during cellular respiration | Putative protein of unknown function; non-essential gene with similarity to Mnn4, a putative membrane protein involved in glycosylation; transcription repressed by Rm101p | Phosphatidylinositol 3-phosphate [PI(3)P] phosphatase, regulates the localization and levels of PI(3)P; involved in cytoplasm to vacuole (CVT) transport; has similarity to the conserved myotubularin dual specificity phosphatase family | Glucose-responsive transcription factor that regulates expression of several glucose transporter (HXT) genes in response to glucose; binds to promoters and acts both as a transcriptional activator and repressor | Member of an oxysterol-binding protein family with overlapping, redundant functions in sterol metabolism and which collectively perform a function essential for viability; GFP-fusion protein localizes to the cell periphery | General amino acid permease; localization to the plasma membrane is regulated by nitrogen source | Putative protein of unknown function; identified by fungal homology and RT-PCR | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLL007C is not an essential gene | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YLR001C is not an essential gene | Sterol regulatory element binding protein, regulates transcription of the sterol biosynthetic genes ERG2 and ERG3; member of the fungus-specific Zn[2]-Cys[6] binuclear cluster family of transcription factors; homologous to Upc2p | Kynureninase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Putative helicase; localized to mitochondria and the nucleus; YLR247C is not an essential gene; null mutant displays increased levels of spontaneous Rad52 foci | Protein involved in vesicular transport, mediates transport between an endosomal compartment and the Golgi, contains a Golgi-localization (GRIP) domain that interacts with activated Arl1p-GTP to localize Imh1p to the Golgi | Homodimeric Zn2Cys6 zinc finger transcription factor; binds to a weak acid response element to induce transcription of PDR12 and FUN34, encoding an acid transporter and a putative ammonia transporter, respectively | Putative protein of unknown function with some similarity to sialidase from Trypanosoma; YMR317W is not an essential gene | Putative protein of unknown function with strong similarity to alanyl-tRNA synthases from Eubacteria; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YNL040W is not an essential gene | Non-essential small GTPase of the Rho/Rac subfamily of Ras-like proteins, likely involved in protein kinase C (Pkc1p)-dependent signal transduction pathway that controls cell integrity | Ubiquitin-specific protease that removes ubiquitin from ubiquitinated proteins, cleaves at the C terminus of ubiquitin fusions; capable of cleaving polyubiquitin and possesses isopeptidase activity | Protein required for ubiquinone (Coenzyme Q) biosynthesis; localizes to the matrix face of the mitochondrial inner membrane in a large complex with ubiquinone biosynthetic enzymes; required for gluconeogenic gene activation | Autoregulatory oleate-specific transcriptional activator of peroxisome proliferation, contains Zn(2)-Cys(6) cluster domain, forms heterodimer with Oaf1p, binds oleate response elements (OREs), activates beta-oxidation genes | Putative protein of unknown function; regulated by the metal-responsive Aft1p transcription factor; highly inducible in zinc-depleted conditions; localizes to the soluble fraction | Peripheral mitochondrial inner membrane protein, located on the matrix face of the membrane; stabilizes the bicistronic AAP1-ATP6 mRNA encoding subunits 6 and 8 of the ATP synthase complex |
| 56 | 54 | 449 | 0.743 | 0.554 | 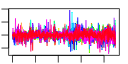 |  | 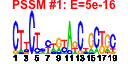 | 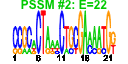 |  | YAL037W YAR042W YAR043C YAR047C YBR134W YBR223C YBR224W YBR225W YBR292C YBR300C YCL069W YCR006C YCR080W YCRX03C YCRX09C YCRX19W YDL039C YDL242W YDR106W YDR135C YDR136C YDR511W YEL008W YEL045C YFL050C YGL176C YGL177W YGL240W YGL262W YGR213C YHL026C YHR130C YHR177W YJL027C YJL216C YKL037W YKL102C YKL118W YKL158W YLR334C YNR071C YOR139C YOR180C YPL021W YAR068W YCR103C YCRX18C YDL071C YDL109C YDR010C YDR048C YER153C YIL102C YMR159C | SWH1 TDP1 VBA3 SRB8 PRM7 ARP10 YCF1 ACN9 ALR2 DOC1 RTA1 AIM26 APE2 DCI1 ECM23 PET122 YIL102C-A ATG16 | Putative protein of unknown function | Protein similar to mammalian oxysterol-binding protein; contains ankyrin repeats; localizes to the Golgi and the nucleus-vacuole junction | Tyrosyl-DNA Phosphodiesterase I, hydrolyzes 3'-phosphotyrosyl bonds to generate 3'-phosphate DNA and tyrosine, involved in the repair of DNA lesions created by topoisomerase I | Putative protein of unknown function; non-essential gene identified in a screen for mutants affected in mannosylphophorylation of cell wall components | Permease of basic amino acids in the vacuolar membrane | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; essential for transcriptional regulation; involved in glucose repression | Pheromone-regulated protein, predicted to have one transmembrane segment; promoter contains Gcn4p binding elements | Component of the dynactin complex, localized to the pointed end of the Arp1p filament; may regulate membrane association of the complex | Vacuolar glutathione S-conjugate transporter of the ATP-binding cassette family, has a role in detoxifying metals such as cadmium, mercury, and arsenite; also transports unconjugated bilirubin; similar to human cystic fibrosis protein CFTR | Protein of the mitochondrial intermembrane space, required for acetate utilization and gluconeogenesis; has orthologs in higher eukaryotes | Probable Mg(2+) transporter; overexpression confers increased tolerance to Al(3+) and Ga(3+) ions | Putative protein of unknown function; deletion mutant is viable and has no detectable phenotype | Processivity factor required for the ubiquitination activity of the anaphase promoting complex (APC), mediates the activity of the APC by contributing to substrate recognition; involved in cyclin proteolysis | Putative protein of unknown function; null mutant displays elevated sensitivity to expression of a mutant huntingtin fragment or of alpha-synuclein; YGL262W is not an essential gene | Protein involved in 7-aminocholesterol resistance; has seven potential membrane-spanning regions | Putative protein of unknown function; YHL026C is not an essential gene; in 2005 the start site was moved 141 nt upstream (see Locus History) | Protein of unknown function, similar to alpha-D-glucosidases; transcriptionally activated by both Pdr8p and Yrm1p, along with transporters and other genes involved in the pleiotropic drug resistance (PDR) phenomenon | Putative protein of unknown function; null mutant is viable and displays increased frequency of mitochondrial genome loss (petite formation) | Zinc-dependent metallopeptidase yscII, may have a role in obtaining leucine from dipeptide substrates; sequence coordinates have changed since RT-PCR analysis showed that the adjacent ORF YKL158W comprises the 5' exon of APE2/YKL157W | Peroxisomal delta(3,5)-delta(2,4)-dienoyl-CoA isomerase, involved in fatty acid metabolism, contains peroxisome targeting signals at amino and carboxy termini | Non-essential protein of unconfirmed function; affects pre-rRNA processing, may act as a negative regulator of the transcription of genes involved in pseudohyphal growth; homologous to Srd1p | Fungal-specific protein of unknown function; induced in respiratory-deficient cells | Putative lipase; involved in lipid metabolism; YDL109C is not an essential gene | Mitochondrial translational activator specific for the COX3 mRNA, acts together with Pet54p and Pet494p; located in the mitochondrial inner membrane | Putative protein of unknown function, identified based on comparisons of the genome sequences of six Saccharomyces species | Protein that interacts with the Atg12p-Atg5p conjugate during formation of the pre-autophagosomal structure; essential for autophagy |
| 167 | 52 | 447 | 0.746 | 0.537 |  |  |  |  |  | YCR085W YJL207C YJR150C YLL059C YML083C YOR024W YOR066W YOR072W YAL067C YAR029W YAR064W YAR069C YBL088C YBR100W YBR144C YCLX03C YCR025C YCR064C YCR106W YDR015C YDR102C YDR106W YDR220C YDR326C YER135C YFL015C YGL088W YGL158W YHR125W YIL025C YIL163C YJL086C YJL160C YJL182C YJR157W YJR162C YKL097C YKL118W YLR184W YLR213C YLR296W YLR334C YLR366W YNL017C YNL204C YNL205C YOL160W YOR082C YOR084W YOR378W YPL187W YPL276W | LAA1 DAN1 MSA1 YOR072W-B SEO1 TEL1 MMS4 RDS1 ARP10 YSP2 RCK1 CRR1 SPS18 LPX1 MF(ALPHA)1 | AP-1 accessory protein; colocalizes with clathrin to the late-Golgi apparatus; involved in TGN-endosome transport; physically interacts with AP-1; similar to the mammalian p200; may interact with ribosomes; YJL207C is a non-essential gene | Cell wall mannoprotein with similarity to Tir1p, Tir2p, Tir3p, and Tir4p; expressed under anaerobic conditions, completely repressed during aerobic growth | Putative protein of unknown function; strong increase in transcript abundance during anaerobic growth compared to aerobic growth; cells deleted for YML083C do not exhibit growth defects in anerobic or anaerobic conditions | Activator of G1-specific transcription factors, MBF and SBF, that regulates both the timing of G1-specific gene transcription, and cell cycle initiation; potential Cdc28p substrate | Putative protein of unknown function; identified by expression profiling and mass spectrometry | Putative permease, member of the allantoate transporter subfamily of the major facilitator superfamily; mutation confers resistance to ethionine sulfoxide | Member of DUP240 gene family but contains no transmembrane domains; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Putative protein of unknown function | Protein kinase primarily involved in telomere length regulation; contributes to cell cycle checkpoint control in response to DNA damage; functionally redundant with Mec1p; homolog of human ataxia telangiectasia (ATM) gene | Subunit of the structure-specific Mms4p-Mus81p endonuclease that cleaves branched DNA; involved in recombination and DNA repair | Zinc cluster transcription factor involved in conferring resistance to cycloheximide | Component of the dynactin complex, localized to the pointed end of the Arp1p filament; may regulate membrane association of the complex | Protein involved in programmed cell death; mutant shows resistance to cell death induced by amiodarone or intracellular acidification | Protein kinase involved in the response to oxidative stress; identified as suppressor of S. pombe cell cycle checkpoint mutations | Putative protein of unknown function; member of the PIR (proteins with internal repeats) family of cell wall proteins; non-essential gene that is required for sporulation; mRNA is weakly cell cycle regulated, peaking in mitosis | Putative glycoside hydrolase of the spore wall envelope; required for normal spore wall assembly, possibly for cross-linking between the glucan and chitosan layers; expressed during sporulation | Protein of unknown function, contains a putative zinc-binding domain; expressed during sporulation | Oleic acid-inducible, peroxisomal matrix localized lipase; transcriptionally activated by Yrm1p along with genes involved in multidrug resistance; peroxisomal import is dependent on the PTS1 receptor, Pex5p and on self-interaction | Putative protein of unknown function; belongs to the DHA2 family of drug:H+ antiporters; YOR378W is not an essential gene | Mating pheromone alpha-factor, made by alpha cells; interacts with mating type a cells to induce cell cycle arrest and other responses leading to mating; also encoded by MF(ALPHA)2, although MF(ALPHA)1 produces most alpha-factor |
| 138 | 51 | 448 | 0.772 | 0.589 |  |  | 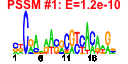 | 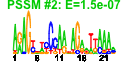 |  | YBL062W YDL209C YDR121W YIL003W YJL071W YJL169W YJL175W YJR129C YKL048C YKR078W YLL004W YLL038C YLR045C YLR059C YLR096W YLR399C YML038C YMR003W YMR132C YMR138W YNL226W YNL290W YNL304W YNR021W YOL093W YOL095C YOR213C YOR301W YOR333C YOR345C YOR357C YPL007C YPL029W YPL063W YPL233W YPR058W YPR062W YBL020W YBR153W YDR113C YDR354W YDR383C YJL181W YLL002W YLR181C YNL259C YPL125W YPL141C YPR101W YPR116W YPR130C | CWC2 DPB4 CFD1 ARG2 ELM1 ORC3 ENT4 STU2 REX2 KIN2 BDF1 YMD8 AIM34 JLP2 CIN4 RFC3 YPT11 TRM10 HMI1 SAS5 RAX1 SNX3 TFC8 SUV3 TIM50 NSL1 YMC1 FCY1 RFT1 RIB7 PDS1 TRP4 NKP1 RTT109 VTA1 ATX1 KAP120 SNT309 | Protein involved in pre-mRNA splicing, component of a complex containing Cef1p; interacts with Prp19p; contains an RNA recognition motif; has similarity to S. pombe Cwf2p | Shared subunit of DNA polymerase (II) epsilon and of ISW2/yCHRAC chromatin accessibility complex; involved in both chromosomal DNA replication and in inheritance of telomeric silencing | Highly conserved, iron-sulfur cluster binding protein localized in the cytoplasm; forms a complex with Nbp35p that is involved in iron-sulfur protein assembly in the cytosol | Acetylglutamate synthase (glutamate N-acetyltransferase), mitochondrial enzyme that catalyzes the first step in the biosynthesis of the arginine precursor ornithine; forms a complex with Arg5,6p | Putative protein of unknown function; predicted S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Serine/threonine protein kinase that regulates cellular morphogenesis, septin behavior, and cytokinesis; required for the regulation of other kinases; forms part of the bud neck ring | Cytoplasmic protein of unknown function, has similarity to Vps5p; potential Cdc28p substrate; contains a Phox homology (PX) domain and specifically binds phosphatidylinositol 3-phosphate (PtdIns-3-P) | Subunit of the origin recognition complex, which directs DNA replication by binding to replication origins and is also involved in transcriptional silencing | Protein of unknown function, contains an N-terminal epsin-like domain | Microtubule-associated protein (MAP) of the XMAP215/Dis1 family; regulates microtubule dynamics during spindle orientation and metaphase chromosome alignment; interacts with spindle pole body component Spc72p | RNA exonuclease, required for U4 snRNA maturation; functions redundantly with Rnh70p in 5.8S rRNA maturation, and with Rnh70p and Rex3p in processing of U5 snRNA and RNase P RNA; member of RNase D family of exonucleases | Serine/threonine protein kinase involved in regulation of exocytosis; localizes to the cytoplasmic face of the plasma membrane; closely related to Kin1p | Protein involved in transcription initiation at TATA-containing promoters; associates with the basal transcription factor TFIID; contains two bromodomains; corresponds to the C-terminal region of mammalian TAF1; redundant with Bdf2p | Putative nucleotide sugar transporter, has similarity to Vrg4p | Protein of unknown function; GFP-fusion protein localizes to the mitochondria; null mutant is viable and displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Protein of unknown function, contains sequence that closely resembles a J domain (typified by the E. coli DnaJ protein) | GTP-binding protein involved in beta-tubulin (Tub2p) folding; isolated as mutant with increased chromosome loss and sensitivity to benomyl | Subunit of heteropentameric Replication factor C (RF-C), which is a DNA binding protein and ATPase that acts as a clamp loader of the proliferating cell nuclear antigen (PCNA) processivity factor for DNA polymerases delta and epsilon | Rab-type small GTPase that interacts with the C-terminal tail domain of Myo2p to mediate distribution of mitochondria to daughter cells | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum; YNR021W is not an essential gene | tRNA methyltransferase, methylates the N-1 position of guanosine in tRNAs | Mitochondrial inner membrane localized ATP-dependent DNA helicase, required for the maintenance of the mitochondrial genome; not required for mitochondrial transcription; has homology to E. coli helicase uvrD | Subunit of the SAS complex (Sas2p, Sas4p, Sas5p), which acetylates free histones and nucleosomes and regulates transcriptional silencing; stimulates Sas2p HAT activity | Protein involved in bud site selection during bipolar budding; localization requires Rax2p; has similarity to members of the insulin-related peptide superfamily | Sorting nexin required to maintain late-Golgi resident enzymes in their proper location by recycling molecules from the prevacuolar compartment; contains a PX domain and sequence similarity to human Snx3p | One of six subunits of RNA polymerase III transcription initiation factor complex (TFIIIC); part of TFIIIC TauB domain that binds BoxB promoter sites of tRNA and other genes; linker between TauB and TauA domains; human homolog is TFIIIC-90 | ATP-dependent RNA helicase, component of the mitochondrial degradosome along with the RNase Dss1p; the degradosome associates with the ribosome and mediates turnover of aberrant or unprocessed RNAs | Constituent of the mitochondrial inner membrane presequence translocase (TIM23 complex); may promote binding of incoming precursor proteins to the intermembrane space domain of Tom22p during translocation | Essential component of the MIND kinetochore complex (Mtw1p Including Nnf1p-Nsl1p-Dsn1p) which joins kinetochore subunits contacting DNA to those contacting microtubules; required for accurate chromosome segregation | Mitochondrial protein, putative inner membrane transporter with a role in oleate metabolism and glutamate biosynthesis; member of the mitochondrial carrier (MCF) family; has similarity with Ymc2p | Cytosine deaminase, zinc metalloenzyme that catalyzes the hydrolytic deamination of cytosine to uracil; of biomedical interest because it also catalyzes the deamination of 5-fluorocytosine (5FC) to form anticancer drug 5-fluorouracil (5FU) | Flippase, essential integral membrane protein that is required for translocation of Man5GlcNac2-PP-Dol from the cytoplasmic side to the lumenal side of the ER membrane; mutation is suppressed by expression human p53 protein | Diaminohydroxyphoshoribosylaminopyrimidine deaminase; catalyzes the second step of the riboflavin biosynthesis pathway | Securin that inhibits anaphase by binding separin Esp1p, also blocks cyclin destruction and mitotic exit, essential for cell cycle arrest in mitosis in the presence of DNA damage or aberrant mitotic spindles; also present in meiotic nuclei | Anthranilate phosphoribosyl transferase of the tryptophan biosynthetic pathway, catalyzes the phosphoribosylation of anthranilate, subject to the general control system of amino acid biosynthesis | Non-essential kinetochore protein, subunit of the Ctf19 central kinetochore complex (Ctf19p-Mcm21p-Okp1p-Mcm22p-Mcm16p-Ctf3p-Chl4p-Mcm19p- Nkp1p-Nkp2p-Ame1p-Mtw1p) | Putative protein of unknown function; expression is cell-cycle regulated as shown by microarray analysis | Histone acetyltransferase critical for cell survival in the presence of DNA damage during S phase, acetylates H3-K56; plays a role in regulation of Ty1 transposition | Multivesicular body (MVB) protein involved in endosomal protein sorting; regulates Vps4p activity by promoting its oligimeriztion; binds to Vps4p, Vps20p, and Vps60p; may act at a late step in MVB formation | Cytosolic copper metallochaperone that transports copper to the secretory vesicle copper transporter Ccc2p for eventual insertion into Fet3p, which is a multicopper oxidase required for high-affinity iron uptake | Karyopherin with a role in the assembly or export of 60S ribosomal subunits | Putative protein kinase; similar to Kin4p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YPL141C is not an essential gene | Component of NineTeen complex (NTC) containing Prp19p involved in mRNA splicing, interacts physically and genetically with Prp19p | Putative protein of unknown function |
| 52 | 45 | 445 | 0.790 | 0.590 |  |  | 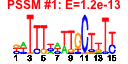 |  |  | YBR076W YBR295W YCL008C YCL027W YCLX07W YCR042C YCR064C YCR068W YCR089W YCRX18C YDR216W YDR403W YEL030W YEL076W-C YHL049C YHR213W YIL067C YIL102C YJL170C YJR035W YJR053W YJR127C YJR128W YKL178C YLR040C YLR122C YLR124W YLR436C YLR462W YLR466W YMR182C YMR207C YMR320W YMR324C YNL279W YNL321W YNR048W YOR034C YOR318C YPL156C YPL192C YPR049C YPR070W YPR203W YDR069C | ECM8 PCA1 STP22 FUS1 TAF2 ATG15 FIG2 ADR1 DIT1 ECM10 YIL102C-A ASG7 RAD26 BFA1 RSF2 STE3 ECM30 YRF1-1 RGM1 HFA1 PRM1 VNX1 CRF1 AKR2 PRM4 PRM3 ATG11 MED1 DOA4 | Non-essential protein of unknown function | Cadmium transporting P-type ATPase; may also have a role in copper and iron homeostasis; S288C and most other lab strains contain a G970R mutation which eliminates normal cadmium transport function | Component of the ESCRT-I complex, which is involved in ubiquitin-dependent sorting of proteins into the endosome; homologous to the mouse and human Tsg101 tumor susceptibility gene; mutants exhibit a Class E Vps phenotype | Membrane protein localized to the shmoo tip, required for cell fusion; expression regulated by mating pheromone; proposed to coordinate signaling, fusion, and polarization events required for fusion; potential Cdc28p substrate | TFIID subunit (150 kDa), involved in RNA polymerase II transcription initiation | Lipase, required for intravacuolar lysis of autophagic bodies; located in the endoplasmic reticulum membrane and targeted to intravacuolar vesicles during autophagy via the multivesicular body (MVB) pathway | Cell wall adhesin, expressed specifically during mating; may be involved in maintenance of cell wall integrity during mating | Carbon source-responsive zinc-finger transcription factor, required for transcription of the glucose-repressed gene ADH2, of peroxisomal protein genes, and of genes required for ethanol, glycerol, and fatty acid utilization | Sporulation-specific enzyme required for spore wall maturation, involved in the production of a soluble LL-dityrosine-containing precursor of the spore wall; transcripts accumulate at the time of prospore enclosure | Heat shock protein of the Hsp70 family, localized in mitochondrial nucleoids, plays a role in protein translocation, interacts with Mge1p in an ATP-dependent manner; overexpression induces extensive mitochondrial DNA aggregations | Putative protein of unknown function | Uncharacterized protein of unknown function | Putative protein of unknown function, identified based on comparisons of the genome sequences of six Saccharomyces species | Protein that regulates signaling from a G protein beta subunit Ste4p and its relocalization within the cell; specific to a-cells and induced by alpha-factor | Protein involved in transcription-coupled repair nucleotide excision repair of UV-induced DNA lesions; homolog of human CSB protein | Component of the GTPase-activating Bfa1p-Bub2p complex involved in multiple cell cycle checkpoint pathways that control exit from mitosis | Zinc-finger protein involved in transcriptional control of both nuclear and mitochondrial genes, many of which specify products required for glycerol-based growth, respiration, and other functions | Receptor for a factor receptor, transcribed in alpha cells and required for mating by alpha cells, couples to MAP kinase cascade to mediate pheromone response; ligand bound receptors are endocytosed and recycled to the plasma membrane; GPCR | Putative protein of unknown function; localizes to the cell wall; predicted to be a GPI-attached protein; upregulated by Mcm1p-Alpha1p transcription factor; partially overlaps the dubious ORF YLR041W; YLR040C is not essential | Putative protein of unknown function with similarity to helicases; YLR462W is within the telomere on the right arm of chromosome XII | Helicase encoded by the Y' element of subtelomeric regions, highly expressed in the mutants lacking the telomerase component TLC1; potentially phosphorylated by Cdc28p | Putative transcriptional repressor with proline-rich zinc fingers; overproduction impairs cell growth | Mitochondrial acetyl-coenzyme A carboxylase, catalyzes the production of malonyl-CoA in mitochondrial fatty acid biosynthesis | Pheromone-regulated multispanning membrane protein involved in membrane fusion during mating; predicted to have 5 transmembrane segments and a coiled coil domain; localizes to the shmoo tip; regulated by Ste12p | Low affinity vacuolar membrane localized monovalent cation/H+ antiporter; member of the calcium exchanger (CAX) family; potential Cdc28p substrate | Transcriptional corepressor involved in the regulation of ribosomal protein gene transcription via the TOR signaling pathway and protein kinase A, phosphorylated by activated Yak1p which promotes accumulation of Crf1p in the nucleus | Ankyrin repeat-containing protein similar to Akr1p; member of a family of putative palmitoyltransferases containing an Asp-His-His-Cys-cysteine rich (DHHC-CRD) domain; possibly involved in constitutive endocytosis of Ste3p | Pheromone-regulated protein, predicted to have 1 transmembrane segment; transcriptionally regulated by Ste12p during mating and by Cat8p during the diauxic shift | Pheromone-regulated protein required for karyogamy; localizes to the inner membrane of the nuclear envelope | Peripheral membrane protein required for delivery of aminopeptidase I (Lap4p) to the vacuole in the cytoplasm-to-vacuole targeting pathway; also required for peroxisomal degradation (pexophagy) | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; essential for transcriptional regulation | Ubiquitin isopeptidase, required for recycling ubiquitin from proteasome-bound ubiquitinated intermediates, acts at the late endosome/prevacuolar compartment to recover ubiquitin from ubiquitinated membrane proteins en route to the vacuole |
| 162 | 40 | 442 | 0.805 | 0.567 |  |  |  |  |  | YCR035C YER148W YGL106W YGR095C YGR228W YIL008W YJL009W YJL011C YKL122C YLR008C YLR101C YLR146C YLR405W YLR420W YML126C YNL108C YNL109W YNL149C YNL150W YPL239W YCR043C YDR086C YDR156W YDR157W YDR397C YGR037C YJL008C YJL117W YJL121C YJR015W YJR024C YKL024C YLR287C YML058W YMR123W YNL303W YOL052C YOR047C YOR332W YPR050C | RRP43 SPT15 MLC1 RRP46 URM1 RPC17 SRP21 PAM18 SPE4 DUS4 URA4 ERG13 PGA2 YAR1 SSS1 RPA14 NCB2 ACB1 CCT8 PHO86 RPE1 MDE1 URA6 HUG1 PKR1 DDR2 STD1 VMA4 | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex with Rrp41p, Rrp42p, Rrp4p and Dis3p; required for efficient maturation of 5.8S, 18S and 25S rRNA | TATA-binding protein, general transcription factor that interacts with other factors to form the preinitiation complex at promoters, essential for viability | Essential light chain for Myo1p, light chain for Myo2p; stabilizes Myo2p by binding to the neck region; interacts with Myo1p, Iqg1p, and Myo2p to coordinate formation and contraction of the actomyosin ring with targeted membrane deposition | Protein involved in rRNA processing; component of the exosome 3->5 exonuclease complex | Ubiquitin-like protein with weak sequence similarity to ubiquitin; depends on the E1-like activating enzyme Uba4p; molecular function of the Urm1p pathway is unknown, but it is required for normal growth, particularly at high temperature | RNA polymerase III subunit C17; physically interacts with C31, C11, and TFIIIB70; may be involved in the recruitment of pol III by the preinitiation complex | Subunit of the signal recognition particle (SRP), which functions in protein targeting to the endoplasmic reticulum membrane; not found in mammalian SRP; forms a pre-SRP structure in the nucleolus that is translocated to the cytoplasm | J-protein co-chaperone of the mitochondrial import motor associated with the presequence translocase, with Ssc1p, Tim44p, Mge1p, and Pam16p; stimulates ATPase activity of Ssc1p to drive mitochondrial import; activity is inhibited by Pam16p | Spermine synthase, required for the biosynthesis of spermine and also involved in biosynthesis of pantothenic acid | Dihydrouridine synthase, member of a widespread family of conserved proteins including Smm1p, Dus1p, and Dus3p | Dihydroorotase, catalyzes the third enzymatic step in the de novo biosynthesis of pyrimidines, converting carbamoyl-L-aspartate into dihydroorotate | 3-hydroxy-3-methylglutaryl-CoA (HMG-CoA) synthase, catalyzes the formation of HMG-CoA from acetyl-CoA and acetoacetyl-CoA; involved in the second step in mevalonate biosynthesis | Putative protein of unknown function with similarity to Tfc7p and prokaryotic phosphotransfer enzymes; null mutant shows alterations in glucose metabolism; GFP-fusion protein localizes to the cytoplasm and nucleus | Essential protein required for maturation of Gas1p and Pho8p; involved in protein trafficking; GFP-fusion protein localizes to the ER and YFP-fusion protein to the nuclear envelope-ER network; null mutants have a cell separation defect | Cytoplasmic ankyrin-repeat containing protein of unknown function, proposed to link the processes of 40S ribosomal subunit biogenesis and adaptation to osmotic and oxidative stress; expression repressed by heat shock | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the Golgi apparatus; YCR043C is not an essential gene | Subunit of the Sec61p translocation complex (Sec61p-Sss1p-Sbh1p) that forms a channel for passage of secretory proteins through the endoplasmic reticulum membrane, and of the Ssh1p complex (Ssh1p-Sbh2p-Sss1p); interacts with Ost4p and Wbp1p | RNA polymerase I subunit A14 | Subunit of a heterodimeric NC2 transcription regulator complex with Bur6p; complex binds to TBP and can repress transcription by preventing preinitiation complex assembly or stimulate activated transcription; homologous to human NC2beta | Acyl-CoA-binding protein, transports newly synthesized acyl-CoA esters from fatty acid synthetase (Fas1p-Fas2p) to acyl-CoA-consuming processes | Subunit of the cytosolic chaperonin Cct ring complex, related to Tcp1p, required for the assembly of actin and tubulins in vivo | Endoplasmic reticulum (ER) resident protein required for ER exit of the high-affinity phosphate transporter Pho84p, specifically required for packaging of Pho84p into COPII vesicles | D-ribulose-5-phosphate 3-epimerase, catalyzes a reaction in the non-oxidative part of the pentose-phosphate pathway; mutants are sensitive to oxidative stress | Putative protein of unknown function; localizes to the endoplasmic reticulum and cytoplasm; predicted to encode a membrane transporter based on phylogenetic analysis; YJR015W is a non-essential gene | Putative methylthio-ribulose-1-phosphate dehydratase; acts in the methionine salvage pathway; potential Smt3p sumoylation substrate; expression downregulated by caspofungin and deletion mutant is caspofungin resistant | Uridylate kinase, catalyzes the seventh enzymatic step in the de novo biosynthesis of pyrimidines, converting uridine monophosphate (UMP) into uridine-5'-diphosphate (UDP) | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR287C is not an essential gene | Protein involved in the Mec1p-mediated checkpoint pathway that responds to DNA damage or replication arrest, transcription is induced by DNA damage | V-ATPase assembly factor, functions with other V-ATPase assembly factors in the ER to efficiently assemble the V-ATPase membrane sector (V0); overproduction confers resistance to Pichia farinosa killer toxin | Multistress response protein, expression is activated by a variety of xenobiotic agents and environmental or physiological stresses | Protein involved in control of glucose-regulated gene expression; interacts with protein kinase Snf1p, glucose sensors Snf3p and Rgt2p, and TATA-binding protein Spt15p; acts as a regulator of the transcription factor Rgt1p | Subunit E of the eight-subunit V1 peripheral membrane domain of the vacuolar H+-ATPase (V-ATPase), an electrogenic proton pump found throughout the endomembrane system; required for the V1 domain to assemble onto the vacuolar membrane |
| 90 | 42 | 442 | 0.808 | 0.574 | 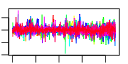 |  |  |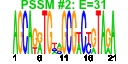  |  | YBR120C YBR294W YCLX09W YDL005C YDL190C YDL194W YDR369C YER184C YGL184C YGR125W YGR238C YHR131C YHR176W YHR211W YIL122W YIL146C YIL170W YIR031C YJR154W YKR017C YKR106W YLL017W YLR364W YLR393W YMR017W YMR095C YNL234W YOL154W YOR178C YOR328W YPL261C YPL264C YPR013C YPR194C YBR295W YCL073C YGL178W YIL169C YLL053C YLR092W YPL272C YPR195C | CBP6 SUL1 MED2 UFD2 SNF3 XRS2 STR3 KEL2 FMO1 FLO5 POG1 ECM37 DAL7 ATP10 SPO20 SNO1 ZPS1 GAC1 PDR10 OPT2 PCA1 MTH1 SUL2 | Mitochondrial translational activator of the COB mRNA; phosphorylated | High affinity sulfate permease; sulfate uptake is mediated by specific sulfate transporters Sul1p and Sul2p, which control the concentration of endogenous activated sulfate intermediates | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; essential for transcriptional regulation | Ubiquitin chain assembly factor (E4) that cooperates with a ubiquitin-activating enzyme (E1), a ubiquitin-conjugating enzyme (E2), and a ubiquitin protein ligase (E3) to conjugate ubiquitin to substrates; also functions as an E3 | Plasma membrane glucose sensor that regulates glucose transport; has 12 predicted transmembrane segments; long cytoplasmic C-terminal tail is required for low glucose induction of hexose transporter genes HXT2 and HXT4 | Protein required for DNA repair; component of the Mre11 complex, which is involved in double strand breaks, meiotic recombination, telomere maintenance, and checkpoint signaling | Putative zinc cluster protein; deletion confers sensitivity to Calcufluor white, and prevents growth on glycerol or lactate as sole carbon source | Cystathionine beta-lyase, converts cystathionine into homocysteine | Putative protein of unknown function; deletion mutant has decreased rapamycin resistance but normal wormannin resistance; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Protein that functions in a complex with Kel1p to negatively regulate mitotic exit, interacts with Tem1p and Lte1p; localizes to regions of polarized growth; potential Cdc28p substrate | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Flavin-containing monooxygenase, localized to the cytoplasmic face of the ER membrane; catalyzes oxidation of biological thiols to maintain the ER redox buffer ratio for correct folding of disulfide-bonded proteins | Lectin-like cell wall protein (flocculin) involved in flocculation, binds to mannose chains on the surface of other cells, confers floc-forming ability that is chymotrypsin resistant but heat labile; similar to Flo1p | Putative transcriptional activator that promotes recovery from pheromone induced arrest; inhibits both alpha-factor induced G1 arrest and repression of CLN1 and CLN2 via SCB/MCB promoter elements; potential Cdc28p substrate; SBF regulated | Non-essential protein of unknown function | Malate synthase, role in allantoin degradation unknown; expression sensitive to nitrogen catabolite repression and induced by allophanate, an intermediate in allantoin degradation | Putative protein of unknown function; contains a RING finger motif | Protein of unconfirmed function; displays a topology characteristic of the Major Facilitators Superfamily of membrane proteins; coding sequence 98% identical to that of YCL073C | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR364W is not an essential gene | Mitochondrial inner membrane protein required for assembly of the F0 sector of mitochondrial F1F0 ATP synthase, interacts genetically with ATP6 | Meiosis-specific subunit of the t-SNARE complex, required for prospore membrane formation during sporulation; similar to but not functionally redundant with Sec9p; SNAP-25 homolog | Protein of unconfirmed function, involved in pyridoxine metabolism; expression is induced during stationary phase; forms a putative glutamine amidotransferase complex with Snz1p, with Sno1p serving as the glutaminase | Similar to globins and has a functional heme-binding domain; involved in glucose signaling or metabolism; regulated by Rgt1p | Putative GPI-anchored protein; transcription is induced under low-zinc conditions, as mediated by the Zap1p transcription factor, and at alkaline pH | Regulatory subunit for Glc7p type-1 protein phosphatase (PP1), tethers Glc7p to Gsy2p glycogen synthase, binds Hsf1p heat shock transcription factor, required for induction of some HSF-regulated genes under heat shock | ATP-binding cassette (ABC) transporter, multidrug transporter involved in the pleiotropic drug resistance network; regulated by Pdr1p and Pdr3p | Putative membrane protein of unknown function; physically interacts with Hsp82p; YPL264C is not an essential gene | Putative zinc finger protein; YPR013C is not an essential gene | Oligopeptide transporter; member of the OPT family, with potential orthologs in S. pombe and C. albicans | Cadmium transporting P-type ATPase; may also have a role in copper and iron homeostasis; S288C and most other lab strains contain a G970R mutation which eliminates normal cadmium transport function | Protein of unconfirmed function; displays a topology characteristic of the Major Facilitators Superfamily of membrane proteins; coding sequence 98% identical to that of YKR106W | Negative regulator of the glucose-sensing signal transduction pathway, required for repression of transcription by Rgt1p; interacts with Rgt1p and the Snf3p and Rgt2p glucose sensors; phosphorylated by Yck1p, triggering Mth1p degradation | Putative protein of unknown function; serine/threonine rich and highly similar to YOL155C, a putative glucan alpha-1,4-glucosidase; transcript is induced in both high and low pH environments; YIL169C is a non-essential gene | Putative protein; in the Sigma 1278B strain background YLL053C is contiguous with AQY2 which encodes an aquaporin | Putative protein of unknown function; gene expression induced in response to ketoconazole; YPL272C is not an essential gene |
| 140 | 48 | 439 | 0.814 | 0.578 | 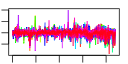 |  |  | 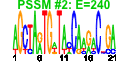 |  | YDR243C YJR060W YJR097W YJR144W YKL024C YKL128C YKL214C YLR018C YLR275W YLR398C YLR419W YMR010W YNL072W YNL228W YNR020C YNR025C YNR041C YOR079C YOR080W YOR104W YOR105W YOR203W YPL001W YPL038W YPL122C YPL175W YPR050C YPR059C YPR125W YPR136C YBR073W YBR156C YDR049W YIL016W YJL195C YJR054W YJR112W YKL074C YKR083C YLR215C YML074C YMR274C YNR027W YOL090W YOR033C YOR242C YPR126C YPR186C | PRP28 CBF1 JJJ3 MGM101 URA6 PMU1 YRA2 POM34 SMD2 SKI2 RNH201 ATP23 COQ2 ATX2 DIA2 PIN2 HAT1 YPL038W-A TFB2 SPT14 YLH47 RDH54 SLI15 SNL1 NNF1 MUD2 DAD2 CDC123 FPR3 RCE1 BUD17 MSH2 EXO1 SSP2 PZF1 | RNA helicase in the DEAD-box family, involved in RNA isomerization at the 5' splice site | Helix-loop-helix protein that binds the motif CACRTG, which is present at several sites including MET gene promoters and centromere DNA element I (CDEI); required for nucleosome positioning at this motif; targets Isw1p to DNA | Protein of unknown function, contains a J-domain, which is a region with homology to the E. coli DnaJ protein | Protein involved in mitochondrial genome maintenance; component of the mitochondrial nucleoid, required for the repair of oxidative mtDNA damage | Uridylate kinase, catalyzes the seventh enzymatic step in the de novo biosynthesis of pyrimidines, converting uridine monophosphate (UMP) into uridine-5'-diphosphate (UDP) | Putative phosphomutase, contains a region homologous to the active site of phosphomutases; overexpression suppresses the histidine auxotrophy of an ade3 ade16 ade17 triple mutant and the temperature sensitivity of a tps2 mutant | Member of the REF (RNA and export factor binding proteins) family; when overexpressed, can substitute for the function of Yra1p in export of poly(A)+ mRNA from the nucleus | Integral membrane protein of the nuclear pore; has an important role in maintaining the architecture of the pore complex | Core Sm protein Sm D2; part of heteroheptameric complex (with Smb1p, Smd1p, Smd3p, Sme1p, Smx3p, and Smx2p) that is part of the spliceosomal U1, U2, U4, and U5 snRNPs; homolog of human Sm D2 | Putative RNA helicase, involved in exosome mediated 3' to 5' mRNA degradation and translation inhibition of non-poly(A) mRNAs; forms complex with Ski3p and Ski8p; required for repressing propagation of dsRNA viruses | Putative helicase with limited sequence similarity to human Rb protein; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YLR419W is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YMR010W is not an essential gene; YMR010W mRNA is transcribed with ADI1 | Ribonuclease H2 catalytic subunit, removes RNA primers during Okazaki fragment synthesis; cooperates with Rad27p nuclease | Putative metalloprotease of the mitochondrial inner membrane, required for processing of Atp6p; has an additional role in assembly of the F0 sector of the F1F0 ATP synthase complex | Para hydroxybenzoate: polyprenyl transferase, catalyzes the second step in ubiquinone (coenzyme Q) biosynthesis | Golgi membrane protein involved in manganese homeostasis; overproduction suppresses the sod1 (copper, zinc superoxide dismutase) null mutation | Origin-binding F-box protein that forms an SCF ubiquitin ligase complex with Skp1p and Cdc53p; plays a role in DNA replication, involved in invasive and pseudohyphal growth | Protein that induces appearance of [PIN+] prion when overproduced | Catalytic subunit of the Hat1p-Hat2p histone acetyltransferase complex that uses the cofactor acetyl coenzyme A, to acetylate free nuclear and cytoplasmic histone H4; involved in telomeric silencing and DNA double-strand break repair | Putative protein of unknown function; identified by fungal homology and RT-PCR | Subunit of TFIIH and nucleotide excision repair factor 3 complexes, involved in transcription initiation, required for nucleotide excision repair, similar to 52 kDa subunit of human TFIIH | UDP-GlcNAc-binding and catalytic subunit of the enzyme that mediates the first step in glycosylphosphatidylinositol (GPI) biosynthesis, mutations cause defects in transcription and in biogenesis of cell wall proteins | Mitochondrial inner membrane protein exposed to the mitochondrial matrix, associates with mitochondrial ribosomes, NOT required for respiratory growth; homolog of human Letm1, a protein implicated in Wolf-Hirschhorn syndrome | DNA-dependent ATPase, stimulates strand exchange by modifying the topology of double-stranded DNA; involved in recombinational repair of DNA double-strand breaks during mitosis and meiosis; proposed to be involved in crossover interference | Subunit of the Ipl1p-Sli15p-Bir1p complex that regulates kinetochore-microtubule attachments, activation of the spindle tension checkpoint, and mitotic spindle disassembly; regulates the activity and localization of the Ipl1p aurora kinase | Zinc finger protein; putative transcription factor that may interact with proteins involved in histone acetylation or deacetylation; may be involved in altering acetylation on histone lysines | Protein of unknown function proposed to be involved in nuclear pore complex biogenesis and maintenance as well as protein folding; has similarity to the mammalian BAG-1 protein | Vacuolar protein of unknown function; potential Cdc28p substrate | Essential component of the MIND kinetochore complex (Mtw1p Including Nnf1p-Nsl1p-Dsn1p) which joins kinetochore subunits contacting DNA to those contacting microtubules; required for accurate chromosome segregation | Protein involved in early pre-mRNA splicing; component of the pre-mRNA-U1 snRNP complex, the commitment complex; interacts with Msl5p/BBP splicing factor and Sub2p; similar to metazoan splicing factor U2AF65 | Essential subunit of the Dam1 complex (aka DASH complex), couples kinetochores to the force produced by MT depolymerization thereby aiding in chromosome segregation; is transferred to the kinetochore prior to mitosis | Protein involved in nutritional control of the cell cycle; regulates abundance of the translation initiation factor eIF2; ortholog of human D123 protein | Nucleolar peptidyl-prolyl cis-trans isomerase (PPIase); FK506 binding protein; phosphorylated by casein kinase II (Cka1p-Cka2p-Ckb1p-Ckb2p) and dephosphorylated by Ptp1p | Type II CAAX prenyl protease involved in the proteolysis and maturation of Ras and the a-factor mating pheromone | Protein involved in bud-site selection; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Protein that forms heterodimers with Msh3p and Msh6p that bind to DNA mismatches to initiate the mismatch repair process; contains a Walker ATP-binding motif required for repair activity; Msh2p-Msh6p binds to and hydrolyzes ATP | 5'-3' exonuclease and flap-endonuclease involved in recombination, double-strand break repair and DNA mismatch repair; member of the Rad2p nuclease family, with conserved N and I nuclease domains | Sporulation specific protein that localizes to the spore wall; required for sporulation at a point after meiosis II and during spore wall formation; SSP2 expression is induced midway in meiosis | Transcription factor IIIA (TFIIIA), essential protein with nine C2H2 Zn-fingers, binds the 5S rRNA gene through the zinc finger domain and directs assembly of a multiprotein initiation complex for RNA polymerase III; also binds DNA |
| 276 | 52 | 436 | 0.817 | 0.615 |  |  | 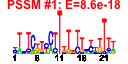 |  |  | YDL066W YNL104C YBR043C YBR096W YBR115C YBR145W YBR256C YCL018W YCLX10C YCR023C YDL198C YDR035W YER052C YER069W YER090W YER175C YFR055W YGL117W YGR239C YHL038C YHR018C YHR029C YHR071W YHR162W YIL074W YIL116W YIR034C YJL088W YJR025C YJR109C YJR111C YJR148W YKL211C YLR027C YLR089C YML116W YMR062C YMR086W YNL129W YNL183C YNL311C YOL030W YOL058W YOL064C YOL119C YOL140W YOR079C YOR080W YOR130C YOR303W YPL202C YPL250C | IDP1 LEU4 QDR3 LYS2 ADH5 RIB5 LEU2 GGC1 ARO3 HOM3 ARG5,6 TRP2 TMT1 IRC7 PEX21 CBP2 ARG4 YHI9 PCL5 HIS5 LYS1 ARG3 BNA1 CPA2 BAT2 TRP3 AAT2 ALT1 ATR1 ECM40 KIC1 NPR1 SKP2 GAS5 ARG1 MET22 MCH4 ARG8 ATX2 DIA2 ORT1 CPA1 AFT2 ICY2 | Mitochondrial NADP-specific isocitrate dehydrogenase, catalyzes the oxidation of isocitrate to alpha-ketoglutarate; not required for mitochondrial respiration and may function to divert alpha-ketoglutarate to biosynthetic processes | Alpha-isopropylmalate synthase (2-isopropylmalate synthase); the main isozyme responsible for the first step in the leucine biosynthesis pathway | Multidrug transporter of the major facilitator superfamily, required for resistance to quinidine, barban, cisplatin, and bleomycin | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the ER | Alpha aminoadipate reductase, catalyzes the reduction of alpha-aminoadipate to alpha-aminoadipate 6-semialdehyde, which is the fifth step in biosynthesis of lysine; activation requires posttranslational phosphopantetheinylation by Lys5p | Alcohol dehydrogenase isoenzyme V; involved in ethanol production | Riboflavin synthase; catalyzes the last step of the riboflavin biosynthesis pathway | Beta-isopropylmalate dehydrogenase (IMDH), catalyzes the third step in the leucine biosynthesis pathway | Putative protein of unknown function; YCR023C is not an essential gene | Mitochondrial GTP/GDP transporter, essential for mitochondrial genome maintenance; has a role in mitochondrial iron transport; member of the mitochondrial carrier family | 3-deoxy-D-arabino-heptulosonate-7-phosphate (DAHP) synthase, catalyzes the first step in aromatic amino acid biosynthesis and is feedback-inhibited by phenylalanine or high concentration of tyrosine or tryptophan | Aspartate kinase (L-aspartate 4-P-transferase); cytoplasmic enzyme that catalyzes the first step in the common pathway for methionine and threonine biosynthesis; expression regulated by Gcn4p and the general control of amino acid synthesis | Protein that is processed in the mitochondrion to yield acetylglutamate kinase and N-acetyl-gamma-glutamyl-phosphate reductase, which catalyze the 2nd and 3rd steps in arginine biosynthesis; enzymes form a complex with Arg2p | Anthranilate synthase, catalyzes the initial step of tryptophan biosynthesis, forms multifunctional hetero-oligomeric anthranilate synthase:indole-3-glycerol phosphate synthase enzyme complex with Trp3p | Trans-aconitate methyltransferase, cytosolic enzyme that catalyzes the methyl esterification of 3-isopropylmalate, an intermediate of the leucine biosynthetic pathway, and trans-aconitate, which inhibits the citric acid cycle | Putative cystathionine beta-lyase; involved in copper ion homeostasis and sulfur metabolism; null mutant displays increased levels of spontaneous Rad52 foci; expression induced by nitrogen limitation in a GLN3, GAT1-dependent manner | Putative protein of unknown function | Peroxin required for targeting of peroxisomal matrix proteins containing PTS2; interacts with Pex7p; partially redundant with Pex18p | Mitochondrial protein required for splicing of the group I intron aI5 of the COB pre-mRNA, binds to the RNA to promote splicing; also involved in but not essential for splicing of the COB bI2 intron and the intron in the 21S rRNA gene | Argininosuccinate lyase, catalyzes the final step in the arginine biosynthesis pathway | Protein of unknown function; null mutant is defective in unfolded protein response; possibly involved in a membrane regulation metabolic pathway; member of the PhzF superfamily, though most likely not involved in phenazine production | Cyclin, interacts with Pho85p cyclin-dependent kinase (Cdk), induced by Gcn4p at level of transcription, specifically required for Gcn4p degradation, may be sensor of cellular protein biosynthetic capacity | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the mitochondrion | Histidinol-phosphate aminotransferase, catalyzes the seventh step in histidine biosynthesis; responsive to general control of amino acid biosynthesis; mutations cause histidine auxotrophy and sensitivity to Cu, Co, and Ni salts | Saccharopine dehydrogenase (NAD+, L-lysine-forming), catalyzes the conversion of saccharopine to L-lysine, which is the final step in the lysine biosynthesis pathway | Ornithine carbamoyltransferase (carbamoylphosphate:L-ornithine carbamoyltransferase), catalyzes the sixth step in the biosynthesis of the arginine precursor ornithine | 3-hydroxyanthranilic acid dioxygenase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Large subunit of carbamoyl phosphate synthetase, which catalyzes a step in the synthesis of citrulline, an arginine precursor | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the mitochondria | Cytosolic branched-chain amino acid aminotransferase, homolog of murine ECA39; highly expressed during stationary phase and repressed during logarithmic phase | Bifunctional enzyme exhibiting both indole-3-glycerol-phosphate synthase and anthranilate synthase activities, forms multifunctional hetero-oligomeric anthranilate synthase:indole-3-glycerol phosphate synthase enzyme complex with Trp2p | Cytosolic aspartate aminotransferase, involved in nitrogen metabolism; localizes to peroxisomes in oleate-grown cells | Putative alanine transaminase (glutamic pyruvic transaminase); the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Multidrug efflux pump of the major facilitator superfamily, required for resistance to aminotriazole and 4-nitroquinoline-N-oxide | Mitochondrial ornithine acetyltransferase, catalyzes the fifth step in arginine biosynthesis; also possesses acetylglutamate synthase activity, regenerates acetylglutamate while forming ornithine | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery; cAMP represses YMR086W expression | Protein kinase of the PAK/Ste20 kinase family, required for cell integrity possibly through regulating 1,6-beta-glucan levels in the wall; physically interacts with Cdc31p (centrin), which is a component of the spindle pole body | Protein kinase that stabilizes several plasma membrane amino acid transporters by antagonizing their ubiquitin-mediated degradation | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; putative F-box protein; analysis of integrated high-throughput datasets predicts involvement in ubiquitin-dependent protein catabolism | 1,3-beta-glucanosyltransferase, has similarity to Gas1p; localizes to the cell wall | Arginosuccinate synthetase, catalyzes the formation of L-argininosuccinate from citrulline and L-aspartate in the arginine biosynthesis pathway; potential Cdc28p substrate | Bisphosphate-3'-nucleotidase, involved in salt tolerance and methionine biogenesis; dephosphorylates 3'-phosphoadenosine-5'-phosphate and 3'-phosphoadenosine-5'-phosphosulfate, intermediates of the sulfate assimilation pathway | Protein with similarity to mammalian monocarboxylate permeases, which are involved in transport of monocarboxylic acids across the plasma membrane; mutant is not deficient in monocarboxylate transport | Acetylornithine aminotransferase, catalyzes the fourth step in the biosynthesis of the arginine precursor ornithine | Golgi membrane protein involved in manganese homeostasis; overproduction suppresses the sod1 (copper, zinc superoxide dismutase) null mutation | Origin-binding F-box protein that forms an SCF ubiquitin ligase complex with Skp1p and Cdc53p; plays a role in DNA replication, involved in invasive and pseudohyphal growth | Ornithine transporter of the mitochondrial inner membrane, exports ornithine from mitochondria as part of arginine biosynthesis; human ortholog is associated with hyperammonaemia-hyperornithinaemia-homocitrullinuria (HHH) syndrome | Small subunit of carbamoyl phosphate synthetase, which catalyzes a step in the synthesis of citrulline, an arginine precursor; translationally regulated by an attenuator peptide encoded by YOR302W within the CPA1 mRNA 5'-leader | Iron-regulated transcriptional activator; activates genes involved in intracellular iron use and required for iron homeostasis and resistance to oxidative stress; similar to Aft1p | Protein of unknown function; mobilized into polysomes upon a shift from a fermentable to nonfermentable carbon source; potential Cdc28p substrate |
| 199 | 46 | 458 | 0.822 | 0.634 |  |  | 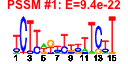 | 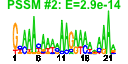 |  | YBR186W YBR231C YBR240C YDL036C YDL077C YDL107W YDL211C YDL241W YDR217C YGR030C YGR096W YIL147C YLR341W YML075C YNL176C YNR077C YOR313C YAR030C YBL106C YBR161W YBR232C YCL008C YDL030W YDL033C YDL105W YDL141W YDR038C YDR039C YDR052C YDR075W YDR314C YDR519W YEL030W YER075C YER095W YFR018C YGL017W YGL063W YGL175C YGR031W YGR068C YGR188C YIR010W YLL013C YNR074C YPR122W | PCH2 SWC5 THI2 PUS9 VAM6 MSS2 RAD9 POP6 TPC1 SLN1 SPO77 HMG1 SPS4 SRO77 CSH1 STP22 PRP9 SLM3 NSE4 BPL1 ENA5 ENA2 DBF4 PPH3 RAD34 FPR2 ECM10 PTP3 RAD51 ATE1 PUS2 SAE2 BUB1 DSN1 PUF3 CPD1 AXL1 | Nucleolar component of the pachytene checkpoint, which prevents chromosome segregation when recombination and chromosome synapsis are defective; also represses meiotic interhomolog recombination in the rDNA | Protein of unknown function, component of the SWR1 complex, which exchanges histone variant H2AZ (Htz1p) for chromatin-bound histone H2A | Zinc finger protein of the Zn(II)2Cys6 type, probable transcriptional activator of thiamine biosynthetic genes | Mitochondrial tRNA pseudouridine synthase involved in pseudouridylation of mitochondrial tRNAs at position 32 | Vacuolar protein that plays a critical role in the tethering steps of vacuolar membrane fusion by facilitating guanine nucleotide exchange on small guanosine triphosphatase Ypt7p | Peripherally bound inner membrane protein of the mitochondrial matrix involved in membrane insertion of C-terminus of Cox2p, interacts genetically and physically with Cox18p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Putative protein of unknown function; YDL241W is not an essential gene | DNA damage-dependent checkpoint protein, required for cell-cycle arrest in G1/S, intra-S, and G2/M; transmits checkpoint signal by activating Rad53p and Chk1p; hyperphosphorylated by Mec1p and Tel1p; potential Cdc28p substrate | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends | Mitochondrial membrane transporter that mediates uptake of the essential cofactor thiamine pyrophosphate (ThPP) into mitochondria; expression appears to be regulated by carbon source; member of the mitochondrial carrier family | Histidine kinase osmosensor that regulates a MAP kinase cascade; transmembrane protein with an intracellular kinase domain that signals to Ypd1p and Ssk1p, thereby forming a phosphorelay system similar to bacterial two-component regulators | Meiosis-specific protein of unknown function, required for spore wall formation during sporulation; dispensable for both nuclear divisions during meiosis | One of two isozymes of HMG-CoA reductase that catalyzes the conversion of HMG-CoA to mevalonate, which is a rate-limiting step in sterol biosynthesis; localizes to the nuclear envelope; overproduction induces the formation of karmellae | Cell cycle-regulated gene of unknown function, promoter bound by Fkh2p | Protein whose expression is induced during sporulation; not required for sporulation; heterologous expression in E. coli induces the SOS response that senses DNA damage | Protein with roles in exocytosis and cation homeostasis; functions in docking and fusion of post-Golgi vesicles with plasma membrane; homolog of Sro7p and Drosophila lethal giant larvae tumor suppressor; interacts with SNARE protein Sec9p | Probable catalytic subunit of a mannosylinositol phosphorylceramide (MIPC) synthase, forms a complex with probable regulatory subunit Csg2p; function in sphingolipid biosynthesis is overlapping with that of Sur1p | Component of the ESCRT-I complex, which is involved in ubiquitin-dependent sorting of proteins into the endosome; homologous to the mouse and human Tsg101 tumor susceptibility gene; mutants exhibit a Class E Vps phenotype | Subunit of the SF3a splicing factor complex, required for spliceosome assembly; acts after the formation of the U1 snRNP-pre-mRNA complex | tRNA-specific 2-thiouridylase, responsible for 2-thiolation of the wobble base of mitochondrial tRNAs; human ortholog is implicated in myoclonus epilepsy associated with ragged red fibers (MERRF) | Nuclear protein that plays a role in the function of the Smc5p-Rhc18p complex | Biotin:apoprotein ligase, covalently modifies proteins with the addition of biotin, required for acetyl-CoA carboxylase (Acc1p) holoenzyme formation | Protein with similarity to P-type ATPase sodium pumps, member of the Na+ efflux ATPase family | P-type ATPase sodium pump, involved in Na+ efflux to allow salt tolerance; likely not involved in Li+ efflux | Regulatory subunit of Cdc7p-Dbf4p kinase complex, required for Cdc7p kinase activity and initiation of DNA replication; phosphorylates the Mcm2-7 family of proteins; cell cycle regulated | Catalytic subunit of an evolutionarily conserved protein phosphatase complex containing Psy2p and the regulatory subunit Psy4p; required for cisplatin resistance; involved in activation of Gln3p | Protein involved in nucleotide excision repair (NER); homologous to RAD4 | Membrane-bound peptidyl-prolyl cis-trans isomerase (PPIase), binds to the drugs FK506 and rapamycin; expression pattern suggests possible involvement in ER protein trafficking | Heat shock protein of the Hsp70 family, localized in mitochondrial nucleoids, plays a role in protein translocation, interacts with Mge1p in an ATP-dependent manner; overexpression induces extensive mitochondrial DNA aggregations | Phosphotyrosine-specific protein phosphatase involved in the inactivation of mitogen-activated protein kinase (MAPK) during osmolarity sensing; dephosporylates Hog1p MAPK and regulates its localization; localized to the cytoplasm | Strand exchange protein, forms a helical filament with DNA that searches for homology; involved in the recombinational repair of double-strand breaks in DNA during vegetative growth and meiosis; homolog of Dmc1p and bacterial RecA protein | Putative protein of unknown function | Arginyl-tRNA-protein transferase, catalyzes post-translational conjugation of arginine to the amino termini of acceptor proteins which are then subject to degradation via the N-end rule pathway | Mitochondrial tRNA:pseudouridine synthase; acts at positions 27 and 28, but not at position 72; efficiently and rapidly targeted to mitochondria, specifically dedicated to mitochondrial tRNA modification | Protein with a role in accurate meiotic and mitotic double-strand break repair; phosphorylated in response to DNA damage and required for normal resistance to DNA-damaging agents | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; YGR068C is not an essential gene. | Protein kinase that forms a complex with Mad1p and Bub3p that is crucial in the checkpoint mechanism required to prevent cell cycle progression into anaphase in the presence of spindle damage, associates with centromere DNA via Skp1p | Essential component of the MIND kinetochore complex (Mtw1p Including Nnf1p-Nsl1p-Dsn1p) which joins kinetochore subunits contacting DNA to those contacting microtubules; important for chromosome segregation | Protein of the mitochondrial outer surface, links the Arp2/3 complex with the mitochore during anterograde mitochondrial movement; also binds to and promotes degradation of mRNAs for select nuclear-encoded mitochondrial proteins | Cyclic nucleotide phosphodiesterase, hydrolyzes ADP-ribose 1'', 2''-cyclic phosphate to ADP-ribose 1''-phosphate; may have a role in tRNA splicing; no detectable phenotype is conferred by null mutation or by overexpression | Haploid specific endoprotease that performs one of two N-terminal cleavages during maturation of a-factor mating pheromone; required for axial budding pattern of haploid cells |
| 96 | 34 | 443 | 0.828 | 0.563 |  |  | 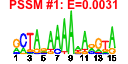 | 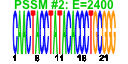 |  | YDL002C YDR088C YDR168W YDR357C YGL091C YGL152C YGL258W YHR146W YIL029C YJL032W YJL131C YJL199C YJR107W YKL061W YLR093C YLR165C YLR211C YLR248W YLR254C YMR255W YNL266W YOL114C YOR097C YOR244W YOR387C YPL159C YPR167C YCL067C YCR039C YDR369C YER061C YMR052W YOL044W YOR111W | NHP10 SLU7 CDC37 NBP35 YGL258W-A CRP1 AIM23 NYV1 PUS5 RCK2 NDL1 GFD1 ESA1 PET20 MET16 HMLALPHA2 XRS2 CEM1 FAR3 PEX15 | Protein related to mammalian high mobility group proteins; likely component of the INO80 complex, which is an ATP-dependent chromatin-remodeling complex | RNA splicing factor, required for ATP-independent portion of 2nd catalytic step of spliceosomal RNA splicing; interacts with Prp18p; contains zinc knuckle domain | Essential Hsp90p co-chaperone; necessary for passage through the START phase of the cell cycle; stabilizes protein kinase nascent chains and participates along with Hsp90p in their folding | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Essential iron-sulfur cluster binding protein localized in the cytoplasm; forms a complex with Cfd1p that is involved in iron-sulfur protein assembly in the cytosol; similar to P-loop NTPases | Putative protein of unknown function | Protein that binds to cruciform DNA structures | Putative protein of unknown function; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO) | Putative protein of unknown function; the authentic non-tagged protein is detected in highly purified mitochondria; null mutant is viable, displays increased frequency of mitochondrial genome loss (petite formation) | Putative protein of unknown function; has sequence or structural similarity to lipases | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endosome | v-SNARE component of the vacuolar SNARE complex involved in vesicle fusion; inhibits ATP-dependent Ca(2+) transport activity of Pmc1p in the vacuolar membrane | Pseudouridine synthase, catalyzes only the formation of pseudouridine (Psi)-2819 in mitochondrial 21S rRNA; not essential for viability | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YLR211C is not an essential gene; ORF contains an intron | Protein kinase involved in the response to oxidative and osmotic stress; identified as suppressor of S. pombe cell cycle checkpoint mutations | Homolog of nuclear distribution factor NudE, NUDEL; interacts with Pac1p and regulates dynein targeting to microtubule plus ends | Coiled-coiled protein of unknown function, identified as a high-copy suppressor of a dbp5 mutation | Putative protein of unknown function with similarity to human ICT1 and prokaryotic factors that may function in translation termination; YOL114C is not an essential gene | Putative protein of unknown function; identified as interacting with Hsp82p in a high-throughput two-hybrid screen; YOR097C is not an essential gene | Histone acetyltransferase catalytic subunit of the native multisubunit complex (NuA4) that acetylates four conserved internal lysines of histone H4 N-terminal tail; required for cell cycle progression | Putative protein of unknown function; regulated by the metal-responsive Aft1p transcription factor; highly inducible in zinc-depleted conditions; localizes to the soluble fraction | Mitochondrial protein, required for respiratory growth under some conditions and for stability of the mitochondrial genome | 3'-phosphoadenylsulfate reductase, reduces 3'-phosphoadenylyl sulfate to adenosine-3',5'-bisphosphate and free sulfite using reduced thioredoxin as cosubstrate, involved in sulfate assimilation and methionine metabolism | Silenced copy of ALPHA2 at HML; homeobox-domain protein that associates with Mcm1p in haploid cells to repress a-specific gene expression and interacts with a1p in diploid cells to repress haploid-specific gene expression | Protein required for DNA repair; component of the Mre11 complex, which is involved in double strand breaks, meiotic recombination, telomere maintenance, and checkpoint signaling | Mitochondrial beta-keto-acyl synthase with possible role in fatty acid synthesis; required for mitochondrial respiration | Protein involved in G1 cell cycle arrest in response to pheromone, in a pathway different from the Far1p-dependent pathway; interacts with Far7p, Far8p, Far9p, Far10p, and Far11p | Phosphorylated tail-anchored type II integral peroxisomal membrane protein required for peroxisome biogenesis, cells lacking Pex15p mislocalize peroxisomal matrix proteins to cytosol, overexpression results in impaired peroxisome assembly |
| 83 | 42 | 433 | 0.829 | 0.580 |  |  | 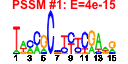 | 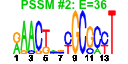 |  | YAR027W YAR028W YDR129C YDR319C YGL006W YHL048W YIR043C YJL094C YJL172W YKL126W YKL157W YKL159C YKL213C YLR028C YLR194C YLR241W YLR414C YMR291W YMR315W YMR316W YNR032W YNR035C YOL016C YOR219C YPL066W YPL067C YPL149W YPL191C YPL270W YPR024W YPR028W YPR198W YDL173W YGR131W YHR028C YHR190W YJR062C YKL094W YMR031C YMR258C YOR059C YOR288C | UIP3 SAC6 YFT2 YGL006W-A COS8 KHA1 CPS1 TOK1 APE2 RCN1 DOA1 ADE16 DIA1 PPG1 ARC35 CMK2 STE13 ATG5 MDL2 YME1 YOP1 SGE1 DAP2 ERG9 NTA1 YJU3 MPD1 | Putative integral membrane protein of unknown function; interacts with Ulp1p at the nuclear periphery; member of DUP240 gene family | Putative integral membrane protein, member of DUP240 gene family; GFP-fusion protein is induced in response to the DNA-damaging agent MMS | Fimbrin, actin-bundling protein; cooperates with Scp1p (calponin/transgelin) in the organization and maintenance of the actin cytoskeleton | Putative protein of unknown function, identified as an ortholog of the highly conserved FIT family of proteins involved in triglyceride droplet biosynthesis; interacts with Sst2p and Hsp82p in high-throughput two-hybrid screens | Putative protein of unknown function; identified by SAGE | Nuclear membrane protein, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins; regulation suggests a potential role in the unfolded protein response | Putative K+/H+ antiporter with a probable role in intracellular cation homeostasis, localized to Golgi vesicles and detected in highly purified mitochondria in high-throughput studies | Vacuolar carboxypeptidase yscS; expression is induced under low-nitrogen conditions | Outward-rectifier potassium channel of the plasma membrane with two pore domains in tandem, each of which forms a functional channel permeable to potassium; carboxy tail functions to prevent inner gate closures; target of K1 toxin | Zinc-dependent metallopeptidase yscII, may have a role in obtaining leucine from dipeptide substrates; sequence coordinates have changed since RT-PCR analysis showed that the adjacent ORF YKL158W comprises the 5' exon of APE2/YKL157W | Protein involved in calcineurin regulation during calcium signaling; has similarity to H. sapiens DSCR1 which is found in the Down Syndrome candidate region | WD repeat protein required for ubiquitin-mediated protein degradation, forms complex with Cdc48p, plays a role in controlling cellular ubiquitin concentration; also promotes efficient NHEJ in postdiauxic/stationary phase | Enzyme of 'de novo' purine biosynthesis containing both 5-aminoimidazole-4-carboxamide ribonucleotide transformylase and inosine monophosphate cyclohydrolase activities, isozyme of Ade17p; ade16 ade17 mutants require adenine and histidine | Structural constituent of the cell wall attached to the plasma membrane by a GPI-anchor; expression is upregulated in response to cell wall stress | Putative protein of unknown function, may be involved in detoxification | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the bud and cytoplasm; Hog1p is required for transcriptional induction in response to cell wall damage; YLR414C is not an essential gene | Putative kinase of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YMR291W is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YMR315W is not an essential gene | Protein of unknown function, involved in invasive and pseudohyphal growth; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Putative serine/threonine protein phosphatase, required for glycogen accumulation; interacts with Tap42p, which binds to and regulates other protein phosphatases | Subunit of the ARP2/3 complex, which is required for the motility and integrity of cortical actin patches; required for cortical localization of calmodulin | Calmodulin-dependent protein kinase; may play a role in stress response, many CA++/calmodulan dependent phosphorylation substrates demonstrated in vitro, amino acid sequence similar to Cmk1p and mammalian Cam Kinase II | Dipeptidyl aminopeptidase, Golgi integral membrane protein that cleaves on the carboxyl side of repeating -X-Ala- sequences, required for maturation of alpha factor, transcription is induced by a-factor | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the bud neck and cytoplasm; null mutant is viable and exhibits growth defect on a non-fermentable (respiratory) carbon source | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YPL067C is not an essential gene | Conserved autophagy-related protein that undergoes conjugation with Atg12p and then associates with Atg16p to form a cytosolic complex essential for autophagosome formation | Putative protein of unknown function; diploid deletion strain exhibits high budding index; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Mitochondrial inner membrane half-type ATP-binding cassette (ABC) transporter | Subunit, with Mgr1p, of the mitochondrial inner membrane i-AAA protease complex, which is responsible for degradation of unfolded or misfolded mitochondrial gene products; mutation causes an elevated rate of mitochondrial turnover | Membrane protein that interacts with Yip1p to mediate membrane traffic; overexpression results in cell death and accumulation of internal cell membranes; regulates vesicular traffic in stressed cells | Plasma membrane multidrug transporter of the major facilitator superfamily, acts as an extrusion permease; partial multicopy suppressor of gal11 mutations | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YDL173W is not an essential gene | Protein of unknown function; expression induced in response to ketoconazole; promoter region contains a sterol regulatory element motif, which has been identified as a Upc2p-binding site | Dipeptidyl aminopeptidase, synthesized as a glycosylated precursor; localizes to the vacuolar membrane; similar to Ste13p | Farnesyl-diphosphate farnesyl transferase (squalene synthase), joins two farnesyl pyrophosphate moieties to form squalene in the sterol biosynthesis pathway | Amidase, removes the amide group from N-terminal asparagine and glutamine residues to generate proteins with N-terminal aspartate and glutamate residues that are targets of ubiquitin-mediated degradation | Serine hydrolase with sequence similarity to monoglyceride lipase (MGL), localizes to lipid particles | Protein of unknown function with similarity to Ykl050cp and Uso1p; the authentic, non-tagged protein is detected in a phosphorylated state in highly purified mitochondria in high-throughput studies; YMR031C is not an essential gene | Protein of unknown function with similarity to F-box proteins; physically interacts with Skp1p; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YMR258C is not an essential gene | hypothetical protein | Member of the protein disulfide isomerase (PDI) family; interacts with and inhibits the chaperone activity of Cne1p; MPD1 overexpression in a pdi1 null mutant suppresses defects in Pdi1p functions such as carboxypeptidase Y maturation |
| 145 | 37 | 432 | 0.830 | 0.627 |  |  | 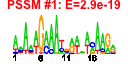 | 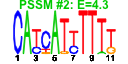 |  | YBL043W YBR054W YDL048C YDR185C YER024W YGL056C YGR138C YHR053C YHR055C YIL121W YIL161W YIR035C YLL027W YLL054C YLR107W YLR375W YMR118C YNL063W YOR302W YOR303W YOR365C YOR377W YPL015C YPR156C YDR403W YDR441C YER046W YIL024C YIL076W YIL119C YJL137C YJL170C YLL057C YLR273C YLR297W YOR339C YPL156C | ECM13 YRO2 STP4 YAT2 SDS23 TPO2 CUP1-1 QDR2 ISA1 REX3 STP3 MTQ1 CPA1 ATF1 HST2 TPO3 DIT1 APT2 SPO73 SEC28 RPI1 GLG2 ASG7 JLP1 PIG1 UBC11 PRM4 | Non-essential protein of unknown function | Putative protein of unknown function; the authentic, non-tagged protein is detected in a phosphorylated state in highly purified mitochondria in high-throughput studies; transcriptionally regulated by Haa1p | Protein containing a Kruppel-type zinc-finger domain; has similarity to Stp1p, Stp2p, and Stp3p | Mitochondrial protein of unknown function; has similarity to Ups1p, which is involved in regulation of alternative topogenesis of the dynamin-related GTPase Mgm1p | Carnitine acetyltransferase; has similarity to Yat1p, which is a carnitine acetyltransferase associated with the mitochondrial outer membrane | One of two S. cerevisiae homologs (Sds23p and Sds24p) of the Schizosaccharomyces pombe Sds23 protein, which genetic studies have implicated in APC/cyclosome regulation | Polyamine transport protein specific for spermine; localizes to the plasma membrane; transcription of TPO2 is regulated by Haa1p; member of the major facilitator superfamily | Metallothionein, binds copper and mediates resistance to high concentrations of copper and cadmium; locus is variably amplified in different strains, with two copies, CUP1-1 and CUP1-2, in the genomic sequence reference strain S288C | Multidrug transporter of the major facilitator superfamily, required for resistance to quinidine, barban, cisplatin, and bleomycin; may have a role in potassium uptake | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; mRNA is enriched in Scp160p-associated mRNPs; YIL161W is a non-essential gene | Putative cytoplasmic protein of unknown function | Mitochondrial matrix protein involved in biogenesis of the iron-sulfur (Fe/S) cluster of Fe/S proteins, isa1 deletion causes loss of mitochondrial DNA and respiratory deficiency; depletion reduces growth on nonfermentable carbon sources | Putative protein of unknown function with similarity to Pip2p, an oleate-specific transcriptional activator of peroxisome proliferation; YLL054C is not an essential gene | RNA exonuclease; required for maturation of the RNA component of RNase MRP; functions redundantly with Rnh70p and Rex2p in processing of U5 snRNA and RNase P RNA; member of RNase D family of exonucleases | Zinc-finger protein of unknown function, possibly involved in pre-tRNA splicing and in uptake of branched-chain amino acids | Protein of unknown function with similarity to succinate dehydrogenase cytochrome b subunit; YMR118C is not an essential gene | S-adenosylmethionine-dependent methyltransferase; methylates translational release factor Mrf1p; similar to E.coli PrmC; is not an essential gene | CPA1 uORF , Arginine attenuator peptide, regulates translation of the CPA1 mRNA | Small subunit of carbamoyl phosphate synthetase, which catalyzes a step in the synthesis of citrulline, an arginine precursor; translationally regulated by an attenuator peptide encoded by YOR302W within the CPA1 mRNA 5'-leader | Putative protein of unknown function; YOR365C is not an essential protein | Alcohol acetyltransferase with potential roles in lipid and sterol metabolism; responsible for the major part of volatile acetate ester production during fermentation | Cytoplasmic member of the silencing information regulator 2 (Sir2) family of NAD(+)-dependent protein deacetylases; modulates nucleolar (rDNA) and telomeric silencing; possesses NAD(+)-dependent histone deacetylase activity in vitro | Polyamine transport protein specific for spermine; localizes to the plasma membrane; member of the major facilitator superfamily | Sporulation-specific enzyme required for spore wall maturation, involved in the production of a soluble LL-dityrosine-containing precursor of the spore wall; transcripts accumulate at the time of prospore enclosure | Apparent pseudogene, not transcribed or translated under normal conditions; encodes a protein with similarity to adenine phosphoribosyltransferase, but artificially expressed protein exhibits no enzymatic activity | Meiosis-specific protein of unknown function, required for spore wall formation during sporulation; dispensible for both nuclear divisions during meiosis | Putative protein of unknown function; non-essential gene; expression directly regulated by the metabolic and meiotic transcriptional regulator Ume6p | Epsilon-COP subunit of the coatomer; regulates retrograde Golgi-to-ER protein traffic; stabilizes Cop1p, the alpha-COP and the coatomer complex; non-essential for cell growth | Putative transcriptional regulator; overexpression suppresses the heat shock sensitivity of wild-type RAS2 overexpression and also suppresses the cell lysis defect of an mpk1 mutation | Self-glucosylating initiator of glycogen synthesis, also glucosylates n-dodecyl-beta-D-maltoside; similar to mammalian glycogenin | Protein that regulates signaling from a G protein beta subunit Ste4p and its relocalization within the cell; specific to a-cells and induced by alpha-factor | Fe(II)-dependent sulfonate/alpha-ketoglutarate dioxygenase, involved in sulfonate catabolism for use as a sulfur source, contains sequence that closely resembles a J domain (typified by the E. coli DnaJ protein) | Putative targeting subunit for the type-1 protein phosphatase Glc7p that tethers it to the Gsy2p glycogen synthase | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole; YLR297W is not an essential gene | Ubiquitin-conjugating enzyme most similar in sequence to Xenopus ubiquitin-conjugating enzyme E2-C, but not a true functional homolog of this E2; unlike E2-C, not required for the degradation of mitotic cyclin Clb2 | Pheromone-regulated protein, predicted to have 1 transmembrane segment; transcriptionally regulated by Ste12p during mating and by Cat8p during the diauxic shift |
| 60 | 54 | 443 | 0.906 | 0.665 |  |  | 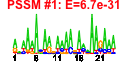 | 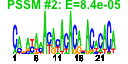 |  | YBL030C YBR153W YBR159W YBR296C YCL025C YDR046C YDR134C YDR342C YDR434W YDR441C YDR492W YDR497C YDR510W YEL047C YER003C YER073W YGR189C YGR218W YGR279C YGR282C YHR092C YHR123W YIL051C YIL119C YJL079C YJL159W YJR048W YKL008C YKL096W YKR093W YLR110C YML106W YML123C YMR016C YMR088C YMR145C YMR272C YNL292W YOR092W YPL057C YPL061W YPL092W YPR183W YDR133C YDR345C YEL046C YFL031W YGL215W YHR175W YJL097W YKL163W YLR332W YNL142W YOR045W | PET9 RIB7 IFA38 PHO89 AGP1 BAP3 HXT7 GPI17 APT2 IZH1 ITR1 SMT3 PMI40 ALD5 CRH1 CRM1 SCW4 BGL2 HXT4 EPT1 MMF1 RPI1 PRY1 HSP150 CYC1 MTH1 CWP2 PTR2 CCW12 URA5 PHO84 SOK2 VBA1 NDE1 SCS7 PUS4 ECM3 SUR1 ALD6 SSU1 DPM1 HXT3 GLY1 HAC1 CLG1 CTR2 PHS1 PIR3 MID2 MEP2 TOM6 | Major ADP/ATP carrier of the mitochondrial inner membrane, exchanges cytosolic ADP for mitochondrially synthesized ATP; phosphorylated; required for viability in many common lab strains carrying a mutation in the polymorphic SAL1 gene | Diaminohydroxyphoshoribosylaminopyrimidine deaminase; catalyzes the second step of the riboflavin biosynthesis pathway | Microsomal beta-keto-reductase; contains oleate response element (ORE) sequence in the promoter region; mutants exhibit reduced VLCFA synthesis, accumulate high levels of dihydrosphingosine, phytosphingosine and medium-chain ceramides | Na+/Pi cotransporter, active in early growth phase; similar to phosphate transporters of Neurospora crassa; transcription regulated by inorganic phosphate concentrations and Pho4p | Low-affinity amino acid permease with broad substrate range, involved in uptake of asparagine, glutamine, and other amino acids; expression is regulated by the SPS plasma membrane amino acid sensor system (Ssy1p-Ptr3p-Ssy5p) | Amino acid permease involved in the uptake of cysteine, leucine, isoleucine and valine | High-affinity glucose transporter of the major facilitator superfamily, nearly identical to Hxt6p, expressed at high basal levels relative to other HXTs, expression repressed by high glucose levels | Transmembrane protein subunit of the glycosylphosphatidylinositol transamidase complex that adds GPIs to newly synthesized proteins; human PIG-Sp homolog | Apparent pseudogene, not transcribed or translated under normal conditions; encodes a protein with similarity to adenine phosphoribosyltransferase, but artificially expressed protein exhibits no enzymatic activity | Membrane protein involved in zinc metabolism, member of the four-protein IZH family; transcription is regulated directly by Zap1p, expression induced by zinc deficiency and fatty acids; deletion increases sensitivity to elevated zinc | Myo-inositol transporter with strong similarity to the minor myo-inositol transporter Itr2p, member of the sugar transporter superfamily; expression is repressed by inositol and choline via Opi1p and derepressed via Ino2p and Ino4p | Ubiquitin-like protein of the SUMO family, conjugated to lysine residues of target proteins; regulates chromatid cohesion, chromosome segregation, APC-mediated proteolysis, DNA replication and septin ring dynamics | Soluble fumarate reductase, required with isoenzyme Osm1p for anaerobic growth; may interact with ribosomes, based on co-purification experiments; authentic, non-tagged protein is detected in purified mitochondria in high-throughput studies | Mannose-6-phosphate isomerase, catalyzes the interconversion of fructose-6-P and mannose-6-P; required for early steps in protein mannosylation | Mitochondrial aldehyde dehydrogenase, involved in regulation or biosynthesis of electron transport chain components and acetate formation; activated by K+; utilizes NADP+ as the preferred coenzyme; constitutively expressed | Cell wall protein that functions in the transfer of chitin to beta(1-6)glucan, putative chitin transglycosidase | Major karyopherin, involved in export of proteins, RNAs, and ribosomal subunits from the nucleus | Cell wall protein with similarity to glucanases; scw4 scw10 double mutants exhibit defects in mating | Endo-beta-1,3-glucanase, major protein of the cell wall, involved in cell wall maintenance | High-affinity glucose transporter of the major facilitator superfamily, expression is induced by low levels of glucose and repressed by high levels of glucose | sn-1,2-diacylglycerol ethanolamine- and cholinephosphotranferase; not essential for viability | Mitochondrial protein involved in maintenance of the mitochondrial genome | Putative transcriptional regulator; overexpression suppresses the heat shock sensitivity of wild-type RAS2 overexpression and also suppresses the cell lysis defect of an mpk1 mutation | Protein of unknown function, has similarity to Pry2p and Pry3p and to the plant PR-1 class of pathogen related proteins | O-mannosylated heat shock protein that is secreted and covalently attached to the cell wall via beta-1,3-glucan and disulfide bridges; required for cell wall stability; induced by heat shock, oxidative stress, and nitrogen limitation | Cytochrome c, isoform 1; electron carrier of the mitochondrial intermembrane space that transfers electrons from ubiquinone-cytochrome c oxidoreductase to cytochrome c oxidase during cellular respiration | Negative regulator of the glucose-sensing signal transduction pathway, required for repression of transcription by Rgt1p; interacts with Rgt1p and the Snf3p and Rgt2p glucose sensors; phosphorylated by Yck1p, triggering Mth1p degradation | Covalently linked cell wall mannoprotein, major constituent of the cell wall; plays a role in stabilizing the cell wall; involved in low pH resistance; precursor is GPI-anchored | Integral membrane peptide transporter, mediates transport of di- and tri-peptides; conserved protein that contains 12 transmembrane domains; PTR2 expression is regulated by the N-end rule pathway via repression by Cup9p | Cell wall mannoprotein, mutants are defective in mating and agglutination, expression is downregulated by alpha-factor | One of two orotate phosphoribosyltransferase isozymes (see also URA10) that catalyze the fifth enzymatic step in de novo biosynthesis of pyrimidines, converting orotate into orotidine-5'-phosphate | High-affinity inorganic phosphate (Pi) transporter and low-affinity manganese transporter; regulated by Pho4p and Spt7p; mutation confers resistance to arsenate; exit from the ER during maturation requires Pho86p | Nuclear protein that plays a regulatory role in the cyclic AMP (cAMP)-dependent protein kinase (PKA) signal transduction pathway; negatively regulates pseudohyphal differentiation; homologous to several transcription factors | Permease of basic amino acids in the vacuolar membrane | Mitochondrial external NADH dehydrogenase, a type II NAD(P)H:quinone oxidoreductase that catalyzes the oxidation of cytosolic NADH; Nde1p and Nde2p provide cytosolic NADH to the mitochondrial respiratory chain | Sphingolipid alpha-hydroxylase, functions in the alpha-hydroxylation of sphingolipid-associated very long chain fatty acids, has both cytochrome b5-like and hydroxylase/desaturase domains, not essential for growth | Pseudouridine synthase, catalyzes only the formation of pseudouridine-55 (Psi55), a highly conserved tRNA modification, in mitochondrial and cytoplasmic tRNAs; PUS4 overexpression leads to translational derepression of GCN4 (Gcd- phenotype) | Non-essential protein of unknown function | Probable catalytic subunit of a mannosylinositol phosphorylceramide (MIPC) synthase, forms a complex with probable regulatory subunit Csg2p; function in sphingolipid biosynthesis is overlapping with that of Csh1p | Cytosolic aldehyde dehydrogenase, activated by Mg2+ and utilizes NADP+ as the preferred coenzyme; required for conversion of acetaldehyde to acetate; constitutively expressed; locates to the mitochondrial outer surface upon oxidative stress | Plasma membrane sulfite pump involved in sulfite metabolism and required for efficient sulfite efflux; major facilitator superfamily protein | Dolichol phosphate mannose (Dol-P-Man) synthase of the ER membrane, catalyzes the formation of Dol-P-Man from Dol-P and GDP-Man; required for glycosyl phosphatidylinositol membrane anchoring, O mannosylation, and protein glycosylation | Low affinity glucose transporter of the major facilitator superfamily, expression is induced in low or high glucose conditions | Threonine aldolase, catalyzes the cleavage of L-allo-threonine and L-threonine to glycine; involved in glycine biosynthesis | bZIP transcription factor (ATF/CREB1 homolog) that regulates the unfolded protein response, via UPRE binding, and membrane biogenesis; ER stress-induced splicing pathway utilizing Ire1p, Trl1p and Ada5p facilitates efficient Hac1p synthesis | Cyclin-like protein that interacts with Pho85p; has sequence similarity to G1 cyclins PCL1 and PCL2 | Putative low-affinity copper transporter of the vacuolar membrane; mutation confers resistance to toxic copper concentrations, while overexpression confers resistance to copper starvation | Protein of unknown function; homolog of mammalian PTPLA; involved in sphingolipid biosynthesis, protein trafficking; required for cell viability | O-glycosylated covalently-bound cell wall protein required for cell wall stability; expression is cell cycle regulated, peaking in M/G1 and also subject to regulation by the cell integrity pathway | O-glycosylated plasma membrane protein that acts as a sensor for cell wall integrity signaling and activates the pathway; interacts with Rom2p, a guanine nucleotide exchange factor for Rho1p, and with cell integrity pathway protein Zeo1p | Ammonium permease involved in regulation of pseudohyphal growth; belongs to a ubiquitous family of cytoplasmic membrane proteins that transport only ammonium (NH4+); expression is under the nitrogen catabolite repression regulation | Component of the TOM (translocase of outer membrane) complex responsible for recognition and initial import steps for all mitochondrially directed proteins; promotes assembly and stability of the TOM complex |
| 54 | 41 | 434 | 0.927 | 0.613 |  |  | 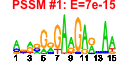 | 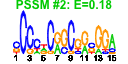 |  | YAL017W YBR083W YBR262C YBR286W YCR075C YCR079W YDL046W YDL059C YDL142C YDL173W YEL038W YER020W YER044C YER058W YER061C YER083C YFR047C YGL035C YGL181W YGR044C YGR086C YGR130C YGR174C YGR230W YGR244C YGR284C YHR028C YHR033W YHR190W YKL163W YLR079W YLR090W YML110C YNL098C YOL109W YOR075W YOR317W YDR100W YDR231C YJL052W YJL059W | PSK1 TEC1 AIM5 APE3 ERS1 PTC6 NPC2 RAD59 CRD1 UTR4 GPA2 ERG28 PET117 CEM1 GET2 BNA6 MIG1 GTS1 RME1 PIL1 CBP4 BNS1 LSC2 ERV29 DAP2 ERG9 PIR3 SIC1 XDJ1 COQ5 RAS2 ZEO1 UFE1 FAA1 TVP15 COX20 TDH1 YHC3 | One of two (see also PSK2) PAS domain containing S/T protein kinases; coordinately regulates protein synthesis and carbohydrate metabolism and storage in response to a unknown metabolite that reflects nutritional status | Transcription factor required for full Ty1 epxression, Ty1-mediated gene activation, and haploid invasive and diploid pseudohyphal growth; TEA/ATTS DNA-binding domain family member | Protein of unknown function; non-tagged protein is detected in purified mitochondria in high-throughput studies; null mutant displays decreased frequency of mitochondrial genome loss and reduced growth rate in minimal glycerol media | Vacuolar aminopeptidase Y, processed to mature form by Prb1p | Protein with similarity to human cystinosin, which is a H(+)-driven transporter involved in L-cystine export from lysosomes and implicated in the disease cystinosis; contains seven transmembrane domains | Phosphoprotein phosphatase type 2C similar to mammalian PP1Ks; involved in mitophagy; localized to mitochondrial inner membrane space; null mutant is sensitive to rapamycin | Functional homolog of human NPC2/He1, which is a cholesterol-binding protein whose deficiency causes Niemann-Pick type C2 disease involving retention of cholesterol in lysosomes | Protein involved in the repair of double-strand breaks in DNA during vegetative growth via recombination and single-strand annealing; anneals complementary single-stranded DNA; homologous to Rad52p | Cardiolipin synthase; produces cardiolipin, which is an important constituent of mitochondrial membranes; required for normal mitochondrial membrane potential and function | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YDL173W is not an essential gene | Protein of unknown function with sequence similarity to 2,3-diketo-5-methylthiopentyl-1-phosphate enolase-phosphatases, found in both the cytoplasm and nucleus | Nucleotide binding alpha subunit of the heterotrimeric G protein that interacts with the receptor Gpr1p, has signaling role in response to nutrients; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | Endoplasmic reticulum membrane protein, may facilitate protein-protein interactions between the Erg26p dehydrogenase and the Erg27p 3-ketoreductase and/or tether these enzymes to the ER, also interacts with Erg6p | Protein required for assembly of cytochrome c oxidase | Mitochondrial beta-keto-acyl synthase with possible role in fatty acid synthesis; required for mitochondrial respiration | Subunit of the GET complex; required for meiotic nuclear division and for the retrieval of HDEL proteins from the Golgi to the ER in an ERD2 dependent fashion; may be involved in cell wall function | Quinolinate phosphoribosyl transferase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Transcription factor involved in glucose repression; sequence specific DNA binding protein containing two Cys2His2 zinc finger motifs; regulated by the SNF1 kinase and the GLC7 phosphatase | Protein containing a zinc-finger in the N-terminus and a long Gln-rich region in the C-terminus; regulates ultradian rhythm, cell size, cell cycle, lifespan, sporulation, heat tolerance, and multidrug transport | Zinc finger protein involved in control of meiosis; prevents meiosis by repressing IME1 expression and promotes mitosis by activating CLN2 expression; directly repressed by a1-a2 regulator; mediates cell type control of sporulation | Primary component of eisosomes, which are large immobile cell cortex structures associated with endocytosis; null mutants show activation of Pkc1p/Ypk1p stress resistance pathways; detected in phosphorylated state in mitochondria | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; specifically phosphorylated in vitro by mammalian diphosphoinositol pentakisphosphate (IP7) | Mitochondrial protein required for assembly of ubiquinol cytochrome-c reductase complex (cytochrome bc1 complex); interacts with Cbp3p and function is partially redundant with that of Cbp3p | Protein with some similarity to Spo12p; overexpression bypasses need for Spo12p, but not required for meiosis | Beta subunit of succinyl-CoA ligase, which is a mitochondrial enzyme of the TCA cycle that catalyzes the nucleotide-dependent conversion of succinyl-CoA to succinate | Protein localized to COPII-coated vesicles, involved in vesicle formation and incorporation of specific secretory cargo | Dipeptidyl aminopeptidase, synthesized as a glycosylated precursor; localizes to the vacuolar membrane; similar to Ste13p | Putative protein of unknown function; epitope-tagged protein localizes to the cytoplasm | Farnesyl-diphosphate farnesyl transferase (squalene synthase), joins two farnesyl pyrophosphate moieties to form squalene in the sterol biosynthesis pathway | O-glycosylated covalently-bound cell wall protein required for cell wall stability; expression is cell cycle regulated, peaking in M/G1 and also subject to regulation by the cell integrity pathway | Inhibitor of Cdc28-Clb kinase complexes that controls G1/S phase transition, preventing premature S phase and ensuring genomic integrity; phosphorylation targets Sic1p for SCF(CDC4)-dependent turnover; functional homolog of mammalian Kip1 | Putative chaperone, homolog of E. coli DnaJ, closely related to Ydj1p; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | 2-hexaprenyl-6-methoxy-1,4-benzoquinone methyltransferase, involved in ubiquinone (Coenzyme Q) biosynthesis; localizes to the matrix face of the mitochondrial inner membrane in a large complex with other ubiquinone biosynthetic enzymes | GTP-binding protein that regulates the nitrogen starvation response, sporulation, and filamentous growth; farnesylation and palmitoylation required for activity and localization to plasma membrane; homolog of mammalian Ras proto-oncogenes | Peripheral membrane protein of the plasma membrane that interacts with Mid2p; regulates the cell integrity pathway mediated by Pkc1p and Slt2p; the authentic protein is detected in a phosphorylated state in highly purified mitochondria | t-SNARE required for ER membrane fusion and vesicular traffic, integral membrane protein that constitutes with Sec20p and Use1p the trimeric acceptor for R/v-SNAREs on Golgi-derived vesicles at the ER; part of Dsl1p complex | Long chain fatty acyl-CoA synthetase with a preference for C12:0-C16:0 fatty acids; involved in the activation of imported fatty acids; localized to both lipid particles and mitochondrial outer membrane; essential for stationary phase | Integral membrane protein localized to late Golgi vesicles along with the v-SNARE Tlg2p | Mitochondrial inner membrane protein, required for proteolytic processing of Cox2p and its assembly into cytochrome c oxidase | Glyceraldehyde-3-phosphate dehydrogenase, isozyme 1, involved in glycolysis and gluconeogenesis; tetramer that catalyzes the reaction of glyceraldehyde-3-phosphate to 1,3 bis-phosphoglycerate; detected in the cytoplasm and cell-wall | Vacuolar membrane protein involved in the ATP-dependent transport of arginine into the vacuole and possibly in balancing ion homeostasis; homolog of human CLN3 involved in Batten disease (juvenile onset neuronal ceroid lipofuscinosis) |
| 208 | 46 | 455 | 0.932 | 0.573 |  |  |  | 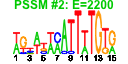 |  | YDR543C YIL150C YIR040C YKR033C YLR135W YNL018C YNL034W YOR298W YOR343C YCL065W YDL028C YDR401W YEL008W YER187W YFL063W YGL177W YIL032C YIL037C YIL054W YKL177W YKL178C YKL221W YKL223W YKL225W YKR034W YKR040C YLR111W YLR235C YLR255C YLR307W YML084W YMR122C YMR326C YNL014W YNL143C YNL318C YNR063W YNR077C YOL085C YOR066W YOR235W YOR313C YOR379C YPL025C YPL275W YPR064W | MCM10 SLX4 MUM3 MPS1 PRM2 STE3 MCH2 DAL80 CDA1 HEF3 HXT14 MSA1 SPS4 | Essential chromatin-associated protein involved in the initiation of DNA replication; required for the association of the MCM2-7 complex with replication origins | Subunit of a complex, with Slx1p, that hydrolyzes 5' branches from duplex DNA in response to stalled or converging replication forks; function overlaps with that of Sgs1p-Top3p | Putative protein of unknown function | Putative protein of unknown function; YNL034W is not an essential gene | Protein of unknown function involved in the organization of the outer spore wall layers; has similarity to the tafazzins superfamily of acyltransferases | Dual-specificity kinase required for spindle pole body (SPB) duplication and spindle checkpoint function; substrates include SPB proteins Spc42p, Spc110p, and Spc98p, mitotic exit network protein Mob1p, and checkpoint protein Mad1p | Putative protein of unknown function; induced in respiratory-deficient cells | Pheromone-regulated protein, predicted to have 4 transmembrane segments and a coiled coil domain; regulated by Ste12p | Receptor for a factor receptor, transcribed in alpha cells and required for mating by alpha cells, couples to MAP kinase cascade to mediate pheromone response; ligand bound receptors are endocytosed and recycled to the plasma membrane; GPCR | Protein with similarity to mammalian monocarboxylate permeases, which are involved in transport of monocarboxylic acids across the plasma membrane; mutant is not deficient in monocarboxylate transport | Negative regulator of genes in multiple nitrogen degradation pathways; expression is regulated by nitrogen levels and by Gln3p; member of the GATA-binding family, forms homodimers and heterodimers with Deh1p | Chitin deacetylase, together with Cda2p involved in the biosynthesis ascospore wall component, chitosan; required for proper rigidity of the ascospore wall | Translational elongation factor EF-3; paralog of YEF3 and member of the ABC superfamily; stimulates EF-1 alpha-dependent binding of aminoacyl-tRNA by the ribosome; normally expressed in zinc deficient cells | Protein with similarity to hexose transporter family members, expression is induced in low glucose and repressed in high glucose; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative zinc-cluster protein of unknown function | Activator of G1-specific transcription factors, MBF and SBF, that regulates both the timing of G1-specific gene transcription, and cell cycle initiation; potential Cdc28p substrate | Protein whose expression is induced during sporulation; not required for sporulation; heterologous expression in E. coli induces the SOS response that senses DNA damage |
| 31 | 39 | 423 | 0.939 | 0.581 |  |  | 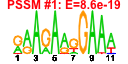 | 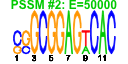 |  | YAL038W YAL044C YBR175W YBR238C YCL036W YCR084C YDL212W YDL228C YDL229W YDR194C YDR233C YDR354W YDR404C YEL033W YEL034W YER056C YER091C YGL077C YGR061C YGR063C YGR175C YGR191W YHR007C YJL198W YJR015W YKL060C YKR080W YLR058C YMR120C YMR243C YMR303C YNL111C YNL220W YOR128C YPR004C YBR220C YDR119W YIL009W YLR153C | CDC19 GCV3 SWD3 GFD2 TUP1 SHR3 SSB1 MSS116 RTN1 TRP4 RPB7 HYP2 FCY2 MET6 HNM1 ADE6 SPT4 ERG1 HIP1 ERG11 PHO90 FBA1 MTD1 SHM2 ADE17 ZRC1 ADH2 CYB5 ADE12 ADE2 AIM45 FAA3 ACS2 | Pyruvate kinase, functions as a homotetramer in glycolysis to convert phosphoenolpyruvate to pyruvate, the input for aerobic (TCA cycle) or anaerobic (glucose fermentation) respiration | H subunit of the mitochondrial glycine decarboxylase complex, required for the catabolism of glycine to 5,10-methylene-THF; expression is regulated by levels of levels of 5,10-methylene-THF in the cytoplasm | Essential subunit of the COMPASS (Set1C) complex, which methylates histone H3 on lysine 4 and is required in transcriptional silencing near telomeres; WD40 beta propeller superfamily member and ortholog of mammalian WDR5 | Mitochondrial membrane protein with similarity to Rmd9p; not required for respiratory growth but causes a synthetic respiratory defect in combination with rmd9 mutations; transcriptionally up-regulated by TOR; deletion increases life span | Protein of unknown function, identified as a high-copy suppressor of a dbp5 mutation | General repressor of transcription, forms complex with Cyc8p, involved in the establishment of repressive chromatin structure through interactions with histones H3 and H4, appears to enhance expression of some genes | Endoplasmic reticulum packaging chaperone, required for incorporation of amino acid permeases into COPII coated vesicles for transport to the cell surface | Cytoplasmic ATPase that is a ribosome-associated molecular chaperone, functions with J-protein partner Zuo1p; may be involved in folding of newly-made polypeptide chains; member of the HSP70 family; interacts with phosphatase subunit Reg1p | DEAD-box protein required for efficient splicing of mitochondrial Group I and II introns; non-polar RNA helicase that also facilities strand annealing | ER membrane protein that interacts with exocyst subunit Sec6p and with Yip3p; also interacts with Sbh1p; null mutant has an altered (mostly cisternal) ER morphology; member of the RTNLA (reticulon-like A) subfamily | Anthranilate phosphoribosyl transferase of the tryptophan biosynthetic pathway, catalyzes the phosphoribosylation of anthranilate, subject to the general control system of amino acid biosynthesis | RNA polymerase II subunit B16; forms two subunit dissociable complex with Rpb4p | Translation initiation factor eIF-5A, promotes formation of the first peptide bond; similar to and functionally redundant with Anb1p; possible role in translation elongation; undergoes an essential hypusination modification | Purine-cytosine permease, mediates purine (adenine, guanine, and hypoxanthine) and cytosine accumulation | Cobalamin-independent methionine synthase, involved in amino acid biosynthesis; requires a minimum of two glutamates on the methyltetrahydrofolate substrate, similar to bacterial metE homologs | Choline/ethanolamine transporter (permease) that also controls the uptake of nitrogen mustard; expression is co-regulated with phospholipid biosynthetic genes and negatively regulated by choline and myo-inositol | Formylglycinamidine-ribonucleotide (FGAM)-synthetase, catalyzes a step in the 'de novo' purine nucleotide biosynthetic pathway | Protein that forms a complex with Spt5p and mediates both activation and inhibition of transcription elongation, and plays a role in pre-mRNA processing; in addition, Spt4p is involved in kinetochore function and gene silencing | Squalene epoxidase, catalyzes the epoxidation of squalene to 2,3-oxidosqualene; plays an essential role in the ergosterol-biosynthesis pathway and is the specific target of the antifungal drug terbinafine | High-affinity histidine permease, also involved in the transport of manganese ions | Lanosterol 14-alpha-demethylase, catalyzes the C-14 demethylation of lanosterol to form 4,4''-dimethyl cholesta-8,14,24-triene-3-beta-ol in the ergosterol biosynthesis pathway; member of the cytochrome P450 family | Low-affinity phosphate transporter; deletion of pho84, pho87, pho89, pho90, and pho91 causes synthetic lethality; transcription independent of Pi and Pho4p activity; overexpression results in vigorous growth | Putative protein of unknown function; localizes to the endoplasmic reticulum and cytoplasm; predicted to encode a membrane transporter based on phylogenetic analysis; YJR015W is a non-essential gene | Fructose 1,6-bisphosphate aldolase, required for glycolysis and gluconeogenesis; catalyzes conversion of fructose 1,6 bisphosphate to glyceraldehyde-3-P and dihydroxyacetone-P; locates to mitochondrial outer surface upon oxidative stress | NAD-dependent 5,10-methylenetetrahydrafolate dehydrogenase, plays a catalytic role in oxidation of cytoplasmic one-carbon units; expression is regulated by Bas1p and Bas2p, repressed by adenine, and may be induced by inositol and choline | Cytosolic serine hydroxymethyltransferase, converts serine to glycine plus 5,10 methylenetetrahydrofolate; major isoform involved in generating precursors for purine, pyrimidine, amino acid, and lipid biosynthesis | Enzyme of 'de novo' purine biosynthesis containing both 5-aminoimidazole-4-carboxamide ribonucleotide transformylase and inosine monophosphate cyclohydrolase activities, isozyme of Ade16p; ade16 ade17 mutants require adenine and histidine | Vacuolar membrane zinc transporter, transports zinc from the cytosol into the vacuole for storage; also has a role in resistance to zinc shock resulting from a sudden influx of zinc into the cytoplasm | Glucose-repressible alcohol dehydrogenase II, catalyzes the conversion of ethanol to acetaldehyde; involved in the production of certain carboxylate esters; regulated by ADR1 | Cytochrome b5, involved in the sterol and lipid biosynthesis pathways; required for sterol C5-6 and fatty acid desaturation | Adenylosuccinate synthase, catalyzes the first step in synthesis of adenosine monophosphate from inosine 5'monophosphate during purine nucleotide biosynthesis; exhibits binding to single-stranded autonomously replicating (ARS) core sequence | Phosphoribosylaminoimidazole carboxylase, catalyzes a step in the 'de novo' purine nucleotide biosynthetic pathway; red pigment accumulates in mutant cells deprived of adenine | Protein with similarity to mammalian electron transfer flavoprotein complex subunit ETF-alpha; interacts with yeast frataxin Yfh1p; null mutant displays increased frequency of mitochondrial genome loss (petite formation) | Putative protein of unknown function; YBR220C is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to vacuolar membrane; YDR119W is not an essential gene | Long chain fatty acyl-CoA synthetase, has a preference for C16 and C18 fatty acids; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | Acetyl-coA synthetase isoform which, along with Acs1p, is the nuclear source of acetyl-coA for histone acetylation; mutants affect global transcription; required for growth on glucose; expressed under anaerobic conditions |
| 228 | 45 | 444 | 0.945 | 0.613 |  |  | 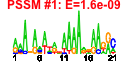 | 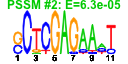 |  | YDL230W YDR452W YHL027W YHR002W YIR034C YJL107C YKL019W YLR121C YNL036W YAL053W YBR004C YBR071W YBR182C YCL021W YER144C YFR044C YGL038C YGL157W YGL179C YGR138C YHR030C YHR043C YJL108C YJL159W YKL104C YKL126W YLL028W YLR042C YLR099C YLR120C YLR137W YLR194C YLR414C YMR238W YMR316W YNL192W YOL011W YOL105C YOR137C YOR385W YPL088W YPL089C YPL221W YPR156C YPR198W | PTP1 PPN1 RIM1 LEU5 LYS1 RAM2 YPS3 NCE103 FLC2 GPI18 SMP1 YCL021W-A UBP5 DUG1 OCH1 TOS3 TPO2 SYN8 DOG2 PRM10 HSP150 GFA1 TOK1 TPO1 ICT1 YAP3 DFG5 DIA1 CHS1 PLB3 WSC3 SIA1 RLM1 FLC1 TPO3 SGE1 | Phosphotyrosine-specific protein phosphatase that dephosphorylates a broad range of substrates in vivo, including Fpr3p; localized to the cytoplasm and the mitochondria | Vacuolar endopolyphosphatase with a role in phosphate metabolism; functions as a homodimer | Single-stranded DNA-binding protein essential for mitochondrial genome maintenance; involved in mitochondrial DNA replication | Mitochondrial carrier protein involved in the accumulation of CoA in the mitochondrial matrix; homolog of human Graves disease protein; does not encode an isozyme of Leu4p, as first hypothesized | Saccharopine dehydrogenase (NAD+, L-lysine-forming), catalyzes the conversion of saccharopine to L-lysine, which is the final step in the lysine biosynthesis pathway | Putative protein of unknown function; expression is induced by activation of the HOG1 mitogen-activated signaling pathway and this induction is Hog1p/Pbs2p dependent; YJL107C and adjacent ORF, YJL108C are merged in related fungi | Alpha subunit of both the farnesyltransferase and type I geranylgeranyltransferase that catalyze prenylation of proteins containing a CAAX consensus motif; essential protein required for membrane localization of Ras proteins and a-factor | Aspartic protease, attached to the plasma membrane via a glycosylphosphatidylinositol (GPI) anchor | Carbonic anhydrase; poorly transcribed under aerobic conditions and at an undetectable level under anaerobic conditions; involved in non-classical protein export pathway | Putative FAD transporter; required for uptake of FAD into endoplasmic reticulum; involved in cell wall maintenance | Functional ortholog of human PIG-V, which is a mannosyltransferase that transfers the second mannose in glycosylphosphatidylinositol biosynthesis; the authentic, non-tagged protein was localized to mitochondria | Putative protein of unknown function; (GFP)-fusion and epitope-tagged proteins localize to the cytoplasm; mRNA expression may be regulated by the cell cycle and/or cell wall stress | Putative transcription factor involved in regulating the response to osmotic stress; member of the MADS-box family of transcription factors | Putative protein of unknown function | Putative ubiquitin-specific protease, closest paralog of Doa4p but has no functional overlap; concentrates at the bud neck | Probable di- and tri-peptidase; forms a complex with Dug2p and Dug3p to degrade glutathione (GSH) and other peptides containing a gamma-glu-X bond in an alternative pathway to GSH degradation by gamma-glutamyl transpeptidase (Ecm38p) | Mannosyltransferase of the cis-Golgi apparatus, initiates the polymannose outer chain elongation of N-linked oligosaccharides of glycoproteins | Oxidoreductase, catalyzes NADPH-dependent reduction of the bicyclic diketone bicyclo[2.2.2]octane-2,6-dione (BCO2,6D) to the chiral ketoalcohol (1R,4S,6S)-6-hydroxybicyclo[2.2.2]octane-2-one (BCO2one6ol) | Protein kinase, related to and functionally redundant with Elm1p and Sak1p for the phosphorylation and activation of Snf1p; functionally orthologous to LKB1, a mammalian kinase associated with Peutz-Jeghers cancer-susceptibility syndrome | Polyamine transport protein specific for spermine; localizes to the plasma membrane; transcription of TPO2 is regulated by Haa1p; member of the major facilitator superfamily | Endosomal SNARE related to mammalian syntaxin 8 | 2-deoxyglucose-6-phosphate phosphatase, member of a family of low molecular weight phosphatases, similar to Dog1p, induced by oxidative and osmotic stress, confers 2-deoxyglucose resistance when overexpressed | Pheromone-regulated protein, predicted to have 5 transmembrane segments | O-mannosylated heat shock protein that is secreted and covalently attached to the cell wall via beta-1,3-glucan and disulfide bridges; required for cell wall stability; induced by heat shock, oxidative stress, and nitrogen limitation | Glutamine-fructose-6-phosphate amidotransferase, catalyzes the formation of glucosamine-6-P and glutamate from fructose-6-P and glutamine in the first step of chitin biosynthesis | Outward-rectifier potassium channel of the plasma membrane with two pore domains in tandem, each of which forms a functional channel permeable to potassium; carboxy tail functions to prevent inner gate closures; target of K1 toxin | Polyamine transporter that recognizes spermine, putrescine, and spermidine; catalyzes uptake of polyamines at alkaline pH and excretion at acidic pH; phosphorylation enhances activity and sorting to the plasma membrane | Protein of unknown function; localizes to the cytoplasm; YLL042C is not an essential gene | Protein of unknown function, null mutation leads to an increase in sensitivity to Calcofluor white; expression of the gene is induced in the presence of isooctane | Basic leucine zipper (bZIP) transcription factor | Structural constituent of the cell wall attached to the plasma membrane by a GPI-anchor; expression is upregulated in response to cell wall stress | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the bud and cytoplasm; Hog1p is required for transcriptional induction in response to cell wall damage; YLR414C is not an essential gene | Putative mannosidase, essential glycosylphosphatidylinositol (GPI)-anchored membrane protein required for cell wall biogenesis in bud formation, involved in filamentous growth, homologous to Dcw1p | Protein of unknown function, involved in invasive and pseudohyphal growth; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Chitin synthase I, requires activation from zymogenic form in order to catalyze the transfer of N-acetylglucosamine (GlcNAc) to chitin; required for repairing the chitin septum during cytokinesis; transcription activated by mating factor | Phospholipase B (lysophospholipase) involved in phospholipid metabolism; hydrolyzes phosphatidylinositol and phosphatidylserine and displays transacylase activity in vitro | Partially redundant sensor-transducer of the stress-activated PKC1-MPK1 signaling pathway involved in maintenance of cell wall integrity; involved in the response to heat shock and other stressors; regulates 1,3-beta-glucan synthesis | Protein of unassigned function involved in activation of the Pma1p plasma membrane H+-ATPase by glucose | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YOR385W is not an essential gene | Putative aryl alcohol dehydrogenase; transcription is activated by paralogous transcription factors Yrm1p and Yrr1p along with genes involved in multidrug resistance | MADS-box transcription factor, component of the protein kinase C-mediated MAP kinase pathway involved in the maintenance of cell integrity; phosphorylated and activated by the MAP-kinase Slt2p | Polyamine transport protein specific for spermine; localizes to the plasma membrane; member of the major facilitator superfamily | Plasma membrane multidrug transporter of the major facilitator superfamily, acts as an extrusion permease; partial multicopy suppressor of gal11 mutations |
| 280 | 52 | 440 | 0.953 | 0.647 |  |  | 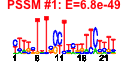 | 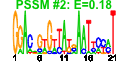 |  | YNL289W YOR201C YBL016W YDL171C YDL212W YDR044W YDR210W YDR261C YDR434W YER186C YGL012W YGL186C YGL232W YGR216C YHR031C YIL038C YIL114C YJL157C YJL196C YJR144W YKL185W YKL209C YKR084C YLR007W YLR113W YLR261C YML127W YMR088C YMR208W YMR220W YNL232W YNL268W YNL290W YNR033W YNR061C YOL002C YOL008W YOL059W YOL136C YOL143C YOL147C YOR002W YOR092W YOR202W YOR323C YOR360C YPL149W YPL179W YPL251W YPR088C YPR128C YPR138C | PCL1 MRM1 FUS3 GLT1 SHR3 HEM13 EXG2 GPI17 ERG4 TPN1 TAN1 GPI1 RRM3 NOT3 POR2 FAR1 ELO1 MGM101 ASH1 STE6 HBS1 NSE1 HOG1 RSC9 VBA1 ERG12 ERG8 CSL4 LYP1 RFC3 ABZ1 IZH2 COQ10 GPD2 PFK27 RIB4 PEX11 ALG6 ECM3 HIS3 PRO2 PDE2 ATG5 PPQ1 SRP54 PDR1 MEP3 | Pho85 cyclin of the Pcl1,2-like subfamily, involved in entry into the mitotic cell cycle and regulation of morphogenesis, localizes to sites of polarized cell growth | Ribose methyltransferase that modifies a functionally critical, conserved nucleotide in mitochondrial 21S rRNA | Mitogen-activated serine/threonine protein kinase involved in mating; phosphoactivated by Ste7p; substrates include Ste12p, Far1p, Bni1p, Sst2p; inhibits invasive growth during mating by phosphorylating Tec1p, promoting its degradation | NAD(+)-dependent glutamate synthase (GOGAT), synthesizes glutamate from glutamine and alpha-ketoglutarate; with Gln1p, forms the secondary pathway for glutamate biosynthesis from ammonia; expression regulated by nitrogen source | Endoplasmic reticulum packaging chaperone, required for incorporation of amino acid permeases into COPII coated vesicles for transport to the cell surface | Coproporphyrinogen III oxidase, an oxygen requiring enzyme that catalyzes the sixth step in the heme biosynthetic pathway; localizes to the mitochondrial inner membrane; transcription is repressed by oxygen and heme (via Rox1p and Hap1p) | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | Exo-1,3-beta-glucanase, involved in cell wall beta-glucan assembly; may be anchored to the plasma membrane via a glycosylphosphatidylinositol (GPI) anchor | Transmembrane protein subunit of the glycosylphosphatidylinositol transamidase complex that adds GPIs to newly synthesized proteins; human PIG-Sp homolog | Putative protein of unknown function | C-24(28) sterol reductase, catalyzes the final step in ergosterol biosynthesis; mutants are viable, but lack ergosterol | Plasma membrane pyridoxine (vitamin B6) transporter; member of the purine-cytosine permease subfamily within the major facilitator superfamily; proton symporter with similarity to Fcy21p, Fcy2p, and Fcy22p | Putative tRNA acetyltransferase, RNA-binding protein required for the formation of the modified nucleoside N(4)-acetylcytidine in serine and leucine tRNAs but not required for the same modification in 18S rRNA | Membrane protein involved in the synthesis of N-acetylglucosaminyl phosphatidylinositol (GlcNAc-PI), the first intermediate in the synthesis of glycosylphosphatidylinositol (GPI) anchors; human and mouse GPI1p are functional homologs | DNA helicase involved in rDNA replication and Ty1 transposition; relieves replication fork pauses at telomeric regions; structurally and functionally related to Pif1p | Subunit of the CCR4-NOT complex, which is a global transcriptional regulator with roles in transcription initiation and elongation and in mRNA degradation | Putative mitochondrial porin (voltage-dependent anion channel), related to Por1p but not required for mitochondrial membrane permeability or mitochondrial osmotic stability | Cyclin-dependent kinase inhibitor that mediates cell cycle arrest in response to pheromone; also forms a complex with Cdc24p, Ste4p, and Ste18p that may specify the direction of polarized growth during mating; potential Cdc28p substrate | Elongase I, medium-chain acyl elongase, catalyzes carboxy-terminal elongation of unsaturated C12-C16 fatty acyl-CoAs to C16-C18 fatty acids | Protein involved in mitochondrial genome maintenance; component of the mitochondrial nucleoid, required for the repair of oxidative mtDNA damage | Zinc-finger inhibitor of HO transcription; mRNA is localized and translated in the distal tip of anaphase cells, resulting in accumulation of Ash1p in daughter cell nuclei and inhibition of HO expression; potential Cdc28p substrate | Plasma membrane ATP-binding cassette (ABC) transporter required for the export of a-factor, catalyzes ATP hydrolysis coupled to a-factor transport; contains 12 transmembrane domains and two ATP binding domains; expressed only in MATa cells | GTP binding protein with sequence similarity to the elongation factor class of G proteins, EF-1alpha and Sup35p; associates with Dom34p, and shares a similar genetic relationship with genes that encode ribosomal protein components | Essential subunit of the Mms21-Smc5-Smc6 complex; nuclear protein required for DNA repair and growth | Mitogen-activated protein kinase involved in osmoregulation via three independent osmosensors; mediates the recruitment and activation of RNA Pol II at Hot1p-dependent promoters; localization regulated by Ptp2p and Ptp3p | Component of the RSC chromatin remodeling complex; DNA-binding protein involved in the synthesis of rRNA and in transcriptional repression and activation of genes regulated by the Target of Rapamycin (TOR) pathway | Permease of basic amino acids in the vacuolar membrane | Mevalonate kinase, acts in the biosynthesis of isoprenoids and sterols, including ergosterol, from mevalonate | Phosphomevalonate kinase, an essential cytosolic enzyme that acts in the biosynthesis of isoprenoids and sterols, including ergosterol, from mevalonate | Subunit of the exosome, which is an essential complex present in both nucleus and cytoplasm that mediates RNA processing and degradation | Lysine permease; one of three amino acid permeases (Alp1p, Can1p, Lyp1p) responsible for uptake of cationic amino acids | Subunit of heteropentameric Replication factor C (RF-C), which is a DNA binding protein and ATPase that acts as a clamp loader of the proliferating cell nuclear antigen (PCNA) processivity factor for DNA polymerases delta and epsilon | Para-aminobenzoate (PABA) synthase, has similarity to Escherichia coli PABA synthase components PabA and PabB | Plasma membrane protein involved in zinc metabolism and osmotin-induced apoptosis; transcription regulated by Zap1p, zinc and fatty acid levels; similar to mammalian adiponectins; deletion increases sensitivity to elevated zinc | Coenzyme Q (ubiquinone) binding protein, functions in the delivery of Q6 to its proper location for electron transport during respiration; START domain protein with homologs in bacteria and eukaryotes | NAD-dependent glycerol 3-phosphate dehydrogenase, homolog of Gpd1p, expression is controlled by an oxygen-independent signaling pathway required to regulate metabolism under anoxic conditions; located in cytosol and mitochondria | 6-phosphofructo-2-kinase, catalyzes synthesis of fructose-2,6-bisphosphate; inhibited by phosphoenolpyruvate and sn-glycerol 3-phosphate, expression induced by glucose and sucrose, transcriptional regulation involves protein kinase A | Lumazine synthase (6,7-dimethyl-8-ribityllumazine synthase, also known as DMRL synthase); catalyzes synthesis of immediate precursor to riboflavin | Peroxisomal membrane protein required for peroxisome proliferation and medium-chain fatty acid oxidation, most abundant protein in the peroxisomal membrane, regulated by Adr1p and Pip2p-Oaf1p, promoter contains ORE and UAS1-like elements | Alpha 1,3 glucosyltransferase, involved in transfer of oligosaccharides from dolichyl pyrophosphate to asparagine residues of proteins during N-linked protein glycosylation; mutations in human ortholog are associated with disease | Non-essential protein of unknown function | Imidazoleglycerol-phosphate dehydratase, catalyzes the sixth step in histidine biosynthesis; mutations cause histidine auxotrophy and sensitivity to Cu, Co, and Ni salts; transcription is regulated by general amino acid control via Gcn4p | Gamma-glutamyl phosphate reductase, catalyzes the second step in proline biosynthesis | High-affinity cyclic AMP phosphodiesterase, component of the cAMP-dependent protein kinase signaling system, protects the cell from extracellular cAMP, contains readthrough motif surrounding termination codon | Conserved autophagy-related protein that undergoes conjugation with Atg12p and then associates with Atg16p to form a cytosolic complex essential for autophagosome formation | Putative protein serine/threonine phosphatase; null mutation enhances efficiency of translational suppressors | Signal recognition particle (SRP) subunit (homolog of mammalian SRP54); contains the signal sequence-binding activity of SRP, interacts with the SRP RNA, and mediates binding of SRP to signal receptor; contains GTPase domain | Zinc cluster protein that is a master regulator involved in recruiting other zinc cluster proteins to pleiotropic drug response elements (PDREs) to fine tune the regulation of multidrug resistance genes | Ammonium permease of high capacity and low affinity; belongs to a ubiquitous family of cytoplasmic membrane proteins that transport only ammonium (NH4+); expression is under the nitrogen catabolite repression regulation ammonia permease |
| 15 | 41 | 441 | 0.962 | 0.592 |  |  |  |  |  | YBR016W YBR035C YCR029C YCR030C YCR076C YCRX17W YDL018C YDL086W YDR063W YDR154C YDR155C YDR487C YER004W YER063W YER163C YER177W YGL242C YGR026W YIL088C YJL052W YJR074W YJR117W YKL007W YKL025C YKL167C YKL190W YKR047W YLR203C YLR330W YML030W YMR086W YMR184W YMR276W YNL138W YNL239W YNR036C YOL049W YOR135C YOR288C YOR321W YOR332W | PDX3 SYP1 ERP3 AIM7 CPR1 RIB3 FMP52 THO1 BMH1 AVT7 TDH1 MOG1 STE24 CAP1 PAN3 MRP49 CNB1 MSS51 CHS5 AIM31 ADD37 DSK2 YSF3 LAP3 GSH2 MPD1 PMT3 VMA4 | Plasma membrane protein of unknown function; has similarity to hydrophilins, which are hydrophilic, glycine-rich proteins involved in the adaptive response to hyperosmotic conditions | Pyridoxine (pyridoxamine) phosphate oxidase, has homologs in E. coli and Myxococcus xanthus; transcription is under the general control of nitrogen metabolism | Protein with a potential role in actin cytoskeletal organization; overexpression suppresses a pfy1 (profilin) null mutation | Putative protein of unknown function; YCR076C is not an essential gene | Protein with similarity to Emp24p and Erv25p, member of the p24 family involved in ER to Golgi transport | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies; YDL086W is not an essential gene | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; null mutant is viable and displays decreased frequency of mitochondrial genome loss (petite formation) | Cytoplasmic peptidyl-prolyl cis-trans isomerase (cyclophilin), catalyzes the cis-trans isomerization of peptide bonds N-terminal to proline residues; binds the drug cyclosporin A | 3,4-dihydroxy-2-butanone-4-phosphate synthase (DHBP synthase), required for riboflavin biosynthesis from ribulose-5-phosphate, also has an unrelated function in mitochondrial respiration | Protein of unknown function, localized to the mitochondrial outer membrane | Conserved nuclear RNA-binding protein; specifically binds to transcribed chromatin in a THO- and RNA-dependent manner, genetically interacts with shuttling hnRNP NAB2; overproduction suppresses transcriptional defect caused by hpr1 mutation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus | 14-3-3 protein, major isoform; controls proteome at post-transcriptional level, binds proteins and DNA, involved in regulation of many processes including exocytosis, vesicle transport, Ras/MAPK signaling, and rapamycin-sensitive signaling | Putative protein of unknown function; deletion mutant is viable | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | Putative transporter, member of a family of seven S. cerevisiae genes (AVT1-7) related to vesicular GABA-glycine transporters | Glyceraldehyde-3-phosphate dehydrogenase, isozyme 1, involved in glycolysis and gluconeogenesis; tetramer that catalyzes the reaction of glyceraldehyde-3-phosphate to 1,3 bis-phosphoglycerate; detected in the cytoplasm and cell-wall | Conserved nuclear protein that interacts with GTP-Gsp1p, which is a Ran homolog of the Ras GTPase family, and stimulates nucleotide release, involved in nuclear protein import, nucleotide release is inhibited by Yrb1p | Highly conserved zinc metalloprotease that functions in two steps of a-factor maturation, C-terminal CAAX proteolysis and the first step of N-terminal proteolytic processing; contains multiple transmembrane spans | Alpha subunit of the capping protein (CP) heterodimer (Cap1p and Cap2p) which binds to the barbed ends of actin filaments preventing further polymerization; localized predominantly to cortical actin patches | Essential subunit of the Pan2p-Pan3p poly(A)-ribonuclease complex, which acts to control poly(A) tail length and regulate the stoichiometry and activity of postreplication repair complexes | Mitochondrial ribosomal protein of the large subunit, not essential for mitochondrial translation | Calcineurin B; the regulatory subunit of calcineurin, a Ca++/calmodulin-regulated type 2B protein phosphatase which regulates Crz1p (a stress-response transcription factor), the other calcineurin subunit is encoded by CNA1 and/or CMP1 | Nuclear encoded protein required for translation of COX1 mRNA; binds to Cox1 protein | Component of the exomer complex, which also contains Csh6p, Bch1p, Bch2p, and Bud7p and is involved in export of selected proteins, such as chitin synthase Chs3p, from the Golgi to the plasma membrane | Putative protein of unknown function; GFP-fusion protein localizes to mitochondria; null mutant is viable and displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth rate in minimal glycerol media | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery; cAMP represses YMR086W expression | Protein of unknown function involved in ER-associated protein degradation; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and is induced in response to the DNA-damaging agent MMS; YMR184W is not an essential gene | Nuclear-enriched ubiquitin-like polyubiquitin-binding protein, required for spindle pole body (SPB) duplication and for transit through the G2/M phase of the cell cycle, involved in proteolysis, interacts with the proteasome | Component of the U2 snRNP, associated with the SF3b complex; conserved in Ashbya gossypii | Cysteine aminopeptidase with homocysteine-thiolactonase activity; protects cells against homocysteine toxicity; has bleomycin hydrolase activity in vitro; transcription is regulated by galactose via Gal4p; orthologous to human BLMH | Mitochondrial protein; may interact with ribosomes based on co-purification experiments; similar to E. coli and human mitochondrial S12 ribosomal proteins | Glutathione synthetase, catalyzes the ATP-dependent synthesis of glutathione (GSH) from gamma-glutamylcysteine and glycine; induced by oxidative stress and heat shock | Member of the protein disulfide isomerase (PDI) family; interacts with and inhibits the chaperone activity of Cne1p; MPD1 overexpression in a pdi1 null mutant suppresses defects in Pdi1p functions such as carboxypeptidase Y maturation | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; acts in a complex with Pmt5p, can instead interact with Pmt1p in some conditions; target for new antifungals | Subunit E of the eight-subunit V1 peripheral membrane domain of the vacuolar H+-ATPase (V-ATPase), an electrogenic proton pump found throughout the endomembrane system; required for the V1 domain to assemble onto the vacuolar membrane |
| 170 | 49 | 449 | 0.965 | 0.630 |  |  | 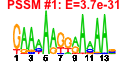 | 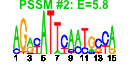 |  | YAL012W YBR021W YBR281C YDR291W YDR405W YER001W YGL125W YGR055W YIL074C YIL093C YJL212C YJR010W YKL001C YKL054C YKL058W YKR069W YLL062C YLR247C YLR303W YMR089C YNL219C YNR029C YOL004W YOL019W YOR006C YPL026C YER152C YFR030W YHR163W YHR183W YHR186C YIR021W YJL060W YJR040W YJR137C YKL101W YKL128C YLL052C YLL061W YLR304C YMR189W YNL121C YNL138W YNL277W YOL138C YPL104W YPL105C YPR086W YPR167C | CYS3 FUR4 DUG2 HRQ1 MRP20 MNN1 MET13 MUP1 SER33 RSM25 OPT1 MET3 MET14 DEF1 TOA2 MET1 MHT1 IRC20 MET17 YTA12 ALG9 SIN3 YOL019W-A TSR3 SKS1 MET10 SOL3 GND1 KOG1 MRS1 BNA3 GEF1 ECM17 HSL1 PMU1 AQY2 MMP1 ACO1 GCV2 TOM70 YSF3 YNL277W-A MSD1 SUA7 MET16 | Cystathionine gamma-lyase, catalyzes one of the two reactions involved in the transsulfuration pathway that yields cysteine from homocysteine with the intermediary formation of cystathionine | Uracil permease, localized to the plasma membrane; expression is tightly regulated by uracil levels and environmental cues | Probable di- and tri-peptidase; forms a complex with Dug1p and Dug3p to degrade glutathione (GSH) and other peptides containing a gamma-glu-X bond in an alternative pathway to GSH degradation by gamma-glutamyl transpeptidase (Ecm38p) | Putative DNA helicase | Mitochondrial ribosomal protein of the large subunit | Alpha-1,3-mannosyltransferase, integral membrane glycoprotein of the Golgi complex, required for addition of alpha1,3-mannose linkages to N-linked and O-linked oligosaccharides, one of five S. cerevisiae proteins of the MNN1 family | Isozyme of methylenetetrahydrofolate reductase, catalyzes the reduction of 5,10-methylenetetrahydrofolate to 5-methyltetrahydrofolate in the methionine biosynthesis pathway | High affinity methionine permease, integral membrane protein with 13 putative membrane-spanning regions; also involved in cysteine uptake | 3-phosphoglycerate dehydrogenase, catalyzes the first step in serine and glycine biosynthesis; isozyme of Ser3p | Mitochondrial ribosomal protein of the small subunit | Proton-coupled oligopeptide transporter of the plasma membrane; also transports glutathione and phytochelatin; member of the OPT family | ATP sulfurylase, catalyzes the primary step of intracellular sulfate activation, essential for assimilatory reduction of sulfate to sulfide, involved in methionine metabolism | Adenylylsulfate kinase, required for sulfate assimilation and involved in methionine metabolism | RNAPII degradation factor, forms a complex with Rad26p in chromatin, enables ubiquitination and proteolysis of RNAPII present in an elongation complex | TFIIA small subunit; involved in transcriptional activation, acts as antirepressor or as coactivator; homologous to smallest subunit of human and Drosophila TFIIA | S-adenosyl-L-methionine uroporphyrinogen III transmethylase, involved in the biosynthesis of siroheme, a prosthetic group used by sulfite reductase; required for sulfate assimilation and methionine biosynthesis | S-methylmethionine-homocysteine methyltransferase, functions along with Sam4p in the conversion of S-adenosylmethionine (AdoMet) to methionine to control the methionine/AdoMet ratio | Putative helicase; localized to mitochondria and the nucleus; YLR247C is not an essential gene; null mutant displays increased levels of spontaneous Rad52 foci | Methionine and cysteine synthase (O-acetyl homoserine-O-acetyl serine sulfhydrylase), required for sulfur amino acid synthesis | Component, with Afg3p, of the mitochondrial inner membrane m-AAA protease that mediates degradation of misfolded or unassembled proteins and is also required for correct assembly of mitochondrial enzyme complexes | Mannosyltransferase, involved in N-linked glycosylation; catalyzes the transfer of mannose from Dol-P-Man to lipid-linked oligosaccharides; mutation of the human ortholog causes type 1 congenital disorders of glycosylation | Putative protein of unknown function, deletion confers reduced fitness in saline | Component of the Sin3p-Rpd3p histone deacetylase complex, involved in transcriptional repression and activation of diverse processes, including mating-type switching and meiosis; involved in the maintenance of chromosomal integrity | Identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus; Null mutant accumulates 20S pre-rRNA | Putative serine/threonine protein kinase; involved in the adaptation to low concentrations of glucose independent of the SNF3 regulated pathway | Putative protein of unknown function; shares amino acid similarity with the aminotransferases Aro8p and Aro9p; YER152C is not an essential gene | Subunit alpha of assimilatory sulfite reductase, which is responsible for the conversion of sulfite into sulfide | 6-phosphogluconolactonase, catalyzes the second step of the pentose phosphate pathway; weak multicopy suppressor of los1-1 mutation; homologous to Sol2p and Sol1p | 6-phosphogluconate dehydrogenase (decarboxylating), catalyzes an NADPH regenerating reaction in the pentose phosphate pathway; required for growth on D-glucono-delta-lactone and adaptation to oxidative stress | Subunit of TORC1, a rapamycin-sensitive complex involved in growth control that contains Tor1p or Tor2p, Lst8p and Tco89p; contains four HEAT repeats and seven WD-40 repeats; may act as a scaffold protein to couple TOR and its effectors | Protein required for the splicing of two mitochondrial group I introns (BI3 in COB and AI5beta in COX1); forms a splicing complex, containing four subunits of Mrs1p and two subunits of the BI3-encoded maturase, that binds to the BI3 RNA | Arylformamidase, involved in biosynthesis of nicotinic acid from tryptophan via kynurenine pathway; potential Cdc28p substrate | Chloride channel localized to late- or post-Golgi vesicles, involved in iron metabolism; highly homologous to voltage-gated chloride channels in vertebrates | Sulfite reductase beta subunit, involved in amino acid biosynthesis, transcription repressed by methionine | Nim1p-related protein kinase that regulates the morphogenesis and septin checkpoints; associates with the assembled septin filament; required along with Hsl7p for bud neck recruitment, phosphorylation, and degradation of Swe1p | Putative phosphomutase, contains a region homologous to the active site of phosphomutases; overexpression suppresses the histidine auxotrophy of an ade3 ade16 ade17 triple mutant and the temperature sensitivity of a tps2 mutant | Water channel that mediates the transport of water across cell membranes, only expressed in proliferating cells, controlled by osmotic signals, may be involved in freeze tolerance; disrupted by a stop codon in many S. cerevisiae strains | High-affinity S-methylmethionine permease, required for utilization of S-methylmethionine as a sulfur source; has similarity to S-adenosylmethionine permease Sam3p | Aconitase, required for the tricarboxylic acid (TCA) cycle and also independently required for mitochondrial genome maintenance; phosphorylated; component of the mitochondrial nucleoid; mutation leads to glutamate auxotrophy | P subunit of the mitochondrial glycine decarboxylase complex, required for the catabolism of glycine to 5,10-methylene-THF; expression is regulated by levels of 5,10-methylene-THF in the cytoplasm | Component (70 kDa) of the TOM (translocase of outer membrane) complex responsible for recognition and initial import steps for all mitochondrially directed proteins; acts as a receptor for incoming precursor proteins | Component of the U2 snRNP, associated with the SF3b complex; conserved in Ashbya gossypii | Putative protein of unknown function | Putative protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Mitochondrial aspartyl-tRNA synthetase, required for acylation of aspartyl-tRNA; yeast and bacterial aspartyl-, asparaginyl-, and lysyl-tRNA synthetases contain regions with high sequence similarity, suggesting a common ancestral gene | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Transcription factor TFIIB, a general transcription factor required for transcription initiation and start site selection by RNA polymerase II | 3'-phosphoadenylsulfate reductase, reduces 3'-phosphoadenylyl sulfate to adenosine-3',5'-bisphosphate and free sulfite using reduced thioredoxin as cosubstrate, involved in sulfate assimilation and methionine metabolism |
| 77 | 46 | 446 | 0.975 | 0.661 |  |  | 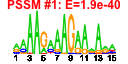 | 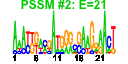 |  | YBL016W YBL074C YBR067C YCL073C YDL037C YDL073W YDL141W YDL160C YDL200C YDL240W YDR038C YDR039C YDR052C YDR112W YEL035C YER064C YER075C YER095W YER165W YGL038C YGR166W YHR117W YHR214W YJL018W YJL117W YKL038W YKR075C YNL014W YNL277W YNR067C YNR075W YBR083W YDL127W YDL189W YDL205C YDR040C YDR216W YDR524C YGL032C YHL026C YHR094C YIL141W YJL025W YKL184W YLR436C YNL269W | FUS3 AAR2 TIP1 BSC1 BPL1 YDL160C-A MGT1 LRG1 ENA5 ENA2 DBF4 UTR5 PTP3 RAD51 PAB1 OCH1 KRE11 TOM71 MPS3 PHO86 RGT1 HEF3 YNL277W-A DSE4 COS10 TEC1 PCL2 RBS1 HEM3 ENA1 ADR1 AGE1 AGA2 HXT1 RRN7 SPE1 ECM30 BSC4 | Mitogen-activated serine/threonine protein kinase involved in mating; phosphoactivated by Ste7p; substrates include Ste12p, Far1p, Bni1p, Sst2p; inhibits invasive growth during mating by phosphorylating Tec1p, promoting its degradation | Component of the U5 snRNP, required for splicing of U3 precursors; originally described as a splicing factor specifically required for splicing pre-mRNA of the MATa1 cistron | Major cell wall mannoprotein with possible lipase activity; transcription is induced by heat- and cold-shock; member of the Srp1p/Tip1p family of serine-alanine-rich proteins | Protein of unconfirmed function; displays a topology characteristic of the Major Facilitators Superfamily of membrane proteins; coding sequence 98% identical to that of YKR106W | Protein of unconfirmed function, similar to cell surface flocculin Muc1p; ORF exhibits genomic organization compatible with a translational readthrough-dependent mode of expression | Putative protein of unknown function; YDL073W is not an essential gene | Biotin:apoprotein ligase, covalently modifies proteins with the addition of biotin, required for acetyl-CoA carboxylase (Acc1p) holoenzyme formation | Putative protein of unknown function | DNA repair methyltransferase (6-O-methylguanine-DNA methylase) involved in protection against DNA alkylation damage | Putative GTPase-activating protein (GAP) involved in the Pkc1p-mediated signaling pathway that controls cell wall integrity; appears to specifically regulate 1,3-beta-glucan synthesis | Protein with similarity to P-type ATPase sodium pumps, member of the Na+ efflux ATPase family | P-type ATPase sodium pump, involved in Na+ efflux to allow salt tolerance; likely not involved in Li+ efflux | Regulatory subunit of Cdc7p-Dbf4p kinase complex, required for Cdc7p kinase activity and initiation of DNA replication; phosphorylates the Mcm2-7 family of proteins; cell cycle regulated | Protein of unknown function; transcription may be regulated by Gcr1p | Non-essential nuclear protein; null mutation has global effects on transcription | Phosphotyrosine-specific protein phosphatase involved in the inactivation of mitogen-activated protein kinase (MAPK) during osmolarity sensing; dephosporylates Hog1p MAPK and regulates its localization; localized to the cytoplasm | Strand exchange protein, forms a helical filament with DNA that searches for homology; involved in the recombinational repair of double-strand breaks in DNA during vegetative growth and meiosis; homolog of Dmc1p and bacterial RecA protein | Poly(A) binding protein, part of the 3'-end RNA-processing complex, mediates interactions between the 5' cap structure and the 3' mRNA poly(A) tail, involved in control of poly(A) tail length, interacts with translation factor eIF-4G | Mannosyltransferase of the cis-Golgi apparatus, initiates the polymannose outer chain elongation of N-linked oligosaccharides of glycoproteins | Protein involved in biosynthesis of cell wall beta-glucans; subunit of the TRAPP (transport protein particle) complex, which is involved in the late steps of endoplasmic reticulum to Golgi transport | Mitochondrial outer membrane protein with similarity to Tom70p; probable minor component of the TOM (translocase of outer membrane) complex responsible for recognition and import of mitochondrially directed proteins | Putative protein of unknown function; predicted to be a glycosylphosphatidylinositol-modified (GPI) protein | Essential integral membrane protein required for spindle pole body duplication and for nuclear fusion, localizes to the spindle pole body half bridge, interacts with DnaJ-like chaperone Jem1p and with centrin homolog Cdc31p | Endoplasmic reticulum (ER) resident protein required for ER exit of the high-affinity phosphate transporter Pho84p, specifically required for packaging of Pho84p into COPII vesicles | Glucose-responsive transcription factor that regulates expression of several glucose transporter (HXT) genes in response to glucose; binds to promoters and acts both as a transcriptional activator and repressor | Protein of unknown function; similar to YOR062Cp and Reg1p; expression regulated by glucose and Rgt1p; GFP-fusion protein is induced in response to the DNA-damaging agent MMS | Translational elongation factor EF-3; paralog of YEF3 and member of the ABC superfamily; stimulates EF-1 alpha-dependent binding of aminoacyl-tRNA by the ribosome; normally expressed in zinc deficient cells | Daughter cell-specific secreted protein with similarity to glucanases, degrades cell wall from the daughter side causing daughter to separate from mother | Protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins | Transcription factor required for full Ty1 epxression, Ty1-mediated gene activation, and haploid invasive and diploid pseudohyphal growth; TEA/ATTS DNA-binding domain family member | G1 cyclin, associates with Pho85p cyclin-dependent kinase (Cdk) to contribute to entry into the mitotic cell cycle, essential for cell morphogenesis; localizes to sites of polarized cell growth | Protein of unknown function, identified as a high copy suppressor of psk1 psk2 mutations that confer temperature-sensitivity for galactose utilization; proposed to bind single-stranded nucleic acids via its R3H domain | Phorphobilinogen deaminase, catalyzes the conversion of 4-porphobilinogen to hydroxymethylbilane, the third step in the heme biosynthetic pathway; localizes to both the cytoplasm and nucleus; expression is regulated by Hap2p-Hap3p | P-type ATPase sodium pump, involved in Na+ and Li+ efflux to allow salt tolerance | Carbon source-responsive zinc-finger transcription factor, required for transcription of the glucose-repressed gene ADH2, of peroxisomal protein genes, and of genes required for ethanol, glycerol, and fatty acid utilization | ADP-ribosylation factor (ARF) GTPase activating protein (GAP) effector, involved in the secretory and endocytic pathways; contains C2C2H2 cysteine/histidine motif | Adhesion subunit of a-agglutinin of a-cells, C-terminal sequence acts as a ligand for alpha-agglutinin (Sag1p) during agglutination, modified with O-linked oligomannosyl chains, linked to anchorage subunit Aga1p via two disulfide bonds | Putative protein of unknown function; YHL026C is not an essential gene; in 2005 the start site was moved 141 nt upstream (see Locus History) | Low-affinity glucose transporter of the major facilitator superfamily, expression is induced by Hxk2p in the presence of glucose and repressed by Rgt1p when glucose is limiting | Protein involved in the transcription of 35S rRNA genes by RNA polymerase I; component of the core factor (CF) complex also composed of Rrn11p, Rrn6p and TATA-binding protein | Ornithine decarboxylase, catalyzes the first step in polyamine biosynthesis; degraded in a proteasome-dependent manner in the presence of excess polyamines | Non-essential protein of unknown function | Protein of unknown function, ORF exhibits genomic organization compatible with a translational readthrough-dependent mode of expression; readthrough is increased upon depletion of Sup35p |
| 214 | 46 | 439 | 1.004 | 0.597 |  |  | 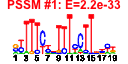 |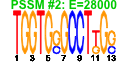  |  | YBL006C YDL217C YJR141W YKL053W YLR316C YMR037C YMR064W YMR233W YNL140C YNL251C YOL059W YOR229W YOR324C YPR011C YPR038W YPR131C YHR014W YHR034C YHR045W YJR049C YKL052C YKL214C YKR045C YLR210W YLR446W YML088W YML114C YMR032W YMR061W YNL004W YNL042W YNL072W YNL191W YNL264C YNR020C YNR049C YOL112W YOR300W YOR325W YOR377W YPL026C YPL038W YPL049C YPL270W YPR037C YPR152C | LDB7 TIM22 TAD3 MSN2 AEP1 TRI1 NRD1 GPD2 WTM2 FRT1 NAT3 SPO13 PIH1 UTR1 ASK1 YRA2 CLB4 UFO1 TAF8 HOF1 RNA14 HRB1 YNL042W-B RNH201 DUG3 PDR17 ATP23 MSO1 MSB4 ATF1 SKS1 YPL038W-A DIG1 MDL2 ERV2 URN1 | Component of the RSC chromatin remodeling complex; interacts with Rsc3p, Rsc30p, Npl6p, and Htl1p to form a module important for a broad range of RSC functions | Component of the mitochondrial Tim54p-Tim22p complex involved in insertion of polytopic proteins into the inner membrane | Putative protein of unknown function; may be involved in mRNA processing; YJR141W is an essential gene | Subunit of tRNA-specific adenosine-34 deaminase, forms a heterodimer with Tad2p that converts adenosine to inosine at the wobble position of several tRNAs | Transcriptional activator related to Msn4p; activated in stress conditions, which results in translocation from the cytoplasm to the nucleus; binds DNA at stress response elements of responsive genes, inducing gene expression | Protein required for expression of the mitochondrial OLI1 gene encoding subunit 9 of F1-F0 ATP synthase | Non-essential sumoylated protein of unknown function with similarity to components of human SWI/SNF complex including SMRD3; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm, nucleus and nucleolus | RNA-binding protein that interacts with the C-terminal domain of the RNA polymerase II large subunit (Rpo21p), required for transcription termination and 3' end maturation of nonpolyadenylated RNAs | NAD-dependent glycerol 3-phosphate dehydrogenase, homolog of Gpd1p, expression is controlled by an oxygen-independent signaling pathway required to regulate metabolism under anoxic conditions; located in cytosol and mitochondria | Transcriptional repressor involved in regulation of meiosis and silencing; contains WD repeats | Tail-anchored endoplasmic reticulum membrane protein that is a substrate of the phosphatase calcineurin, interacts with homolog Frt2p, promotes cell growth in conditions of high Na+, alkaline pH, and cell wall stress | Putative transporter, member of the mitochondrial carrier family; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Catalytic subunit of the NatB N-terminal acetyltransferase, which catalyzes acetylation of the amino-terminal methionine residues of all proteins beginning with Met-Asp or Met-Glu and of some proteins beginning with Met-Asn or Met-Met | Meiosis-specific protein, involved in maintaining sister chromatid cohesion during meiosis I as well as promoting proper attachment of kinetochores to the spindle during meiosis I and meiosis II | Protein of unresolved function; may function in protein folding and/or rRNA processing, interacts with a chaperone (Hsp82p), two chromatin remodeling factors (Rvb1p, Rvb2p) and two rRNA processing factors (Rrp43p, Nop58p) | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum | ATP-NADH kinase; phosphorylates both NAD and NADH; active as a hexamer; enhances the activity of ferric reductase (Fre1p) | Essential subunit of the Dam1 complex (aka DASH complex), couples kinetochores to the force produced by MT depolymerization thereby aiding in chromosome segregation; phosphorylated during the cell cycle by cyclin-dependent kinases | Member of the REF (RNA and export factor binding proteins) family; when overexpressed, can substitute for the function of Yra1p in export of poly(A)+ mRNA from the nucleus | Putative protein of unknown function; epitope-tagged protein localizes to the cytoplasm | B-type cyclin involved in cell cycle progression; activates Cdc28p to promote the G2/M transition; may be involved in DNA replication and spindle assembly; accumulates during S phase and G2, then targeted for ubiquitin-mediated degradation | Putative protein of unknown function with similarity to hexokinases; transcript is upregulated during sporulation and the unfolded protein response; YLR446W is not an essential gene | F-box receptor protein, subunit of the Skp1-Cdc53-F-box receptor (SCF) E3 ubiquitin ligase complex; binds to phosphorylated Ho endonuclease, allowing its ubiquitylation by SCF and subsequent degradation | TFIID subunit (65 kDa), involved in RNA polymerase II transcription initiation | Bud neck-localized, SH3 domain-containing protein required for cytokinesis; regulates actomyosin ring dynamics and septin localization; interacts with the formins, Bni1p and Bnr1p, and with Cyk3p, Vrp1p, and Bni5p | Cleavage and polyadenylation factor I (CF I) component involved in cleavage and polyadenylation of mRNA 3' ends; bridges interaction between Rna15p and Hrp1p in the CF I complex | Poly(A+) RNA-binding protein, involved in the export of mRNAs from the nucleus to the cytoplasm; similar to Gbp2p and Npl3p | Putative protein of unknown function | Ribonuclease H2 catalytic subunit, removes RNA primers during Okazaki fragment synthesis; cooperates with Rad27p nuclease | Probable glutamine amidotransferase, forms a complex with Dug1p and Dug2p to degrade glutathione (GSH) and other peptides containing a gamma-glu-X bond in an alternative pathway to GSH degradation by gamma-glutamyl transpeptidase (Ecm38p) | Phosphatidylinositol transfer protein (PITP), downregulates Plb1p-mediated turnover of phosphatidylcholine, found in the cytosol and microsomes, homologous to Pdr16p, deletion affects phospholipid composition | Putative metalloprotease of the mitochondrial inner membrane, required for processing of Atp6p; has an additional role in assembly of the F0 sector of the F1F0 ATP synthase complex | Probable component of the secretory vesicle docking complex, acts at a late step in secretion; shows genetic and physical interactions with Sec1p and is enriched in microsomal membrane fractions; required for sporulation | GTPase-activating protein of the Ras superfamily that acts primarily on Sec4p, localizes to the bud site and bud tip, has similarity to Msb3p; msb3 msb4 double mutation causes defects in secretion and actin organization | Alcohol acetyltransferase with potential roles in lipid and sterol metabolism; responsible for the major part of volatile acetate ester production during fermentation | Putative serine/threonine protein kinase; involved in the adaptation to low concentrations of glucose independent of the SNF3 regulated pathway | Putative protein of unknown function; identified by fungal homology and RT-PCR | Regulatory protein of unknown function, constitutively-expressed, involved in the regulation of mating-specific genes and the invasive growth pathway, required for MAP-kinase imposed repression, inhibits pheromone-responsive transcription | Mitochondrial inner membrane half-type ATP-binding cassette (ABC) transporter | Flavin-linked sulfhydryl oxidase localized to the endoplasmic reticulum lumen, involved in disulfide bond formation within the ER | Pre-mRNA splicing factor associated with the U2-U5-U6 snRNPs, the RES complex, and the Prp19-associated complex (NTC); null mutation displays synthetic genetic interactions with mutations affecting U2 snRNA and pre-mRNA splicing factors |
| 193 | 41 | 445 | 1.007 | 0.563 |  |  | 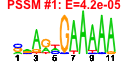 |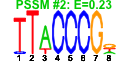  |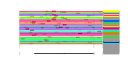  | YBR096W YDL015C YDR045C YDR433W YDR519W YEL027W YGL001C YGL200C YGR064W YGR078C YJR139C YKL153W YLL014W YLR043C YLR065C YMR214W YNL043C YNL090W YPL028W YBR175W YBR206W YBR243C YCR090C YDL143W YDL219W YDL228C YDL236W YDR209C YEL033W YER148W YGR026W YGR063C YGR101W YHR049W YIL145C YJL009W YJR076C YLR008C YLR208W YPL010W YPL246C | TSC13 RPC11 FPR2 CUP5 ERG26 EMP24 PAC10 HOM6 TRX2 SCJ1 RHO2 ERG10 SWD3 ALG7 PET9 DTD1 PHO13 SPT15 SPT4 PCP1 FSH1 PAN6 CDC11 PAM18 SEC13 RET3 RBD2 | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the ER | Enoyl reductase that catalyzes the last step in each cycle of very long chain fatty acid elongation, localizes to the ER, highly enriched in a structure marking nuclear-vacuolar junctions, coimmunoprecipitates with elongases Fen1p and Sur4p | RNA polymerase III subunit C11; mediates pol III RNA cleavage activity and is important for termination of transcription; homologous to TFIIS | Membrane-bound peptidyl-prolyl cis-trans isomerase (PPIase), binds to the drugs FK506 and rapamycin; expression pattern suggests possible involvement in ER protein trafficking | Proteolipid subunit of the vacuolar H(+)-ATPase V0 sector (subunit c; dicyclohexylcarbodiimide binding subunit); required for vacuolar acidification and important for copper and iron metal ion homeostasis | C-3 sterol dehydrogenase, catalyzes the second of three steps required to remove two C-4 methyl groups from an intermediate in ergosterol biosynthesis | Integral membrane component of endoplasmic reticulum-derived COPII-coated vesicles, which function in ER to Golgi transport | Part of the heteromeric co-chaperone GimC/prefoldin complex, which promotes efficient protein folding | Homoserine dehydrogenase (L-homoserine:NADP oxidoreductase), dimeric enzyme that catalyzes the third step in the common pathway for methionine and threonine biosynthesis; enzyme has nucleotide-binding, dimerization and catalytic regions | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic reticulum; YLL014W is not an essential gene | Cytoplasmic thioredoxin isoenzyme of the thioredoxin system which protects cells against oxidative and reductive stress, forms LMA1 complex with Pbi2p, acts as a cofactor for Tsa1p, required for ER-Golgi transport and vacuole inheritance | Putative protein of unknown function; YLR065C is not an essential gene | One of several homologs of bacterial chaperone DnaJ, located in the ER lumen where it cooperates with Kar2p to mediate maturation of proteins | Non-essential small GTPase of the Rho/Rac subfamily of Ras-like proteins, involved in the establishment of cell polarity and in microtubule assembly | Acetyl-CoA C-acetyltransferase (acetoacetyl-CoA thiolase), cytosolic enzyme that transfers an acetyl group from one acetyl-CoA molecule to another, forming acetoacetyl-CoA; involved in the first step in mevalonate biosynthesis | Essential subunit of the COMPASS (Set1C) complex, which methylates histone H3 on lysine 4 and is required in transcriptional silencing near telomeres; WD40 beta propeller superfamily member and ortholog of mammalian WDR5 | UDP-N-acetyl-glucosamine-1-P transferase, transfers Glc-Nac-P from UDP-GlcNac to Dol-P in the ER in the first step of the dolichol pathway of protein asparagine-linked glycosylation; inhibited by tunicamycin | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YCR090C is not an essential gene | Major ADP/ATP carrier of the mitochondrial inner membrane, exchanges cytosolic ADP for mitochondrially synthesized ATP; phosphorylated; required for viability in many common lab strains carrying a mutation in the polymorphic SAL1 gene | D-Tyr-tRNA(Tyr) deacylase, functions in protein translation, may affect nonsense suppression via alteration of the protein synthesis machinery; ubiquitous among eukaryotes | Alkaline phosphatase specific for p-nitrophenyl phosphate, involved in dephosphorylation of histone II-A and casein | TATA-binding protein, general transcription factor that interacts with other factors to form the preinitiation complex at promoters, essential for viability | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | Protein that forms a complex with Spt5p and mediates both activation and inhibition of transcription elongation, and plays a role in pre-mRNA processing; in addition, Spt4p is involved in kinetochore function and gene silencing | Mitochondrial serine protease required for the processing of various mitochondrial proteins and maintenance of mitochondrial DNA and morphology; belongs to the rhomboid-GlpG superfamily of intramembrane peptidases | Serine hydrolase that localizes to both the nucleus and cytoplasm; sequence is similar to Fsh2p and Fsh3p | Pantothenate synthase, also known as pantoate-beta-alanine ligase, required for pantothenic acid biosynthesis, deletion causes pantothenic acid auxotrophy, homologous to E. coli panC | Component of the septin ring of the mother-bud neck that is required for cytokinesis; septins recruit proteins to the neck and can act as a barrier to diffusion at the membrane, and they comprise the 10nm filaments seen with EM | J-protein co-chaperone of the mitochondrial import motor associated with the presequence translocase, with Ssc1p, Tim44p, Mge1p, and Pam16p; stimulates ATPase activity of Ssc1p to drive mitochondrial import; activity is inhibited by Pam16p | Component of both the Nup84 nuclear pore sub-complex and of the COPII complex (Sar1p, Sec13p, Sec16p, Sec23p, Sec24p, Sec31p, Sfb2p, and Sfb3p) which is important for the formation of ER to Golgi transport vesicles | Zeta subunit of the coatomer complex (COPI), which coats Golgi-derived transport vesicles; involved in retrograde transport between Golgi and ER | Possible rhomboid protease, has similarity to eukaryotic rhomboid proteases including Pcp1p |
| 281 | 48 | 449 | 1.007 | 0.597 | 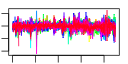 |  | 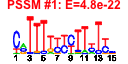 | 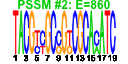 |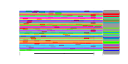  | YBR074W YDL195W YER164W YGR090W YHR206W YIL092W YKL182W YLL048C YNR016C YOR011W YBR075W YBR291C YCR037C YDR291W YDR407C YDR419W YHR085W YHR120W YHR205W YJL039C YJL135W YKL106W YKL116C YKL165C YLR212C YLR214W YLR313C YLR385C YML010W YML046W YML047C YMR319C YNL197C YNL207W YNL221C YNR051C YOR116C YOR127W YOR199W YOR341W YOR342C YOR345C YOR359W YPL030W YPL063W YPR018W YPR069C YPR097W | SEC31 CHD1 UTP22 SKN7 FAS1 BAT1 ACC1 AUS1 CTP1 PHO87 HRQ1 TRS120 RAD30 IPI1 MSH1 SCH9 NUP192 AAT1 PRR1 MCD4 TUB4 FRE1 SPH1 SWC7 SPT5 PRP39 PRM6 FET4 WHI3 RIO2 POP1 BRE5 RPO31 RGA1 RPA190 VTS1 TRM44 TIM50 RLF2 SPE3 | Putative metalloprotease | Essential phosphoprotein component (p150) of the COPII coat of secretory pathway vesicles, in complex with Sec13p; required for ER-derived transport vesicle formation | Nucleosome remodeling factor that functions in regulation of transcription elongation; contains a chromo domain, a helicase domain and a DNA-binding domain; component of both the SAGA and SILK complexes | Possible U3 snoRNP protein involved in maturation of pre-18S rRNA, based on computational analysis of large-scale protein-protein interaction data | Nuclear response regulator and transcription factor, part of a branched two-component signaling system; required for optimal induction of heat-shock genes in response to oxidative stress; involved in osmoregulation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and to the nucleus | Beta subunit of fatty acid synthetase, which catalyzes the synthesis of long-chain saturated fatty acids; contains acetyltransacylase, dehydratase, enoyl reductase, malonyl transacylase, and palmitoyl transacylase activities | Mitochondrial branched-chain amino acid aminotransferase, homolog of murine ECA39; highly expressed during logarithmic phase and repressed during stationary phase | Acetyl-CoA carboxylase, biotin containing enzyme that catalyzes the carboxylation of acetyl-CoA to form malonyl-CoA; required for de novo biosynthesis of long-chain fatty acids | Transporter of the ATP-binding cassette family, involved in uptake of sterols and anaerobic growth | Mitochondrial inner membrane citrate transporter, member of the mitochondrial carrier family | Low-affinity inorganic phosphate (Pi) transporter, involved in activation of PHO pathway; expression is independent of Pi concentration and Pho4p activity; contains 12 membrane-spanning segments | Putative DNA helicase | One of 10 subunits of the transport protein particle (TRAPP) complex of the cis-Golgi which mediates vesicle docking and fusion; involved in endoplasmic reticulum (ER) to Golgi membrane traffic | DNA polymerase eta, involved in the predominantly error-free bypass replication of DNA lesions, catalyzes the efficient and accurate synthesis of DNA opposite cyclobutane pyrimidine dimers; homolog of human POLH and bacterial DinB proteins | Essential component of the Rix1 complex (with Rix1p and Ipi3p) that is required for processing of ITS2 sequences from 35S pre-rRNA; Rix1 complex associates with Mdn1p in pre-60S ribosomal particles | DNA-binding protein of the mitochondria involved in repair of mitochondrial DNA, has ATPase activity and binds to DNA mismatches; has homology to E. coli MutS; transcription is induced during meiosis | Protein kinase involved in transcriptional activation of osmostress-responsive genes; regulates G1 progression, cAPK activity, nitrogen activation of the FGM pathway; involved in life span regulation; homologous to mammalian Akt/PKB | Essential structural subunit of the nuclear pore complex (NPC), localizes to the nuclear periphery of nuclear pores, homologous to human p205 | Mitochondrial aspartate aminotransferase, catalyzes the conversion of oxaloacetate to aspartate in aspartate and asparagine biosynthesis | Serine/threonine protein kinase that inhibits pheromone induced signalling downstream of MAPK, possibly at the level of the Ste12p transcription factor | Protein involved in glycosylphosphatidylinositol (GPI) anchor synthesis; multimembrane-spanning protein that localizes to the endoplasmic reticulum; highly conserved among eukaryotes | Gamma-tubulin, involved in nucleating microtubules from both the cytoplasmic and nuclear faces of the spindle pole body | Ferric reductase and cupric reductase, reduces siderophore-bound iron and oxidized copper prior to uptake by transporters; expression induced by low copper and iron levels | Protein involved in shmoo formation and bipolar bud site selection; homologous to Spa2p, localizes to sites of polarized growth in a cell cycle dependent- and Spa2p-dependent manner, interacts with MAPKKs Mkk1p, Mkk2p, and Ste7p | Protein of unknown function, component of the Swr1p complex that incorporates Htz1p into chromatin | Protein that forms a complex with Spt4p and mediates both activation and inhibition of transcription elongation; Spt4p-Spt5p complex also plays a role in pre-mRNA processing | U1 snRNP protein involved in splicing, contains multiple tetriatricopeptide repeats | Pheromone-regulated protein, predicted to have 2 transmembrane segments; regulated by Ste12p during mating | Low-affinity Fe(II) transporter of the plasma membrane | RNA binding protein that sequesters CLN3 mRNA in cytoplasmic foci; cytoplasmic retention factor for Cdc28p and associated cyclins; regulates cell fate and dose-dependently regulates the critical cell size required for passage through Start | Essential serine kinase involved in the processing of the 20S pre-rRNA into mature 18S rRNA; has similarity to Rio1p | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends; binds to the RPR1 RNA subunit in RNase P | Ubiquitin protease cofactor, forms deubiquitination complex with Ubp3p that coregulates anterograde and retrograde transport between the endoplasmic reticulum and Golgi compartments; null is sensitive to brefeldin A | RNA polymerase III subunit C160, part of core enzyme; similar to bacterial beta-prime subunit | GTPase-activating protein for the polarity-establishment protein Cdc42p; implicated in control of septin organization, pheromone response, and haploid invasive growth | RNA polymerase I subunit; largest subunit of RNA polymerase I | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and the nucleus | Post-transcriptional gene regulator, RNA-binding protein containing a SAM domain; shows genetic interactions with Vti1p, which is a v-SNARE involved in cis-Golgi membrane traffic | Putative tRNA U44 2'-O-methyltransferase; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; non-essential gene | Constituent of the mitochondrial inner membrane presequence translocase (TIM23 complex); may promote binding of incoming precursor proteins to the intermembrane space domain of Tom22p during translocation | Largest subunit (p90) of the Chromatin Assembly Complex (CAF-1) with Cac2p and Msi1p that assembles newly synthesized histones onto recently replicated DNA; involved in the maintenance of transcriptionally silent chromatin | Spermidine synthase, involved in biosynthesis of spermidine and also in biosynthesis of pantothenic acid; spermidine is required for growth of wild-type cells | Protein that contains a Phox homology (PX) domain and binds phosphoinositides; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
| 68 | 52 | 439 | 1.020 | 0.552 |  |  |  |  |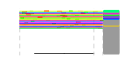  | YAR037W YAR040C YAR061W YAR064W YBL034C YBR100W YCR001W YCR014C YCR047C YCR048W YCR050C YDL210W YIL013C YIL054W YIL132C YIL163C YJL220W YKR032W YKR073C YLR236C YLR402W YLR416C YLR445W YML084W YML089C YML103C YMR122C YMR326C YNL017C YNL143C YOL015W YOL046C YOL132W YOR012W YOR064C YOR082C YOR107W YOR314W YOR379C YPL025C YPL102C YPL130W YPR077C YPR157W YPR195C YAR047C YDL214C YDR011W YMR219W YNL285W YOR139C YPR039W | STU1 MMS4 POL4 BUD23 ARE1 UGA4 PDR11 CSM2 NUP188 IRC10 GAS4 YNG1 RGS2 SPO19 PRR2 SNQ2 ESC1 | Putative protein of unknown function | Component of the mitotic spindle that binds to interpolar microtubules via its association with beta-tubulin (Tub2p); required for interpolar microtubules to provide an outward force on the spindle poles | Subunit of the structure-specific Mms4p-Mus81p endonuclease that cleaves branched DNA; involved in recombination and DNA repair | DNA polymerase IV, undergoes pair-wise interactions with Dnl4p-Lif1p and Rad27p to mediate repair of DNA double-strand breaks by non-homologous end joining (NHEJ); homologous to mammalian DNA polymerase beta | Protein involved in bud-site selection; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Acyl-CoA:sterol acyltransferase, isozyme of Are2p; endoplasmic reticulum enzyme that contributes the major sterol esterification activity in the absence of oxygen | Non-essential protein of unknown function; deletion mutant is synthetically sick or lethal with alpha-synuclein | Permease that serves as a gamma-aminobutyrate (GABA) transport protein involved in the utilization of GABA as a nitrogen source; catalyzes the transport of putrescine and delta-aminolevulinic acid (ALA); localized to the vacuolar membrane | ATP-binding cassette (ABC) transporter, multidrug transporter involved in multiple drug resistance; mediates sterol uptake when sterol biosynthesis is compromisedregulated by Pdr1p; required for anaerobic growth | Protein required for accurate chromosome segregation during meiosis | Putative protein of unknown function; transcription is regulated by Ume6p and induced in response to alpha factor | Subunit of the nuclear pore complex (NPC), involved in the structural organization of the complex and of the nuclear envelope, also involved in nuclear envelope permeability, interacts with Pom152p and Nic96p | hypothetical protein; null mutant displays increased levels of spontaneous Rad52 foci | 1,3-beta-glucanosyltransferase, involved with Gas2p in spore wall assembly; has similarity to Gas1p; localizes to the cell wall | Subunit of the NuA3 histone acetyltransferase complex that acetylates histone H3; contains PHD finger domain that interacts with methylated histone H3, has similarity to the human tumor suppressor ING1 | Negative regulator of glucose-induced cAMP signaling; directly activates the GTPase activity of the heterotrimeric G protein alpha subunit Gpa2p | Meiosis-specific protein of unknown function, involved in completion of nuclear divisions; identified as a weak high-copy suppressor of the spo1-1 ts mutation; putative GPI-dependent cell-wall protein | Serine/threonine protein kinase that inhibits pheromone induced signalling downstream of MAPK, possibly at the level of the Ste12p transcription factor | Plasma membrane ATP-binding cassette (ABC) transporter, multidrug transporter involved in multidrug resistance and resistance to singlet oxygen species | Protein localized to the nuclear periphery, involved in telomeric silencing; interacts with PAD4-domain of Sir4p |
| 82 | 52 | 442 | 1.040 | 0.644 |  |  |  |  |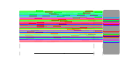  | YAL019W YAL033W YAL039C YBL031W YBR032W YBR190W YBR202W YCR059C YCR063W YDL038C YDL042C YDR024W YDR397C YDR408C YDR445C YDR482C YEL021W YER018C YER019W YFR018C YGL149W YGL235W YGR082W YIL064W YIL065C YIR023W YJL075C YLL052C YLR139C YLR141W YLR400W YLR434C YMR011W YNL210W YNL309W YNR017W YOL101C YOR325W YOR376W YPL080C YBL034C YBL053W YBL070C YCL069W YDL039C YDR301W YDR454C YER072W YGR190C YIL090W YJL119C YPL082C | FUN30 POP5 CYC3 MYO4 MCM7 YIH1 BUD31 SIR2 NCB2 ADE8 CWC21 URA3 SPC25 ISC1 TOM20 FIS1 DAL81 AQY2 SLS1 RRN5 HXT2 MER1 STB1 TIM23 IZH4 YOR376W-A STU1 VBA3 PRM7 CFT1 GUK1 VTC1 ICE2 MOT1 | Protein whose overexpression affects chromosome stability, potential Cdc28p substrate; homolog of Snf2p; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends | Cytochrome c heme lyase (holocytochrome c synthase), attaches heme to apo-cytochrome c (Cyc1p or Cyc7p) in the mitochondrial intermembrane space; human ortholog may have a role in microphthalmia with linear skin defects (MLS) | One of two type V myosin motors (along with MYO2) involved in actin-based transport of cargos; required for mRNA transport, including ASH1 mRNA, and facilitating the growth and movement of ER tubules into the growing bud along with She3p | Component of the hexameric MCM complex, which is important for priming origins of DNA replication in G1 and becomes an active ATP-dependent helicase that promotes DNA melting and elongation when activated by Cdc7p-Dbf4p in S-phase | Protein that inhibits activation of Gcn2p, an eIF2 alpha subunit protein kinase, by competing for Gcn1p binding, thus impacting gene expression in response to starvation; has sequence and functional similarity to the mouse IMPACT gene | Protein involved in bud-site selection; analysis of integrated high-throughput datasets predicts an involvement in RNA splicing; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Putative protein of unknown function; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO) | Conserved NAD+ dependent histone deacetylase of the Sirtuin family involved in regulation of lifespan; plays roles in silencing at HML, HMR, telomeres, and the rDNA locus; negatively regulates initiation of DNA replication | Subunit of a heterodimeric NC2 transcription regulator complex with Bur6p; complex binds to TBP and can repress transcription by preventing preinitiation complex assembly or stimulate activated transcription; homologous to human NC2beta | Phosphoribosyl-glycinamide transformylase, catalyzes a step in the 'de novo' purine nucleotide biosynthetic pathway | Component of a complex containing Cef1p, putatively involved in pre-mRNA splicing; may bind RNA; has similarity to S. pombe Cwf21p | Orotidine-5'-phosphate (OMP) decarboxylase, catalyzes the sixth enzymatic step in the de novo biosynthesis of pyrimidines, converting OMP into uridine monophosphate (UMP); converts 5-FOA into 5-fluorouracil, a toxic compound | Component of the evolutionarily conserved kinetochore-associated Ndc80 complex (Ndc80p-Nuf2p-Spc24p-Spc25p); involved in chromosome segregation, spindle checkpoint activity and kinetochore clustering | Mitochondrial membrane localized inositol phosphosphingolipid phospholipase C, hydrolyzes complex sphingolipids to produce ceramide; activated by phosphatidylserine, cardiolipin, and phosphatidylglycerol; mediates Na+ and Li+ halotolerance | Putative protein of unknown function | Putative protein of unknown function; potential Cdc28p substrate | Component of the TOM (translocase of outer membrane) complex responsible for recognition and initial import steps for all mitochondrially directed proteins; acts as a receptor for incoming precursor proteins | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family | Mitochondrial outer membrane protein involved in membrane fission, required for localization of Dnm1p and Mdv1p during mitochondrial division; mediates ethanol-induced apoptosis and ethanol-induced mitochondrial fragmentation | Positive regulator of genes in multiple nitrogen degradation pathways; contains DNA binding domain but does not appear to bind the dodecanucleotide sequence present in the promoter region of many genes involved in allantoin catabolism | Water channel that mediates the transport of water across cell membranes, only expressed in proliferating cells, controlled by osmotic signals, may be involved in freeze tolerance; disrupted by a stop codon in many S. cerevisiae strains | Mitochondrial membrane protein that coordinates expression of mitochondrially-encoded genes; may facilitate delivery of mRNA to membrane-bound translation machinery | Protein involved in transcription of rDNA by RNA polymerase I; transcription factor, member of UAF (upstream activation factor) family along with Rrn9p and Rrn10p | High-affinity glucose transporter of the major facilitator superfamily, expression is induced by low levels of glucose and repressed by high levels of glucose | Protein with RNA-binding motifs required for meiosis-specific mRNA splicing; required for chromosome pairing and meiotic recombination | Protein with a role in regulation of MBF-specific transcription at Start, phosphorylated by Cln-Cdc28p kinases in vitro; unphosphorylated form binds Swi6p and binding is required for Stb1p function; expression is cell-cycle regulated | Essential protein of the mitochondrial inner membrane, component of the mitochondrial import system | Membrane protein involved in zinc metabolism, member of the four-protein IZH family, expression induced by fatty acids and altered zinc levels; deletion reduces sensitivity to excess zinc; possible role in sterol metabolism | Putative protein of unknown function; identified by fungal homology and RT-PCR | Component of the mitotic spindle that binds to interpolar microtubules via its association with beta-tubulin (Tub2p); required for interpolar microtubules to provide an outward force on the spindle poles | Permease of basic amino acids in the vacuolar membrane | Pheromone-regulated protein, predicted to have one transmembrane segment; promoter contains Gcn4p binding elements | RNA-binding subunit of the mRNA cleavage and polyadenylation factor; involved in poly(A) site recognition and required for both pre-mRNA cleavage and polyadenylation, 51% sequence similarity with mammalian AAUAA-binding subunit of CPSF | Guanylate kinase, converts GMP to GDP; required for growth and mannose outer chain elongation of cell wall N-linked glycoproteins | Vacuolar transporter chaperon (VTC) involved in distributing V-ATPase and other membrane proteins; together with other VTC proteins, forms a heterotetrameric complex that associates with the SNARE Nyv1p and the V0 sector of the V-ATPase | Integral ER membrane protein with type-III transmembrane domains; mutations cause defects in cortical ER morphology in both the mother and daughter cells | Essential abundant protein involved in regulation of transcription, removes Spt15p (TBP) from DNA via its C-terminal ATPase activity, forms a complex with TBP that binds TATA DNA with high affinity but with altered specificity |
| 120 | 48 | 452 | 1.098 | 0.599 |  |  |  |  |  | YAL028AW YAR029W YAR060C YAR062W YBL065W YBL084C YBL097W YBR150C YCRX01W YDL030W YDL049C YDL218W YDL220C YDR250C YDR314C YDR360W YDR515W YGR022C YGR205W YGR241C YGR281W YHR136C YJL150W YJR037W YKL055C YLR176C YLR425W YMR094W YPL110C YPL182C YAR043C YBR178W YDL034W YDL056W YDL242W YDR058C YDR269C YDR283C YDR290W YDR338C YDR408C YER097W YFL019C YGR233C YIL162W YJR120W YOL037C YOR318C | CDC27 BRN1 TBS1 PRP9 KNH1 CDC13 RAD34 SLF1 YAP1802 YOR1 SPL2 OAR1 PEX2 TUS1 CTF13 GDE1 MBP1 TGL2 GCN2 ADE8 PHO81 SUC2 | Member of DUP240 gene family but contains no transmembrane domains; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Subunit of the Anaphase-Promoting Complex/Cyclosome (APC/C), which is a ubiquitin-protein ligase required for degradation of anaphase inhibitors, including mitotic cyclins, during the metaphase/anaphase transition | Essential protein required for chromosome condensation, likely to function as an intrinsic component of the condensation machinery, may influence multiple aspects of chromosome transmission and dynamics | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Subunit of the SF3a splicing factor complex, required for spliceosome assembly; acts after the formation of the U1 snRNP-pre-mRNA complex | Protein with similarity to Kre9p, which is involved in cell wall beta 1,6-glucan synthesis; overproduction suppresses growth defects of a kre9 null mutant | Putative protein of unknown function; YDL218W transcription is regulated by Azf1p and induced by starvation and aerobic conditions | Single stranded DNA-binding protein found at TG1-3 telomere G-tails; regulates telomere replication through recruitment of specific sub-complexes, but the essential function is telomere capping | Protein involved in nucleotide excision repair (NER); homologous to RAD4 | RNA binding protein that associates with polysomes; proposed to be involved in regulating mRNA translation; involved in the copper-dependent mineralization of copper sulfide complexes on cell surface in cells cultured in copper salts | ATP-binding protein of unknown function; crystal structure resembles that of E.coli pantothenate kinase and other small kinases | Protein involved in clathrin cage assembly; binds Pan1p and clathrin; homologous to Yap1801p, member of the AP180 protein family | Plasma membrane ATP-binding cassette (ABC) transporter, multidrug transporter mediates export of many different organic anions including oligomycin | Protein with similarity to cyclin-dependent kinase inhibitors, overproduction suppresses a plc1 null mutation; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Mitochondrial 3-oxoacyl-[acyl-carrier-protein] reductase, may comprise a type II mitochondrial fatty acid synthase along with Mct1p | RING-finger peroxin, peroxisomal membrane protein with a C-terminal zinc-binding RING domain, forms translocation subcomplex with Pex10p and Pex12p which functions in peroxisomal matrix protein import | Guanine nucleotide exchange factor (GEF) that functions to modulate Rho1p activity as part of the cell integrity signaling pathway; multicopy suppressor of tor2 mutation and ypk1 ypk2 double mutation; potential Cdc28p substrate | Subunit of the CBF3 complex, which binds to the CDE III element of centromeres, bending the DNA upon binding, and may be involved in sister chromatid cohesion during mitosis | Glycerophosphocholine (GroPCho) phosphodiesterase; hydrolyzes GroPCho to choline and glycerolphosphate, for use as a phosphate source and as a precursor for phosphocholine synthesis; may interact with ribosomes | Transcription factor involved in regulation of cell cycle progression from G1 to S phase, forms a complex with Swi6p that binds to MluI cell cycle box regulatory element in promoters of DNA synthesis genes | Protein with lipolytic activity towards triacylglycerols and diacylglycerols when expressed in E. coli; role in yeast lipid degradation is unclear | Protein kinase, phosphorylates the alpha-subunit of translation initiation factor eIF2 (Sui2p) in response to starvation; activated by uncharged tRNAs and the Gcn1p-Gcn20p complex | hypothetical protein | Phosphoribosyl-glycinamide transformylase, catalyzes a step in the 'de novo' purine nucleotide biosynthetic pathway | Cyclin-dependent kinase (CDK) inhibitor, regulates Pho80p-Pho85p and Pcl7p-Pho85p cyclin-CDK complexes in response to phosphate levels; inhibitory activity for Pho80p-Pho85p requires myo-D-inositol heptakisphosphate (IP7) generated by Vip1p | Invertase, sucrose hydrolyzing enzyme; a secreted, glycosylated form is regulated by glucose repression, and an intracellular, nonglycosylated enzyme is produced constitutively | Protein of unknown function; essential for growth under anaerobic conditions; mutation causes decreased expression of ATP2, impaired respiration, defective sterol uptake, and altered levels/localization of ABC transporters Aus1p and Pdr11p |
| 296 | 40 | 472 | 1.130 | 0.595 |  |  |  | 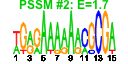 |  | YNL221C YCL058C YCLX01W YDL151C YDL213C YDR110W YDR121W YDR279W YDR528W YER042W YHR040W YIL003W YIL047C YJL011C YJL212C YJR055W YJR129C YKL027W YKL083W YKL125W YLL012W YLL022C YLR400W YML108W YMR177W YMR301C YNL095C YNL149C YNL228W YNL240C YNR046W YOL054W YOL109W YOR006C YPL013C YPL108W YPL193W YPL239W YPR136C YPR169W | POP1 NOP6 FOB1 DPB4 RNH202 HLR1 MXR1 BCD1 CFD1 SYG1 RPC17 OPT1 HIT1 RRN3 YEH1 HIF1 MMT1 ATM1 PGA2 NAR1 TRM112 PSH1 ZEO1 TSR3 MRPS16 RSA1 YAR1 JIP5 | Subunit of both RNase MRP, which cleaves pre-rRNA, and nuclear RNase P, which cleaves tRNA precursors to generate mature 5' ends; binds to the RPR1 RNA subunit in RNase P | Putative RNA-binding protein implicated in ribosome biogenesis; contains an RNA recognition motif (RRM) and has similarity to hydrophilins; NOP6 may be a fungal-specific gene as no homologs have been yet identified in higher eukaryotes | Nucleolar protein required for DNA replication fork blocking and recombinational hotspot activities; binds to the replication fork barrier site in the rDNA region; related to retroviral integrases | Shared subunit of DNA polymerase (II) epsilon and of ISW2/yCHRAC chromatin accessibility complex; involved in both chromosomal DNA replication and in inheritance of telomeric silencing | Ribonuclease H2 subunit, required for RNase H2 activity | Protein involved in regulation of cell wall composition and integrity and response to osmotic stress; overproduction suppresses a lysis sensitive PKC mutation; similar to Lre1p, which functions antagonistically to protein kinase A | Peptide methionine sulfoxide reductase, reverses the oxidation of methionine residues; involved in oxidative damage repair, providing resistance to oxidative stress and regulation of lifespan | Essential protein required for the accumulation of box C/D snoRNA | Highly conserved, iron-sulfur cluster binding protein localized in the cytoplasm; forms a complex with Nbp35p that is involved in iron-sulfur protein assembly in the cytosol | Plasma membrane protein of unknown function; truncation and overexpression suppresses lethality of G-alpha protein deficiency | RNA polymerase III subunit C17; physically interacts with C31, C11, and TFIIIB70; may be involved in the recruitment of pol III by the preinitiation complex | Proton-coupled oligopeptide transporter of the plasma membrane; also transports glutathione and phytochelatin; member of the OPT family | Protein of unknown function, required for growth at high temperature | Putative protein of unknown function; predicted S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Protein of unknown function, localized to the mitochondrial outer membrane | Protein required for transcription of rDNA by RNA polymerase I; transcription factor independent of DNA template; involved in recruitment of RNA polymerase I to rDNA | Steryl ester hydrolase, one of three gene products (Yeh1p, Yeh2p, Tgl1p) responsible for steryl ester hydrolase activity and involved in sterol homeostasis; localized to lipid particle membranes | Non-essential component of the HAT-B histone acetyltransferase complex (Hat1p-Hat2p-Hif1p), localized to the nucleus; has a role in telomeric silencing | Putative protein of unknown function whose structure defines a new subfamily of the split beta-alpha-beta sandwiches; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YML108W is not an essential gene | Putative metal transporter involved in mitochondrial iron accumulation; closely related to Mmt2p | Mitochondrial inner membrane ATP-binding cassette (ABC) transporter, exports mitochondrially synthesized precursors of iron-sulfur (Fe/S) clusters to the cytosol | Putative protein of unknown function predicted to contain a transmembrane domain; YNL095C is not an essential gene | Essential protein required for maturation of Gas1p and Pho8p; involved in protein trafficking; GFP-fusion protein localizes to the ER and YFP-fusion protein to the nuclear envelope-ER network; null mutants have a cell separation defect | Nuclear architecture related protein; component of the cytosolic iron-sulfur (FeS) protein assembly machinery, required for maturation of cytosolic and nuclear FeS proteins; homologous to human Narf | Subunit of an adoMet-dependent tRNA methyltransferase (MTase) complex (Trm11p-Trm112p), required for the methylation of the guanosine nucleotide at position 10 (m2G10) in tRNAs; putative zinc binding subunit of other MTase-related proteins | Nuclear protein, putative RNA polymerase II elongation factor; isolated as Pob3p/Spt16p-binding protein | Peripheral membrane protein of the plasma membrane that interacts with Mid2p; regulates the cell integrity pathway mediated by Pkc1p and Slt2p; the authentic protein is detected in a phosphorylated state in highly purified mitochondria | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus; Null mutant accumulates 20S pre-rRNA | Mitochondrial ribosomal protein of the small subunit | Cytoplasmic protein of unknown function; non-essential gene that is induced in a GDH1 deleted strain with altered redox metabolism; GFP-fusion protein is induced in response to the DNA-damaging agent MMS | Protein involved in the assembly of 60S ribosomal subunits; functionally interacts with Dbp6p; functions in a late nucleoplasmic step of the assembly | Cytoplasmic ankyrin-repeat containing protein of unknown function, proposed to link the processes of 40S ribosomal subunit biogenesis and adaptation to osmotic and oxidative stress; expression repressed by heat shock | Nucleolar protein of unknown function, exhibits a physical interaction with Bre1p |
| 156 | 43 | 448 | 1.158 | 0.638 |  |  |  |  |  | YAL053W YBR043C YBR071W YDR005C YDR210W YER062C YGR032W YIL023C YJL217W YJR089W YKL218C YKR012C YLR007W YLR099C YLR231C YLR348C YLR349W YMR238W YMR295C YNL046W YNL190W YNL192W YOL147C YOR327C YPR106W YPR124W YBR090C YDR134C YDR378C YER041W YGL056C YGR049W YIR035C YJL079C YJL094C YJL211C YLR094C YNR055C YOL103W YOR105W YPL066W YPL067C YPL233W | FLC2 QDR3 MAF1 HOR2 GSC2 YKE4 BIR1 SRY1 NSE1 ICT1 BNA5 DIC1 DFG5 CHS1 PEX11 SNC2 ISR1 HNM1 NHP6B LSM6 YEN1 SDS23 SCM4 PRY1 KHA1 GIS3 HOL1 YOL103W-B NSL1 | Putative FAD transporter; required for uptake of FAD into endoplasmic reticulum; involved in cell wall maintenance | Multidrug transporter of the major facilitator superfamily, required for resistance to quinidine, barban, cisplatin, and bleomycin | Putative protein of unknown function; (GFP)-fusion and epitope-tagged proteins localize to the cytoplasm; mRNA expression may be regulated by the cell cycle and/or cell wall stress | Negative regulator of RNA polymerase III; component of several signaling pathways that repress polymerase III transcription in response to changes in cellular environment; targets the initiation factor TFIIIB | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery | One of two redundant DL-glycerol-3-phosphatases (RHR2/GPP1 encodes the other) involved in glycerol biosynthesis; induced in response to hyperosmotic stress and oxidative stress, and during the diauxic transition | Catalytic subunit of 1,3-beta-glucan synthase, involved in formation of the inner layer of the spore wall; activity positively regulated by Rho1p and negatively by Smk1p; has similarity to an alternate catalytic subunit, Fks1p (Gsc1p) | Zinc transporter; localizes to the ER; null mutant is sensitive to calcofluor white, leads to zinc accumulation in cytosol; ortholog of the mouse KE4 and member of the ZIP (ZRT, IRT-like Protein) family | Cytoplasmic protein of unknown function; expression induced by calcium shortage and via the copper sensing transciption factor Mac1p during conditons of copper deficiency; mRNA is cell cycle regulated, peaking in G1 phase | Essential chromosomal passenger protein involved in coordinating cell cycle events for proper chromosome segregation; C-terminal region binds Sli15p, and the middle region, upon phosphorylation, localizes Cbf2p to the spindle at anaphase | 3-hydroxyaspartate dehydratase, deaminates L-threo-3-hydroxyaspartate to form oxaloacetate and ammonia; required for survival in the presence of hydroxyaspartate | Essential subunit of the Mms21-Smc5-Smc6 complex; nuclear protein required for DNA repair and growth | Protein of unknown function, null mutation leads to an increase in sensitivity to Calcofluor white; expression of the gene is induced in the presence of isooctane | Kynureninase, required for biosynthesis of nicotinic acid from tryptophan via kynurenine pathway | Mitochondrial dicarboxylate carrier, integral membrane protein, catalyzes a dicarboxylate-phosphate exchange across the inner mitochondrial membrane, transports cytoplasmic dicarboxylates into the mitochondrial matrix | Putative mannosidase, essential glycosylphosphatidylinositol (GPI)-anchored membrane protein required for cell wall biogenesis in bud formation, involved in filamentous growth, homologous to Dcw1p | Protein of unknown function that associates with ribosomes; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery and bud; YMR295C is not an essential gene | Putative protein of unknown function; expression depends on Swi5p; GFP-fusion protein localizes to the endoplasmic reticulum; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO) | Cell wall protein of unknown function; proposed role as a hydrophilin induced by osmotic stress; contains a putative GPI-attachment site | Chitin synthase I, requires activation from zymogenic form in order to catalyze the transfer of N-acetylglucosamine (GlcNAc) to chitin; required for repairing the chitin septum during cytokinesis; transcription activated by mating factor | Peroxisomal membrane protein required for peroxisome proliferation and medium-chain fatty acid oxidation, most abundant protein in the peroxisomal membrane, regulated by Adr1p and Pip2p-Oaf1p, promoter contains ORE and UAS1-like elements | Vesicle membrane receptor protein (v-SNARE) involved in the fusion between Golgi-derived secretory vesicles with the plasma membrane; member of the synaptobrevin/VAMP family of R-type v-SNARE proteins | Predicted protein kinase, overexpression causes sensitivity to staurosporine, which is a potent inhibitor of protein kinase C | Choline/ethanolamine transporter (permease) that also controls the uptake of nitrogen mustard; expression is co-regulated with phospholipid biosynthetic genes and negatively regulated by choline and myo-inositol | High-mobility group non-histone chromatin protein, functionally redundant with Nhp6Ap; homologous to mammalian high mobility group proteins 1 and 2; acts to recruit transcription factor Rcs1p to certain promoters | Lsm (Like Sm) protein; part of heteroheptameric complexes (Lsm2p-7p and either Lsm1p or 8p): cytoplasmic Lsm1p complex involved in mRNA decay; nuclear Lsm8p complex part of U6 snRNP and possibly involved in processing tRNA, snoRNA, and rRNA | Protein of unknown function, has similarity to endonuclease Rth1p; potentially phosphorylated by Cdc28p | One of two S. cerevisiae homologs (Sds23p and Sds24p) of the Schizosaccharomyces pombe Sds23 protein, which genetic studies have implicated in APC/cyclosome regulation | Potential regulatory effector of CDC4 function, suppresses a temperature-sensitive allele of CDC4, tripartite protein structure in which a charged region separates two uncharged domains, not essential for mitosis or meiosis | Putative cytoplasmic protein of unknown function | Protein of unknown function, has similarity to Pry2p and Pry3p and to the plant PR-1 class of pathogen related proteins | Putative K+/H+ antiporter with a probable role in intracellular cation homeostasis, localized to Golgi vesicles and detected in highly purified mitochondria in high-throughput studies | Protein of unknown function | Putative transporter in the major facilitator superfamily (DHA1 family) of multidrug resistance transporters; mutations in membrane-spanning domains permit cation and histidinol uptake | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the bud neck and cytoplasm; null mutant is viable and exhibits growth defect on a non-fermentable (respiratory) carbon source | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm; YPL067C is not an essential gene | Essential component of the MIND kinetochore complex (Mtw1p Including Nnf1p-Nsl1p-Dsn1p) which joins kinetochore subunits contacting DNA to those contacting microtubules; required for accurate chromosome segregation |
| 117 | 48 | 446 | 1.165 | 0.664 |  |  |  |  |  | YAL062W YBL069W YBR004C YBR160W YCL062W YCL063W YCR046C YDL132W YDL163W YDR326C YDR407C YER042W YGL063W YHR046C YHR060W YJL194W YJR112W YKL077W YKR013W YLR042C YLR322W YLR380W YML108W YMR039C YMR245W YNL057W YNL191W YNL225C YNR027W YNR042W YOL027C YOL070C YOL090W YOR196C YOR197W YOR348C YPR031W YCL064C YDR222W YDR492W YDR518W YFR005C YHR011W YKR079C YML027W YML103C YNL078W YPL158C | GDH3 AST1 GPI18 CDC28 VAC17 IMG1 CDC53 YSP2 TRS120 MXR1 PUS2 INM1 VMA22 CDC6 NNF1 PRY2 CSR1 SUB1 DUG3 CNM67 BUD17 MDM38 NBA1 MSH2 LIP5 MCA1 PUT4 NTO1 CHA1 IZH1 EUG1 SAD1 DIA4 TRZ1 YOX1 NUP188 NIS1 AIM44 | NADP(+)-dependent glutamate dehydrogenase, synthesizes glutamate from ammonia and alpha-ketoglutarate; rate of alpha-ketoglutarate utilization differs from Gdh1p; expression regulated by nitrogen and carbon sources | Peripheral membrane protein that interacts with the plasma membrane ATPase Pma1p and has a role in its targeting to the plasma membrane, possibly by influencing its incorporation into lipid rafts | Functional ortholog of human PIG-V, which is a mannosyltransferase that transfers the second mannose in glycosylphosphatidylinositol biosynthesis; the authentic, non-tagged protein was localized to mitochondria | Catalytic subunit of the main cell cycle cyclin-dependent kinase (CDK); alternately associates with G1 cyclins (CLNs) and G2/M cyclins (CLBs) which direct the CDK to specific substrates | Protein involved in vacuole inheritance; acts as a vacuole-specific receptor for myosin Myo2p | Mitochondrial ribosomal protein of the large subunit, required for respiration and for maintenance of the mitochondrial genome | Cullin, structural protein of SCF complexes (which also contain Skp1p, Cdc34p, Hrt1p and an F-box protein) involved in ubiquitination; SCF promotes the G1-S transition by targeting G1 cyclins and the Cln-CDK inhibitor Sic1p for degradation | Protein involved in programmed cell death; mutant shows resistance to cell death induced by amiodarone or intracellular acidification | One of 10 subunits of the transport protein particle (TRAPP) complex of the cis-Golgi which mediates vesicle docking and fusion; involved in endoplasmic reticulum (ER) to Golgi membrane traffic | Peptide methionine sulfoxide reductase, reverses the oxidation of methionine residues; involved in oxidative damage repair, providing resistance to oxidative stress and regulation of lifespan | Mitochondrial tRNA:pseudouridine synthase; acts at positions 27 and 28, but not at position 72; efficiently and rapidly targeted to mitochondria, specifically dedicated to mitochondrial tRNA modification | Inositol monophosphatase, involved in biosynthesis of inositol and in phosphoinositide second messenger signaling; INM1 expression increases in the presence of inositol and decreases upon exposure to antibipolar drugs lithium and valproate | Integral membrane protein that is required for vacuolar H+-ATPase (V-ATPase) function, although not an actual component of the V-ATPase complex; functions in the assembly of the V-ATPase; localized to the yeast endoplasmic reticulum (ER) | Essential ATP-binding protein required for DNA replication, component of the pre-replicative complex (pre-RC) which requires ORC to associate with chromatin and is in turn required for Mcm2-7p DNA association; homologous to S. pombe Cdc18p | Essential component of the MIND kinetochore complex (Mtw1p Including Nnf1p-Nsl1p-Dsn1p) which joins kinetochore subunits contacting DNA to those contacting microtubules; required for accurate chromosome segregation | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Protein of unknown function, has similarity to Pry1p and Pry3p and to the plant PR-1 class of pathogen related proteins | Protein of unknown function; localizes to the cytoplasm; YLL042C is not an essential gene | Phosphatidylinositol transfer protein with a potential role in lipid turnover; interacts specifically with thioredoxin peroxidase (Tsa2p) and may have a role in oxidative stress resistance | Putative protein of unknown function whose structure defines a new subfamily of the split beta-alpha-beta sandwiches; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and nucleus; YML108W is not an essential gene | Transcriptional coactivator, facilitates elongation by influencing enzymes that modify RNAP II, acts in a peroxide resistance pathway involving Rad2p; suppressor of TFIIB mutations | Probable glutamine amidotransferase, forms a complex with Dug1p and Dug2p to degrade glutathione (GSH) and other peptides containing a gamma-glu-X bond in an alternative pathway to GSH degradation by gamma-glutamyl transpeptidase (Ecm38p) | Component of the spindle pole body outer plaque; required for spindle orientation and mitotic nuclear migration | Protein involved in bud-site selection; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Mitochondrial inner membrane protein, involved in membrane integration of a subset of mitochondrial proteins; required for K+/H+ exchange; associates with mitochondrial ribosomes; human ortholog Letm1 implicated in Wolf-Hirschhorn syndrome | Protein of unknown function; may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cell periphery, cytoplasm, and bud neck; potential Cdc28p substrate | Protein that forms heterodimers with Msh3p and Msh6p that bind to DNA mismatches to initiate the mismatch repair process; contains a Walker ATP-binding motif required for repair activity; Msh2p-Msh6p binds to and hydrolyzes ATP | Protein involved in biosynthesis of the coenzyme lipoic acid, has similarity to E. coli lipoic acid synthase | Putative cysteine protease similar to mammalian caspases, involved in regulation of apoptosis upon hydrogen peroxide treatment | Proline permease, required for high-affinity transport of proline; also transports the toxic proline analog azetidine-2-carboxylate (AzC); PUT4 transcription is repressed in ammonia-grown cells | Subunit of the NuA3 histone acetyltransferase complex that acetylates histone H3; contains PHD finger domain that interacts with methylated histone H3 | Catabolic L-serine (L-threonine) deaminase, catalyzes the degradation of both L-serine and L-threonine; required to use serine or threonine as the sole nitrogen source, transcriptionally induced by serine and threonine | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Membrane protein involved in zinc metabolism, member of the four-protein IZH family; transcription is regulated directly by Zap1p, expression induced by zinc deficiency and fatty acids; deletion increases sensitivity to elevated zinc | Protein disulfide isomerase of the endoplasmic reticulum lumen, function overlaps with that of Pdi1p; may interact with nascent polypeptides in the ER | Conserved zinc-finger domain protein involved in pre-mRNA splicing, required for assembly of U4 snRNA into the U4/U6 particle | Probable mitochondrial seryl-tRNA synthetase, mutant displays increased invasive and pseudohyphal growth | tRNase Z, involved in RNA processing, has two putative nucleotide triphosphate-binding motifs (P-loop) and a conserved histidine motif, homolog of the human candidate prostate cancer susceptibility gene ELAC2 | Homeodomain-containing transcriptional repressor, binds to Mcm1p and to early cell cycle boxes (ECBs) in the promoters of cell cycle-regulated genes expressed in M/G1 phase; expression is cell cycle-regulated; potential Cdc28p substrate | Subunit of the nuclear pore complex (NPC), involved in the structural organization of the complex and of the nuclear envelope, also involved in nuclear envelope permeability, interacts with Pom152p and Nic96p | Protein localized in the bud neck at G2/M phase; physically interacts with septins; possibly involved in a mitotic signaling network | Protein of unknown function; null mutant displays increased frequency of mitochondrial genome loss (petite formation) and reduced growth rate in minimal glycerol media |
| 284 | 44 | 444 | 1.176 | 0.644 | 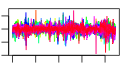 |  | 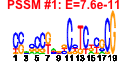 | 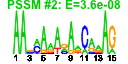 |  | YPL114W YAR014C YBL014C YBR231C YDL010W YDL129W YDL207W YDL211C YEL076W-C YFR034C YGL089C YHR076W YHR146W YIL064W YIL065C YJL012C YJL194W YKL053W YKL070W YKL112W YKL202W YKR031C YLR236C YLR374C YLR380W YLR458W YML098W YMR039C YMR245W YNR032W YNR047W YOL005C YOL015W YOR010C YOR213C YOR298W YOR324C YOR373W YPL024W YPL124W YPL182C YPR034W YPR054W YPR076W | BUD14 RRN6 SWC5 GRX6 GLE1 PHO4 MF(ALPHA)2 PTC7 CRP1 FIS1 VTC4 CDC6 ABF1 SPO14 CSR1 TAF13 SUB1 PPG1 RPB11 IRC10 TIR2 SAS5 MUM3 FRT1 TRM2 RMI1 SPC29 ARP7 SMK1 | Protein involved in bud-site selection, Bud14p-Glc7p complex functions as a cortical regulator of dynein; diploid mutants display a random budding pattern instead of the wild-type bipolar pattern | Protein involved in the transcription of 35S rRNA genes by RNA polymerase I; component of the core factor (CF) complex also composed of Rrn11p, Rrn7p and TATA-binding protein | Protein of unknown function, component of the SWR1 complex, which exchanges histone variant H2AZ (Htz1p) for chromatin-bound histone H2A | Monothiol glutaredoxin that binds an iron-sulfur cluster; more similar in activity to dithiol than other monothiol glutaredoxins; forms homodimers; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm and the nucleus; YDL129W is not an essential gene | Cytoplasmic nucleoporin required for polyadenylated RNA export but not for protein import; component of Nup82p nuclear pore subcomplex; contains a nuclear export signal | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the vacuole | Basic helix-loop-helix (bHLH) transcription factor of the myc-family; binds cooperatively with Pho2p to the PHO5 promoter; function is regulated by phosphorylation at multiple sites and by phosphate availability | Mating pheromone alpha-factor, made by alpha cells; interacts with mating type a cells to induce cell cycle arrest and other responses leading to mating; also encoded by MF(ALPHA)1, which is more highly expressed than MF(ALPHA)2 | Mitochondrially localized type 2C protein phosphatase; expression induced by growth on ethanol and by sustained osmotic stress; possible role in carbon source utilization in low oxygen environments | Protein that binds to cruciform DNA structures | Putative S-adenosylmethionine-dependent methyltransferase of the seven beta-strand family | Mitochondrial outer membrane protein involved in membrane fission, required for localization of Dnm1p and Mdv1p during mitochondrial division; mediates ethanol-induced apoptosis and ethanol-induced mitochondrial fragmentation | Vacuolar membrane protein involved in vacuolar polyphosphate accumulation; functions as a regulator of vacuolar H+-ATPase activity and vacuolar transporter chaperones; involved in non-autophagic vacuolar fusion | Essential ATP-binding protein required for DNA replication, component of the pre-replicative complex (pre-RC) which requires ORC to associate with chromatin and is in turn required for Mcm2-7p DNA association; homologous to S. pombe Cdc18p | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | DNA binding protein with possible chromatin-reorganizing activity involved in transcriptional activation, gene silencing, and DNA replication and repair | Phospholipase D, catalyzes the hydrolysis of phosphatidylcholine, producing choline and phosphatidic acid; involved in Sec14p-independent secretion; required for meiosis and spore formation; differently regulated in secretion and meiosis | Phosphatidylinositol transfer protein with a potential role in lipid turnover; interacts specifically with thioredoxin peroxidase (Tsa2p) and may have a role in oxidative stress resistance | TFIID subunit (19 kDa), involved in RNA polymerase II transcription initiation, similar to histone H4 with atypical histone fold motif of Spt3-like transcription factors | Transcriptional coactivator, facilitates elongation by influencing enzymes that modify RNAP II, acts in a peroxide resistance pathway involving Rad2p; suppressor of TFIIB mutations | Putative serine/threonine protein phosphatase, required for glycogen accumulation; interacts with Tap42p, which binds to and regulates other protein phosphatases | Putative protein kinase that, when overexpressed, interferes with pheromone-induced growth arrest; localizes to the cytoplasm; potential Cdc28p substrate | RNA polymerase II subunit B12.5; part of central core; similar to Rpc19p and bacterial alpha subunit | hypothetical protein; null mutant displays increased levels of spontaneous Rad52 foci | Putative cell wall mannoprotein of the Srp1p/Tip1p family of serine-alanine-rich proteins; transcription is induced by cold shock and anaerobiosis | Subunit of the SAS complex (Sas2p, Sas4p, Sas5p), which acetylates free histones and nucleosomes and regulates transcriptional silencing; stimulates Sas2p HAT activity | Protein of unknown function involved in the organization of the outer spore wall layers; has similarity to the tafazzins superfamily of acyltransferases | Tail-anchored endoplasmic reticulum membrane protein that is a substrate of the phosphatase calcineurin, interacts with homolog Frt2p, promotes cell growth in conditions of high Na+, alkaline pH, and cell wall stress | tRNA methyltransferase, 5-methylates the uridine residue at position 54 of tRNAs and may also have a role in tRNA stabilization or maturation; endo-exonuclease with a role in DNA repair | Involved in response to DNA damage; null mutants have increased rates of recombination and delayed S phase; interacts physically and genetically with Sgs1p (RecQ family member) and Top3p (topoisomerase III) | Inner plaque spindle pole body (SPB) component, links the central plaque component Spc42p to the inner plaque component Spc110p; required for SPB duplication | Component of both the SWI/SNF and RSC chromatin remodeling complexes; actin-related protein involved in transcriptional regulation | Middle sporulation-specific mitogen-activated protein kinase (MAPK) required for production of the outer spore wall layers; negatively regulates activity of the glucan synthase subunit Gsc2p |
| 232 | 40 | 450 | 1.187 | 0.597 | 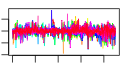 |  | 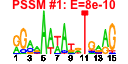 | 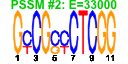 |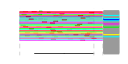  | YEL067C YGR126W YIR042C YJL022W YLL015W YLL016W YLR123C YOR363C YPR014C YBR066C YBR226C YBR240C YCL027W YDL049C YDR253C YDR255C YDR336W YER028C YER188W YFR022W YGL249W YGR016W YGR032W YGR070W YGR226C YGR238C YHL010C YIL086C YIL170W YIL171W YIR032C YJL007C YJL038C YLL017W YLL054C YLR053C YLR081W YNL034W YOR177C YOR394W | BPT1 PIP2 NRG2 THI2 FUS1 KNH1 MET32 RMD5 MIG3 ROG3 ZIP2 GSC2 ROM1 KEL2 DAL3 GAL2 MPC54 PAU21 | Putative protein of unknown function; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and the nucleus and is induced in response to the DNA-damaging agent MMS | Putative protein of unknown function; YIR042C is a non-essential gene | ABC type transmembrane transporter of MRP/CFTR family, found in vacuolar membrane, involved in the transport of unconjugated bilirubin and in heavy metal detoxification via glutathione conjugates, along with Ycf1p | Autoregulatory oleate-specific transcriptional activator of peroxisome proliferation, contains Zn(2)-Cys(6) cluster domain, forms heterodimer with Oaf1p, binds oleate response elements (OREs), activates beta-oxidation genes | Transcriptional repressor that mediates glucose repression and negatively regulates filamentous growth; has similarity to Nrg1p | Zinc finger protein of the Zn(II)2Cys6 type, probable transcriptional activator of thiamine biosynthetic genes | Membrane protein localized to the shmoo tip, required for cell fusion; expression regulated by mating pheromone; proposed to coordinate signaling, fusion, and polarization events required for fusion; potential Cdc28p substrate | Protein with similarity to Kre9p, which is involved in cell wall beta 1,6-glucan synthesis; overproduction suppresses growth defects of a kre9 null mutant | Zinc-finger DNA-binding protein, involved in transcriptional regulation of the methionine biosynthetic genes, similar to Met31p | Cytosolic protein required for sporulation; also required for the ubiquitination of the gluconeogenetic enzyme fructose-1,6-bisphosphatase, which is degraded rapidly after the switch from gluconeogenesis to glycolysis | Putative protein of unknown function; sumoylated under stress conditions in a genome wide study; YDR336W is not an essential gene | Probable transcriptional repressor involved in response to toxic agents such as hydroxyurea that inhibit ribonucleotide reductase; phosphorylation by Snf1p or the Mec1p pathway inactivates Mig3p, allowing induction of damage response genes | Protein that binds to Rsp5p, which is a hect-type ubiquitin ligase, via its 2 PY motifs; has similarity to Rod1p; mutation suppresses the temperature sensitivity of an mck1 rim11 double mutant | Meiosis-specific protein involved in normal synaptonemal complex formation and pairing between homologous chromosomes during meiosis | Putative protein of unknown function | Catalytic subunit of 1,3-beta-glucan synthase, involved in formation of the inner layer of the spore wall; activity positively regulated by Rho1p and negatively by Smk1p; has similarity to an alternate catalytic subunit, Fks1p (Gsc1p) | GDP/GTP exchange protein (GEP) for Rho1p; mutations are synthetically lethal with mutations in rom2, which also encodes a GEP | Protein that functions in a complex with Kel1p to negatively regulate mitotic exit, interacts with Tem1p and Lte1p; localizes to regions of polarized growth; potential Cdc28p substrate | Putative protein of unknown function, contains a zinc finger region and has homology to human BRAP2, which is a cytoplasmic protein that binds nuclear localization sequences | Ureidoglycolate hydrolase, converts ureidoglycolate to glyoxylate and urea in the third step of allantoin degradation; expression sensitive to nitrogen catabolite repression | Putative protein of unknown function; expression induced during sporulation and repressed during vegetative growth by Sum1p and Hst1p; similar to adjacent open reading frame, YJL037W | Putative protein of unknown function with similarity to Pip2p, an oleate-specific transcriptional activator of peroxisome proliferation; YLL054C is not an essential gene | Galactose permease, required for utilization of galactose; also able to transport glucose | Putative protein of unknown function; YNL034W is not an essential gene | Component of the meiotic outer plaque, a membrane-organizing center which is assembled on the cytoplasmic face of the spindle pole body during meiosis II and triggers the formation of the prospore membrane; potential Cdc28p substrate | hypothetical protein |
| 252 | 50 | 452 | 1.296 | 0.607 |  |  | 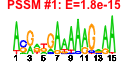 | 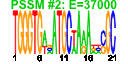 |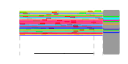  | YCR018C YCR037C YDL093W YDL183C YGL075C YGR035C YLR032W YLR385C YOR083W YAR002AC YBL044W YBR027C YBR134W YBR186W YBR292C YBR296C YCL022C YCL023C YCL039W YCRX06W YCRX07W YCRX08W YDL187C YDL239C YDL241W YDR111C YDR112W YDR135C YDR160W YDR274C YDR281C YDR488C YDR537C YDR538W YEL045C YER085C YFR054C YGL126W YGL209W YGR069W YGR164W YHL037C YHL045W YHR035W YHR130C YHR136C YIL012W YIL150C YJL058C YMR244W | SRD1 PHO87 PMT5 MPS2 RAD5 SWC7 WHI5 PCH2 PHO89 GID7 ADY3 ALT2 YCF1 SSY1 PHM6 PAC11 PAD1 SCS3 MIG2 SPL2 MCM10 BIT61 | Protein involved in the processing of pre-rRNA to mature rRNA; contains a C2/C2 zinc finger motif; srd1 mutation suppresses defects caused by the rrp1-1 mutation | Low-affinity inorganic phosphate (Pi) transporter, involved in activation of PHO pathway; expression is independent of Pi concentration and Pho4p activity; contains 12 membrane-spanning segments | Protein O-mannosyltransferase, transfers mannose residues from dolichyl phosphate-D-mannose to protein serine/threonine residues; acts in a complex with Pmt3p, can instead interact with Pmt2p in some conditions; target for new antifungals | Putative protein of unknown function; YDL183C is not an essential gene | Essential membrane protein localized at the nuclear envelope and spindle pole body (SPB), required for insertion of the newly duplicated SPB into the nuclear envelope; potentially phosphorylated by Cdc28p | Putative protein of unknown function, potential Cdc28p substrate; transcription is activated by paralogous transcription factors Yrm1p and Yrr1p along with genes involved in multidrug resistance | DNA helicase proposed to promote replication fork regression during postreplication repair by template switching; contains RING finger domain | Protein of unknown function, component of the Swr1p complex that incorporates Htz1p into chromatin | Repressor of G1 transcription that binds to SCB binding factor (SBF) at SCB target promoters in early G1; phosphorylation of Whi5p by the CDK, Cln3p/Cdc28p relieves repression and promoter binding by Whi5; periodically expressed in G1 | Putative protein of unknown function; YBL044W is not an essential protein | Nucleolar component of the pachytene checkpoint, which prevents chromosome segregation when recombination and chromosome synapsis are defective; also represses meiotic interhomolog recombination in the rDNA | Na+/Pi cotransporter, active in early growth phase; similar to phosphate transporters of Neurospora crassa; transcription regulated by inorganic phosphate concentrations and Pho4p | Protein of unknown function, involved in proteasome-dependent catabolite inactivation of fructose-1,6-bisphosphatase; contains six WD40 repeats; computational analysis suggests that Gid7p and Moh1p have similar functions | Protein required for spore wall formation, thought to mediate assembly of a Don1p-containing structure at the leading edge of the prospore membrane via interaction with spindle pole body components; potentially phosphorylated by Cdc28p | Putative protein of unknown function; YDL241W is not an essential gene | Putative alanine transaminase (glutamic pyruvic transaminase) | Vacuolar glutathione S-conjugate transporter of the ATP-binding cassette family, has a role in detoxifying metals such as cadmium, mercury, and arsenite; also transports unconjugated bilirubin; similar to human cystic fibrosis protein CFTR | Component of the SPS plasma membrane amino acid sensor system (Ssy1p-Ptr3p-Ssy5p), which senses external amino acid concentration and transmits intracellular signals that result in regulation of expression of amino acid permease genes | Protein of unknown function, expression is regulated by phosphate levels | Dynein intermediate chain, acts in the cytoplasmic dynein pathway, forms cortical cytoplasmic microtubule capture site with Num1p; null mutant is defective in nuclear migration, essential in the absence of CIN8 | Phenylacrylic acid decarboxylase, confers resistance to cinnamic acid, decarboxylates aromatic carboxylic acids to the corresponding vinyl derivatives; homolog of E. coli UbiX | Putative protein of unknown function | Protein required for inositol prototrophy, identified as an ortholog of the FIT family of proteins involved in triglyceride droplet biosynthesis; disputed role in the synthesis of inositol phospholipids from inositol | Protein containing zinc fingers, involved in repression, along with Mig1p, of SUC2 (invertase) expression by high levels of glucose; binds to Mig1p-binding sites in SUC2 promoter | Putative protein of unknown function; not an essential gene | Protein with similarity to cyclin-dependent kinase inhibitors, overproduction suppresses a plc1 null mutation; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Essential chromatin-associated protein involved in the initiation of DNA replication; required for the association of the MCM2-7 complex with replication origins | Subunit of TORC2 (Tor2p-Lst8p-Avo1-Avo2-Tsc11p-Bit61p-Slm1p-Slm2p), a membrane-associated complex that regulates cell cycle-dependent actin cytoskeletal dynamics during polarized growth and cell wall integrity |
| 37 | 51 | 432 | 1.321 | 0.640 |  |  | 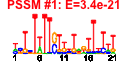 | 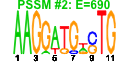 |  | YBL020W YBR069C YBR161W YDL128W YDR044W YDR072C YDR232W YDR345C YDR367W YDR518W YFL031W YFR055W YGL213C YGL215W YGR049W YGR216C YGR276C YHL014C YHR032W YHR049W YHR094C YHR175W YIL145C YJL193W YJR001W YLR130C YLR332W YMR013C YMR274C YNL024C YNL078W YNL259C YNL268W YNR055C YNR061C YOL002C YOL014W YOL103W YOL105C YOR045W YOR099W YOR251C YOR360C YPL053C YPL179W YPL251W YPR138C YDR531W YHR007C YHR061C YLR407W | RFT1 TAT1 CSH1 VCX1 HEM13 IPT1 HEM1 HXT3 EUG1 HAC1 IRC7 SKI8 CLG1 SCM4 GPI1 RNH70 YLF2 FSH1 HXT1 CTR2 PAN6 AVT1 ZRT2 MID2 SEC59 RCE1 YNL024C-A NIS1 ATX1 LYP1 HOL1 IZH2 YOL103W-B WSC3 TOM6 KTR1 PDE2 KTR6 PPQ1 MEP3 ERG11 GIC1 | Flippase, essential integral membrane protein that is required for translocation of Man5GlcNac2-PP-Dol from the cytoplasmic side to the lumenal side of the ER membrane; mutation is suppressed by expression human p53 protein | Amino acid transport protein for valine, leucine, isoleucine, and tyrosine, low-affinity tryptophan and histidine transporter; overexpression confers FK506 and FTY720 resistance | Probable catalytic subunit of a mannosylinositol phosphorylceramide (MIPC) synthase, forms a complex with probable regulatory subunit Csg2p; function in sphingolipid biosynthesis is overlapping with that of Sur1p | Vacuolar H+/Ca2+ exchanger involved in control of cytosolic Ca2+ concentration; has similarity to sodium/calcium exchangers, including the bovine Na+/Ca2+,K+ antiporter | Coproporphyrinogen III oxidase, an oxygen requiring enzyme that catalyzes the sixth step in the heme biosynthetic pathway; localizes to the mitochondrial inner membrane; transcription is repressed by oxygen and heme (via Rox1p and Hap1p) | Inositolphosphotransferase 1, involved in synthesis of mannose-(inositol-P)2-ceramide (M(IP)2C), which is the most abundant sphingolipid in cells, mutation confers resistance to the antifungals syringomycin E and DmAMP1 in some growth media | 5-aminolevulinate synthase, catalyzes the first step in the heme biosynthetic pathway; an N-terminal signal sequence is required for localization to the mitochondrial matrix; expression is regulated by Hap2p-Hap3p | Low affinity glucose transporter of the major facilitator superfamily, expression is induced in low or high glucose conditions | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm in a punctate pattern | Protein disulfide isomerase of the endoplasmic reticulum lumen, function overlaps with that of Pdi1p; may interact with nascent polypeptides in the ER | bZIP transcription factor (ATF/CREB1 homolog) that regulates the unfolded protein response, via UPRE binding, and membrane biogenesis; ER stress-induced splicing pathway utilizing Ire1p, Trl1p and Ada5p facilitates efficient Hac1p synthesis | Putative cystathionine beta-lyase; involved in copper ion homeostasis and sulfur metabolism; null mutant displays increased levels of spontaneous Rad52 foci; expression induced by nitrogen limitation in a GLN3, GAT1-dependent manner | Protein involved in exosome mediated 3' to 5' mRNA degradation and translation inhibition of non-poly(A) mRNAs as well as double-strand break formation during meiotic recombination; required for repressing propagation of dsRNA viruses | Cyclin-like protein that interacts with Pho85p; has sequence similarity to G1 cyclins PCL1 and PCL2 | Potential regulatory effector of CDC4 function, suppresses a temperature-sensitive allele of CDC4, tripartite protein structure in which a charged region separates two uncharged domains, not essential for mitosis or meiosis | Membrane protein involved in the synthesis of N-acetylglucosaminyl phosphatidylinositol (GlcNAc-PI), the first intermediate in the synthesis of glycosylphosphatidylinositol (GPI) anchors; human and mouse GPI1p are functional homologs | 3' exoribonuclease, required for 5S rRNA and tRNA-Arg3 maturation | Protein of unknown function, has weak similarity to E. coli GTP-binding protein gtp1; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Putative protein of unknown function; putative substrate of the cAMP-dependent protein kinase (PKA) | Serine hydrolase that localizes to both the nucleus and cytoplasm; sequence is similar to Fsh2p and Fsh3p | Low-affinity glucose transporter of the major facilitator superfamily, expression is induced by Hxk2p in the presence of glucose and repressed by Rgt1p when glucose is limiting | Putative low-affinity copper transporter of the vacuolar membrane; mutation confers resistance to toxic copper concentrations, while overexpression confers resistance to copper starvation | Pantothenate synthase, also known as pantoate-beta-alanine ligase, required for pantothenic acid biosynthesis, deletion causes pantothenic acid auxotrophy, homologous to E. coli panC | Putative protein of unknown function, predicted to encode a triose phosphate transporter subfamily member based on phylogenetic analysis; similar to YOR307C/SLY41; deletion mutant has a respiratory growth defect | Vacuolar transporter, imports large neutral amino acids into the vacuole; member of a family of seven S. cerevisiae genes (AVT1-7) related to vesicular GABA-glycine transporters | Low-affinity zinc transporter of the plasma membrane; transcription is induced under low-zinc conditions by the Zap1p transcription factor | O-glycosylated plasma membrane protein that acts as a sensor for cell wall integrity signaling and activates the pathway; interacts with Rom2p, a guanine nucleotide exchange factor for Rho1p, and with cell integrity pathway protein Zeo1p | Dolichol kinase, catalyzes the terminal step in dolichyl monophosphate (Dol-P) biosynthesis; required for viability and for normal rates of lipid intermediate synthesis and protein N-glycosylation | Type II CAAX prenyl protease involved in the proteolysis and maturation of Ras and the a-factor mating pheromone | Putative protein of unknown function; YNL024C-A is an essential gene | Protein localized in the bud neck at G2/M phase; physically interacts with septins; possibly involved in a mitotic signaling network | Cytosolic copper metallochaperone that transports copper to the secretory vesicle copper transporter Ccc2p for eventual insertion into Fet3p, which is a multicopper oxidase required for high-affinity iron uptake | Lysine permease; one of three amino acid permeases (Alp1p, Can1p, Lyp1p) responsible for uptake of cationic amino acids | Putative transporter in the major facilitator superfamily (DHA1 family) of multidrug resistance transporters; mutations in membrane-spanning domains permit cation and histidinol uptake | Putative protein of unknown function | Plasma membrane protein involved in zinc metabolism and osmotin-induced apoptosis; transcription regulated by Zap1p, zinc and fatty acid levels; similar to mammalian adiponectins; deletion increases sensitivity to elevated zinc | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Partially redundant sensor-transducer of the stress-activated PKC1-MPK1 signaling pathway involved in maintenance of cell wall integrity; involved in the response to heat shock and other stressors; regulates 1,3-beta-glucan synthesis | Component of the TOM (translocase of outer membrane) complex responsible for recognition and initial import steps for all mitochondrially directed proteins; promotes assembly and stability of the TOM complex | Alpha-1,2-mannosyltransferase involved in O- and N-linked protein glycosylation; type II membrane protein; member of the KRE2/MNT1 mannosyltransferase family | Putative protein of unknown function with similarity to human thiosulfate sulfurtransferase, an enzyme that catalyzes the transfer of sulfur from thiosulfate to cyanide, forming sulfite and thiocyanate; YOR251C is a non-essential gene | High-affinity cyclic AMP phosphodiesterase, component of the cAMP-dependent protein kinase signaling system, protects the cell from extracellular cAMP, contains readthrough motif surrounding termination codon | Probable mannosylphosphate transferase involved in the synthesis of core oligosaccharides in protein glycosylation pathway; member of the KRE2/MNT1 mannosyltransferase family | Putative protein serine/threonine phosphatase; null mutation enhances efficiency of translational suppressors | Ammonium permease of high capacity and low affinity; belongs to a ubiquitous family of cytoplasmic membrane proteins that transport only ammonium (NH4+); expression is under the nitrogen catabolite repression regulation ammonia permease | Pantothenate kinase (ATP:D-pantothenate 4'-phosphotransferase, EC 2.7.1.33); catalyzes the first committed step in the universal biosynthetic pathway leading to coenzyme A1 | Lanosterol 14-alpha-demethylase, catalyzes the C-14 demethylation of lanosterol to form 4,4''-dimethyl cholesta-8,14,24-triene-3-beta-ol in the ergosterol biosynthesis pathway; member of the cytochrome P450 family | Protein of unknown function involved in initiation of budding and cellular polarization, interacts with Cdc42p via the Cdc42/Rac-interactive binding (CRIB) domain | Putative protein of unknown function; YLR407W is not an essential gene |
| 248 | 46 | 446 | 1.332 | 0.658 | 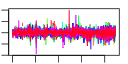 |  | 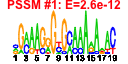 | 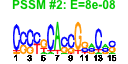 |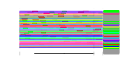  | YDR264C YFL066C YGR180C YJL026W YLR108C YLR214W YLR302C YLR315W YPL274W YBL100C YBR207W YCLX09W YDR270W YDR271C YDR455C YDR515W YDR534C YEL065W YER081W YER088C YER145C YGR065C YGR087C YGR281W YHL035C YHL040C YHL047C YIL117C YKR052C YLL051C YLR034C YLR136C YMR058W YMR275C YMR276W YNL176C YNL321W YOL122C YOL158C YOR226C YOR316C YOR382W YOR383C YOR389W YPR024W YPR123C | AKR1 RNR4 RNR2 FRE1 NKP2 SAM3 FTH1 CCC2 SLF1 FIT1 SIT1 SER3 DOT6 FTR1 VHT1 PDC6 YOR1 VMR1 ARN1 TAF1 PRM5 MRS4 FRE6 SMF3 TIS11 FET3 RDS1 DSK2 VNX1 SMF1 ENB1 ISU2 COT1 FIT2 FIT3 YME1 | Palmitoyl transferase involved in protein palmitoylation; acts as a negative regulator of pheromone response pathway; required for endocytosis of pheromone receptors; involved in cell shape control; contains ankyrin repeats | Helicase-like protein encoded within the telomeric Y' element | Ribonucleotide-diphosphate reductase (RNR), small subunit; the RNR complex catalyzes the rate-limiting step in dNTP synthesis and is regulated by DNA replication and DNA damage checkpoint pathways via localization of the small subunits | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; YLR108C is not an esssential gene | Ferric reductase and cupric reductase, reduces siderophore-bound iron and oxidized copper prior to uptake by transporters; expression induced by low copper and iron levels | Non-essential kinetochore protein, subunit of the Ctf19 central kinetochore complex (Ctf19p-Mcm21p-Okp1p-Mcm22p-Mcm16p-Ctf3p-Chl4p-Mcm19p- Nkp1p-Nkp2p-Ame1p-Mtw1p) | High-affinity S-adenosylmethionine permease, required for utilization of S-adenosylmethionine as a sulfur source; has similarity to S-methylmethionine permease Mmp1p | Putative high affinity iron transporter involved in transport of intravacuolar stores of iron; forms complex with Fet5p; expression is regulated by iron; proposed to play indirect role in endocytosis | Cu(+2)-transporting P-type ATPase, required for export of copper from the cytosol into an extracytosolic compartment; has similarity to human proteins involved in Menkes and Wilsons diseases | RNA binding protein that associates with polysomes; proposed to be involved in regulating mRNA translation; involved in the copper-dependent mineralization of copper sulfide complexes on cell surface in cells cultured in copper salts | Mannoprotein that is incorporated into the cell wall via a glycosylphosphatidylinositol (GPI) anchor, involved in the retention of siderophore-iron in the cell wall | Ferrioxamine B transporter, member of the ARN family of transporters that specifically recognize siderophore-iron chelates; transcription is induced during iron deprivation and diauxic shift; potentially phosphorylated by Cdc28p | 3-phosphoglycerate dehydrogenase, catalyzes the first step in serine and glycine biosynthesis; isozyme of Ser33p | Protein of unknown function, involved in telomeric gene silencing and filamentation | High affinity iron permease involved in the transport of iron across the plasma membrane; forms complex with Fet3p; expression is regulated by iron | High-affinity plasma membrane H+-biotin (vitamin H) symporter; mutation results in fatty acid auxotrophy; 12 transmembrane domain containing major facilitator subfamily member; mRNA levels negatively regulated by iron deprivation and biotin | Minor isoform of pyruvate decarboxylase, decarboxylates pyruvate to acetaldehyde, involved in amino acid catabolism; transcription is glucose- and ethanol-dependent, and is strongly induced during sulfur limitation | Plasma membrane ATP-binding cassette (ABC) transporter, multidrug transporter mediates export of many different organic anions including oligomycin | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; member of the ATP-binding cassette (ABC) family; potential Cdc28p substrate; detected in purified mitochondria in high-throughput studies | Transporter, member of the ARN family of transporters that specifically recognize siderophore-iron chelates; responsible for uptake of iron bound to ferrirubin, ferrirhodin, and related siderophores | TFIID subunit (145 kDa), involved in RNA polymerase II transcription initiation, has histone acetyltransferase activity, involved in promoter binding and G1/S progression | Pheromone-regulated protein, predicted to have 1 transmembrane segment; induced during cell integrity signaling | Mitochondrial iron transporter of the mitochondrial carrier family (MCF), very similar to and functionally redundant with Mrs3p; functions under low-iron conditions; may transport other cations in addition to iron | Putative ferric reductase with similarity to Fre2p; expression induced by low iron levels | Putative divalent metal ion transporter involved in iron homeostasis; transcriptionally regulated by metal ions; member of the Nramp family of metal transport proteins | mRNA-binding protein expressed during iron starvation; binds to a sequence element in the 3'-untranslated regions of specific mRNAs to mediate their degradation; involved in iron homeostasis | Ferro-O2-oxidoreductase required for high-affinity iron uptake and involved in mediating resistance to copper ion toxicity, belongs to class of integral membrane multicopper oxidases | Zinc cluster transcription factor involved in conferring resistance to cycloheximide | Nuclear-enriched ubiquitin-like polyubiquitin-binding protein, required for spindle pole body (SPB) duplication and for transit through the G2/M phase of the cell cycle, involved in proteolysis, interacts with the proteasome | Cell cycle-regulated gene of unknown function, promoter bound by Fkh2p | Low affinity vacuolar membrane localized monovalent cation/H+ antiporter; member of the calcium exchanger (CAX) family; potential Cdc28p substrate | Divalent metal ion transporter with a broad specificity for di-valent and tri-valent metals; post-translationally regulated by levels of metal ions; member of the Nramp family of metal transport proteins | Endosomal ferric enterobactin transporter, expressed under conditions of iron deprivation; member of the major facilitator superfamily; expression is regulated by Rcs1p and affected by chloroquine treatment | Conserved protein of the mitochondrial matrix, required for synthesis of mitochondrial and cytosolic iron-sulfur proteins, performs a scaffolding function in mitochondria during Fe/S cluster assembly; isu1 isu2 double mutant is inviable | Vacuolar transporter that mediates zinc transport into the vacuole; overexpression confers resistance to cobalt and rhodium | Putative protein of unknown function; expression regulated by copper levels | Subunit, with Mgr1p, of the mitochondrial inner membrane i-AAA protease complex, which is responsible for degradation of unfolded or misfolded mitochondrial gene products; mutation causes an elevated rate of mitochondrial turnover |
| 259 | 37 | 443 | 1.353 | 0.633 | 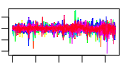 |  |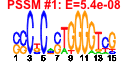  |  |  | YCLX10C YOL054W YBL012C YBR174C YBR244W YCRX19W YDL077C YDL190C YDR355C YGL039W YGR039W YJL028W YJL114W YJR122W YKL044W YKR053C YKR061W YKR071C YLL047W YLR108C YLR202C YLR302C YML090W YMR003W YNL013C YNL105W YNL134C YNL276C YNL296W YOL046C YOR088W YOR104W YOR225W YOR237W YOR282W YPL111W YPR014C | PSH1 GPX2 VAM6 UFD2 IBA57 YSR3 KTR2 DRE2 AIM34 YVC1 PIN2 HES1 CAR1 | Nuclear protein, putative RNA polymerase II elongation factor; isolated as Pob3p/Spt16p-binding protein | Phospholipid hydroperoxide glutathione peroxidase induced by glucose starvation that protects cells from phospholipid hydroperoxides and nonphospholipid peroxides during oxidative stress | Vacuolar protein that plays a critical role in the tethering steps of vacuolar membrane fusion by facilitating guanine nucleotide exchange on small guanosine triphosphatase Ypt7p | Ubiquitin chain assembly factor (E4) that cooperates with a ubiquitin-activating enzyme (E1), a ubiquitin-conjugating enzyme (E2), and a ubiquitin protein ligase (E3) to conjugate ubiquitin to substrates; also functions as an E3 | Oxidoreductase, catalyzes NADPH-dependent reduction of the bicyclic diketone bicyclo[2.2.2]octane-2,6-dione (BCO2,6D) to the chiral ketoalcohol (1R,4S,6S)-6-hydroxybicyclo[2.2.2]octane-2-one (BCO2one6ol) | Protein of unknown function; may interact with ribosomes, based on co-purification experiments | Retrotransposon TYA Gag gene co-transcribed with TYB Pol; translated as TYA or TYA-TYB polyprotein; Gag is a nucleocapsid protein that is the structural constituent of virus-like particles (VLPs); similar to retroviral Gag | Mitochondrial matrix protein involved in the incorporation of iron-sulfur clusters into mitochondrial aconitase-type proteins; activates the radical-SAM family members Bio2p and Lip5p; interacts with Ccr4p in the two-hybrid system | Dihydrosphingosine 1-phosphate phosphatase, membrane protein involved in sphingolipid metabolism; has similarity to Lcb3p | Mannosyltransferase involved in N-linked protein glycosylation; member of the KRE2/MNT1 mannosyltransferase family | Protein of unknown function; mutation displays synthetic lethal interaction with the pol3-13 allele of CDC2 | Protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; YLR108C is not an esssential gene | Protein of unknown function; GFP-fusion protein localizes to the mitochondria; null mutant is viable and displays decreased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Putative protein of unknown function with similarity to dehydrogenases from other model organisms; green fluorescent protein (GFP)-fusion protein localizes to both the cytoplasm and nucleus and is induced by the DNA-damaging agent MMS | Vacuolar cation channel, mediates release of Ca(2+) from the vacuole in response to hyperosmotic shock | Protein that induces appearance of [PIN+] prion when overproduced | Protein implicated in the regulation of ergosterol biosynthesis; one of a seven member gene family with a common essential function and non-essential unique functions; similar to human oxysterol binding protein (OSBP) | Arginase, responsible for arginine degradation, expression responds to both induction by arginine and nitrogen catabolite repression; disruption enhances freeze tolerance |
| 256 | 45 | 445 | 1.359 | 0.642 |  |  |  |  |  | YDR008C YDR327W YER174C YER186C YJR055W YLR261C YNL035C YOL008W YOR263C YPL225W YBL071C YBR138C YCL012W YDL005C YDL216C YDL230W YDR185C YDR445C YFL066C YGR092W YHL028W YHR100C YJL035C YJR113C YKL019W YKL190W YKR047W YLR112W YLR315W YLR316C YLR317W YLR348C YLR349W YLR399C YNL028W YNL046W YNL225C YOL095C YOR013W YOR175C YPL035C YPL095C YPR011C YPR038W YPR106W | GRX4 HIT1 COQ10 YBL071C-B BUD3 MED2 RRI1 PTP1 DBF2 WSC4 TAD2 RSM7 RAM2 CNB1 NKP2 TAD3 DIC1 BDF1 CNM67 HMI1 ALE1 EEB1 ISR1 | Hydroperoxide and superoxide-radical responsive glutathione-dependent oxidoreductase; monothiol glutaredoxin subfamily member along with Grx3p and Grx5p; protects cells from oxidative damage | Putative protein of unknown function | Protein of unknown function, required for growth at high temperature | Putative protein of unknown function with similarity to proteins containing WD-40 domains; green fluorescent protein (GFP)-fusion protein localizes to the nucleus; YNL035C is not an essential gene | Coenzyme Q (ubiquinone) binding protein, functions in the delivery of Q6 to its proper location for electron transport during respiration; START domain protein with homologs in bacteria and eukaryotes | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; green fluorescent protein (GFP)-fusion protein localizes to the cytoplasm | Putative protein of unknown function; identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching | Cytoplasmic protein of unknown function, potentially phosphorylated by Cdc28p; YBR138C is not an essential gene | Protein involved in bud-site selection and required for axial budding pattern; localizes with septins to bud neck in mitosis and may constitute an axial landmark for next round of budding | Subunit of the RNA polymerase II mediator complex; associates with core polymerase subunits to form the RNA polymerase II holoenzyme; essential for transcriptional regulation | Catalytic subunit of the COP9 signalosome (CSN) complex that acts as an isopeptidase in cleaving the ubiquitin-like protein Nedd8 from SCF ubiquitin ligases; metalloendopeptidase involved in the adaptation to pheromone signaling | Phosphotyrosine-specific protein phosphatase that dephosphorylates a broad range of substrates in vivo, including Fpr3p; localized to the cytoplasm and the mitochondria | Mitochondrial protein of unknown function; has similarity to Ups1p, which is involved in regulation of alternative topogenesis of the dynamin-related GTPase Mgm1p | Helicase-like protein encoded within the telomeric Y' element | Ser/Thr kinase involved in transcription and stress response; functions as part of a network of genes in exit from mitosis; localization is cell cycle regulated; activated by Cdc15p during the exit from mitosis | ER membrane protein involved in the translocation of soluble secretory proteins and insertion of membrane proteins into the ER membrane; may also have a role in the stress response but has only partial functional overlap with WSC1-3 | Putative protein of unknown function, required for respiratory growth; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Subunit of tRNA-specific adenosine-34 deaminase, forms a heterodimer with Tad3p that converts adenosine to inosine at the wobble position of several tRNAs | Mitochondrial ribosomal protein of the small subunit, has similarity to E. coli S7 ribosomal protein | Alpha subunit of both the farnesyltransferase and type I geranylgeranyltransferase that catalyze prenylation of proteins containing a CAAX consensus motif; essential protein required for membrane localization of Ras proteins and a-factor | Calcineurin B; the regulatory subunit of calcineurin, a Ca++/calmodulin-regulated type 2B protein phosphatase which regulates Crz1p (a stress-response transcription factor), the other calcineurin subunit is encoded by CNA1 and/or CMP1 | Non-essential kinetochore protein, subunit of the Ctf19 central kinetochore complex (Ctf19p-Mcm21p-Okp1p-Mcm22p-Mcm16p-Ctf3p-Chl4p-Mcm19p- Nkp1p-Nkp2p-Ame1p-Mtw1p) | Subunit of tRNA-specific adenosine-34 deaminase, forms a heterodimer with Tad2p that converts adenosine to inosine at the wobble position of several tRNAs | Mitochondrial dicarboxylate carrier, integral membrane protein, catalyzes a dicarboxylate-phosphate exchange across the inner mitochondrial membrane, transports cytoplasmic dicarboxylates into the mitochondrial matrix | Protein involved in transcription initiation at TATA-containing promoters; associates with the basal transcription factor TFIID; contains two bromodomains; corresponds to the C-terminal region of mammalian TAF1; redundant with Bdf2p | Putative protein of unknown function; expression depends on Swi5p; GFP-fusion protein localizes to the endoplasmic reticulum; deletion confers sensitivity to 4-(N-(S-glutathionylacetyl)amino) phenylarsenoxide (GSAO) | Component of the spindle pole body outer plaque; required for spindle orientation and mitotic nuclear migration | Mitochondrial inner membrane localized ATP-dependent DNA helicase, required for the maintenance of the mitochondrial genome; not required for mitochondrial transcription; has homology to E. coli helicase uvrD | Lysophospholipid acyltransferase, partially redundant with Slc1p; part of MBOAT family of membrane-bound O-acyltransferases; key component of Lands cycle; may have role in fatty acid exchange at sn-2 position of mature glycerophospholipids | Acyl-coenzymeA:ethanol O-acyltransferase responsible for the major part of medium-chain fatty acid ethyl ester biosynthesis during fermentation; possesses short chain esterase activity; may be involved in lipid metabolism and detoxification | Putative transporter, member of the mitochondrial carrier family; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies | Predicted protein kinase, overexpression causes sensitivity to staurosporine, which is a potent inhibitor of protein kinase C |
| 4 | 45 | 436 | 1.379 | 0.616 | 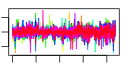 |  |  | 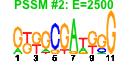 |  | YAL048C YAR068W YBR068C YBR207W YBR235W YDL019C YDL028C YDL034W YDL056W YDL109C YDL214C YDR011W YDR269C YDR270W YDR271C YDR455C YDR456W YDR534C YEL065W YFR030W YGL146C YGR087C YGR097W YHL028W YHL035C YHL040C YHL047C YIL082W YIL169C YJL101C YKR052C YLL051C YLL053C YLR034C YLR136C YLR338W YML087C YOL118C YOL152W YOL158C YOR291W YOR316C YOR382W YOR383C YOR384W | GEM1 BAP2 FTH1 OSH2 MPS1 MBP1 PRR2 SNQ2 CCC2 NHX1 FIT1 SIT1 MET10 PDC6 ASK10 WSC4 VMR1 ARN1 TAF1 YIL082W-A GSH1 MRS4 FRE6 SMF3 TIS11 AIM33 FRE7 ENB1 COT1 FIT2 FIT3 FRE5 | Evolutionarily-conserved tail-anchored outer mitochondrial membrane GTPase which regulates mitochondrial morphology; cells lacking Gem1p contain collapsed, globular, or grape-like mitochondria; not required for pheromone-induced cell death | Fungal-specific protein of unknown function; induced in respiratory-deficient cells | High-affinity leucine permease, functions as a branched-chain amino acid permease involved in the uptake of leucine, isoleucine and valine; contains 12 predicted transmembrane domains | Putative high affinity iron transporter involved in transport of intravacuolar stores of iron; forms complex with Fet5p; expression is regulated by iron; proposed to play indirect role in endocytosis | Putative ion transporter, similar to mammalian electroneutral Na(+)-(K+)-C1- cotransporter family; YBR235W is not an essential gene | Member of an oxysterol-binding protein family with seven members in S. cerevisiae; family members have overlapping, redundant functions in sterol metabolism and collectively perform a function essential for viability | Dual-specificity kinase required for spindle pole body (SPB) duplication and spindle checkpoint function; substrates include SPB proteins Spc42p, Spc110p, and Spc98p, mitotic exit network protein Mob1p, and checkpoint protein Mad1p | Transcription factor involved in regulation of cell cycle progression from G1 to S phase, forms a complex with Swi6p that binds to MluI cell cycle box regulatory element in promoters of DNA synthesis genes | Putative lipase; involved in lipid metabolism; YDL109C is not an essential gene | Serine/threonine protein kinase that inhibits pheromone induced signalling downstream of MAPK, possibly at the level of the Ste12p transcription factor | Plasma membrane ATP-binding cassette (ABC) transporter, multidrug transporter involved in multidrug resistance and resistance to singlet oxygen species | Cu(+2)-transporting P-type ATPase, required for export of copper from the cytosol into an extracytosolic compartment; has similarity to human proteins involved in Menkes and Wilsons diseases | Endosomal Na+/H+ exchanger, required for intracellular sequestration of Na+; required for osmotolerance to acute hypertonic shock | Mannoprotein that is incorporated into the cell wall via a glycosylphosphatidylinositol (GPI) anchor, involved in the retention of siderophore-iron in the cell wall | Ferrioxamine B transporter, member of the ARN family of transporters that specifically recognize siderophore-iron chelates; transcription is induced during iron deprivation and diauxic shift; potentially phosphorylated by Cdc28p | Subunit alpha of assimilatory sulfite reductase, which is responsible for the conversion of sulfite into sulfide | Putative protein of unknown function, contains two putative transmembrane spans, shows no significant homology to any other known protein sequence, YGL146C is not an essential gene | Minor isoform of pyruvate decarboxylase, decarboxylates pyruvate to acetaldehyde, involved in amino acid catabolism; transcription is glucose- and ethanol-dependent, and is strongly induced during sulfur limitation | Component of the RNA polymerase II holoenzyme, phosphorylated in response to oxidative stress; has a role in destruction of Ssn8p, which relieves repression of stress-response genes | ER membrane protein involved in the translocation of soluble secretory proteins and insertion of membrane proteins into the ER membrane; may also have a role in the stress response but has only partial functional overlap with WSC1-3 | Protein of unknown function that may interact with ribosomes, based on co-purification experiments; member of the ATP-binding cassette (ABC) family; potential Cdc28p substrate; detected in purified mitochondria in high-throughput studies | Transporter, member of the ARN family of transporters that specifically recognize siderophore-iron chelates; responsible for uptake of iron bound to ferrirubin, ferrirhodin, and related siderophores | TFIID subunit (145 kDa), involved in RNA polymerase II transcription initiation, has histone acetyltransferase activity, involved in promoter binding and G1/S progression | Retrotransposon TYA Gag and TYB Pol genes; transcribed/translated as one unit; polyprotein is processed to make a nucleocapsid-like protein (Gag), reverse transcriptase (RT), protease (PR), and integrase (IN); similar to retroviral genes | Putative protein of unknown function; serine/threonine rich and highly similar to YOL155C, a putative glucan alpha-1,4-glucosidase; transcript is induced in both high and low pH environments; YIL169C is a non-essential gene | Gamma glutamylcysteine synthetase catalyzes the first step in glutathione (GSH) biosynthesis; expression induced by oxidants, cadmium, and mercury | Mitochondrial iron transporter of the mitochondrial carrier family (MCF), very similar to and functionally redundant with Mrs3p; functions under low-iron conditions; may transport other cations in addition to iron | Putative ferric reductase with similarity to Fre2p; expression induced by low iron levels | Putative protein; in the Sigma 1278B strain background YLL053C is contiguous with AQY2 which encodes an aquaporin | Putative divalent metal ion transporter involved in iron homeostasis; transcriptionally regulated by metal ions; member of the Nramp family of metal transport proteins | mRNA-binding protein expressed during iron starvation; binds to a sequence element in the 3'-untranslated regions of specific mRNAs to mediate their degradation; involved in iron homeostasis | Putative protein of unknown function; null mutant displays increased frequency of mitochondrial genome loss (petite formation) and severe growth defect in minimal glycerol media | Putative ferric reductase with similarity to Fre2p; expression induced by low copper levels | Endosomal ferric enterobactin transporter, expressed under conditions of iron deprivation; member of the major facilitator superfamily; expression is regulated by Rcs1p and affected by chloroquine treatment | Putative protein of unknown function; shares sequence similarity with the type V P-type ATPase Spf1p; YOR291W is not an essential protein | Vacuolar transporter that mediates zinc transport into the vacuole; overexpression confers resistance to cobalt and rhodium | Putative ferric reductase with similarity to Fre2p; expression induced by low iron levels; the authentic, non-tagged protein is detected in highly purified mitochondria in high-throughput studies |
| 28 | 51 | 457 | 1.384 | 0.600 | 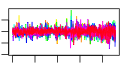 |  | 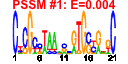 | 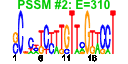 |  | YAR002AC YBL012C YBL053W YBL070C YBL071C YBL106C YBR027C YBR174C YBR232C YCL006C YCL022C YCR049C YCR103C YCRX06W YCRX07W YCRX08W YCRX12W YCRX20C YDL041W YDL094C YDL121C YDR274C YDR281C YDR301W YDR355C YDR431W YDR481C YDR491C YDR537C YER145C YFR034C YFR056C YGL015C YGL217C YGL236C YGR164W YGR176W YGR190C YHR048W YIL141W YJL119C YJL195C YKL116C YKL131W YLR407W YNL028W YNL184C YNL296W YGL263W YGR050C YGR242W | YBL071C-B SRO77 PHM6 CFT1 PHO8 FTR1 PHO4 MTO1 YHK8 PRR1 COS12 | Putative protein of unknown function; identified by gene-trapping, microarray-based expression analysis, and genome-wide homology searching | Protein with roles in exocytosis and cation homeostasis; functions in docking and fusion of post-Golgi vesicles with plasma membrane; homolog of Sro7p and Drosophila lethal giant larvae tumor suppressor; interacts with SNARE protein Sec9p | Putative protein of unknown function; green fluorescent protein (GFP)-fusion protein localizes to the endoplasmic retiuculum; YDL121C is not an essential protein | Protein of unknown function, expression is regulated by phosphate levels | RNA-binding subunit of the mRNA cleavage and polyadenylation factor; involved in poly(A) site recognition and required for both pre-mRNA cleavage and polyadenylation, 51% sequence similarity with mammalian AAUAA-binding subunit of CPSF | Repressible alkaline phosphatase, a glycoprotein localized to the vacuole; regulated by levels of inorganic phosphate and by a system consisting of Pho4p, Pho9p, Pho80p, Pho81p and Pho85p; dephosphorylates phosphotyrosyl peptides | High affinity iron permease involved in the transport of iron across the plasma membrane; forms complex with Fet3p; expression is regulated by iron | Basic helix-loop-helix (bHLH) transcription factor of the myc-family; binds cooperatively with Pho2p to the PHO5 promoter; function is regulated by phosphorylation at multiple sites and by phosphate availability | hypothetical protein | Mitochondrial protein, forms a heterodimer complex with Mss1p that performs the 5-carboxymethylaminomethyl modification of the wobble uridine base in mitochondrial tRNAs; required for respiration in paromomycin-resistant 15S rRNA mutants | Presumed antiporter of the DHA1 family of multidrug resistance transporters; contains 12 predicted transmembrane spans; expression of gene is up-regulated in cells exhibiting reduced susceptibility to azoles | Serine/threonine protein kinase that inhibits pheromone induced signalling downstream of MAPK, possibly at the level of the Ste12p transcription factor | Putative protein of unknown function; YLR407W is not an essential gene | Protein of unknown function, member of the DUP380 subfamily of conserved, often subtelomerically-encoded proteins |
| 282 | 40 | 449 | 1.811 | 0.676 |  |  | 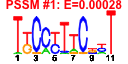 |  |  | YCL058C YIL047C YAL040C YBR067C YBR074W YCLX11W YDL036C YDL066W YDL073W YDR089W YDR094W YDR149C YDR194C YDR430C YDR432W YDR433W YDR461W YFL032W YGL040C YGL149W YGL239C YGR035C YGR259C YHR001W YHR218W YJL175W YKL030W YLR440C YNL065W YNL184C YNL226W YNL231C YNR042W YOL027C YOR011W YOR123C YOR262W YPL075W YPL274W YPR035W | SYG1 CLN3 TIP1 PUS9 IDP1 MSS116 CYM1 NPL3 MFA1 HEM2 OSH7 SEC39 AQR1 PDR16 MDM38 AUS1 LEO1 GCR1 SAM3 GLN1 | Plasma membrane protein of unknown function; truncation and overexpression suppresses lethality of G-alpha protein deficiency | G1 cyclin involved in cell cycle progression; activates Cdc28p kinase to promote the G1 to S phase transition; plays a role in regulating transcription of the other G1 cyclins, CLN1 and CLN2; regulated by phosphorylation and proteolysis | Major cell wall mannoprotein with possible lipase activity; transcription is induced by heat- and cold-shock; member of the Srp1p/Tip1p family of serine-alanine-rich proteins | Putative metalloprotease | Mitochondrial tRNA pseudouridine synthase involved in pseudouridylation of mitochondrial tRNAs at position 32 | Mitochondrial NADP-specific isocitrate dehydrogenase, catalyzes the oxidation of isocitrate to alpha-ketoglutarate; not required for mitochondrial respiration and may function to divert alpha-ketoglutarate to biosynthetic processes | Putative protein of unknown function; YDL073W is not an essential gene | Protein of unknown function; deletion confers resistance to Nickel | DEAD-box protein required for efficient splicing of mitochondrial Group I and II introns; non-polar RNA helicase that also facilities strand annealing | Lysine-specific metalloprotease of the mitochondrial intermembrane space, member of the pitrilysin family; degrades proteins and presequence peptides cleaved from imported proteins; required for normal mitochondrial morphology | RNA-binding protein that carries poly(A)+ mRNA from the nucleus into the cytoplasm; dissociation from mRNAs is promoted by Mtr10p; phosphorylated by Sky1p in the cytoplasm | Mating pheromone a-factor, made by a cells; interacts with alpha cells to induce cell cycle arrest and other responses leading to mating; biogenesis involves C-terminal modification, N-terminal proteolysis, and export; also encoded by MFA2 | Delta-aminolevulinate dehydratase, a homo-octameric enzyme, catalyzes the conversion of delta-aminolevulinic acid to porphobilinogen, the second step in the heme biosynthetic pathway; localizes to both the cytoplasm and nucleus | Putative protein of unknown function, potential Cdc28p substrate; transcription is activated by paralogous transcription factors Yrm1p and Yrr1p along with genes involved in multidrug resistance | Member of an oxysterol-binding protein family with seven members in S. cerevisiae; family members have overlapping, redundant functions in sterol metabolism and collectively perform a function essential for viability | Helicase-like protein encoded within the telomeric Y' element | Protein of unknown function proposed to be involved in protein secretion; interacts with Dsl1p and localizes to the ER and nuclear envelope | Plasma membrane multidrug transporter of the major facilitator superfamily, confers resistance to short-chain monocarboxylic acids and quinidine; involved in the excretion of excess amino acids | Phosphatidylinositol transfer protein (PITP) controlled by the multiple drug resistance regulator Pdr1p, localizes to lipid particles and microsomes, controls levels of various lipids, may regulate lipid synthesis, homologous to Pdr17p | Mitochondrial inner membrane protein, involved in membrane integration of a subset of mitochondrial proteins; required for K+/H+ exchange; associates with mitochondrial ribosomes; human ortholog Letm1 implicated in Wolf-Hirschhorn syndrome | Transporter of the ATP-binding cassette family, involved in uptake of sterols and anaerobic growth | Component of the Paf1 complex, which associates with RNA polymerase II and is involved in histone methylation | Cytoplasmic protein of unknown function; essential gene with similarity to YLR243W; contains an ATP/GTP binding site motif | Transcriptional activator of genes involved in glycolysis; DNA-binding protein that interacts and functions with the transcriptional activator Gcr2p | High-affinity S-adenosylmethionine permease, required for utilization of S-adenosylmethionine as a sulfur source; has similarity to S-methylmethionine permease Mmp1p | Glutamine synthetase (GS), synthesizes glutamine from glutamate and ammonia; with Glt1p, forms the secondary pathway for glutamate biosynthesis from ammonia; expression regulated by nitrogen source and by amino acid limitation |






































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































































